Abstract
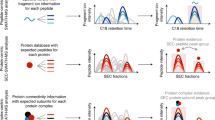
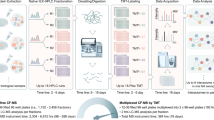
Targeted proteomics approaches are of value for deep and accurate quantification of protein abundance. Extending such methods to quantify large numbers of proteins requires the construction of predefined targeted assays. We developed a targeted proteomics platform—in vitro proteome–assisted multiple reaction monitoring (MRM) for protein absolute quantification (iMPAQT)—by using >18,000 human recombinant proteins, thus enabling protein absolute quantification on a genome-wide scale. Our platform comprises experimentally confirmed MRM assays of mass tag (mTRAQ)-labeled peptides to allow for rapid and straightforward measurement of the absolute abundance of predefined sets of proteins by mass spectrometry. We applied iMPAQT to delineate the quantitative metabolic landscape of normal and transformed human fibroblasts. Oncogenic transformation gave rise to relatively small but global changes in metabolic pathways resulting in aerobic glycolysis (Warburg effect) and increased rates of macromolecule synthesis. iMPAQT should facilitate quantitative biology studies based on protein abundance measurements.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ishii, N. et al. Multiple high-throughput analyses monitor the response of E. coli to perturbations. Science 316, 593–597 (2007).
Ideker, T. et al. Integrated genomic and proteomic analyses of a systematically perturbed metabolic network. Science 292, 929–934 (2001).
Buescher, J.M. et al. Global network reorganization during dynamic adaptations of Bacillus subtilis metabolism. Science 335, 1099–1103 (2012).
Anderson, N.L. et al. A human proteome detection and quantitation project. Mol. Cell. Proteomics 8, 883–886 (2009).
Keshishian, H., Addona, T., Burgess, M., Kuhn, E. & Carr, S.A. Quantitative, multiplexed assays for low abundance proteins in plasma by targeted mass spectrometry and stable isotope dilution. Mol. Cell. Proteomics 6, 2212–2229 (2007).
Lange, V., Picotti, P., Domon, B. & Aebersold, R. Selected reaction monitoring for quantitative proteomics: a tutorial. Mol. Syst. Biol. 4, 222 (2008).
Picotti, P., Bodenmiller, B., Mueller, L.N., Domon, B. & Aebersold, R. Full dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell 138, 795–806 (2009).
Picotti, P. et al. High-throughput generation of selected reaction-monitoring assays for proteins and proteomes. Nat. Methods 7, 43–46 (2010).
Kennedy, J.J. et al. Demonstrating the feasibility of large-scale development of standardized assays to quantify human proteins. Nat. Methods 11, 149–155 (2014).
Deutsch, E.W., Lam, H. & Aebersold, R. PeptideAtlas: a resource for target selection for emerging targeted proteomics workflows. EMBO Rep. 9, 429–434 (2008).
Wilhelm, M. et al. Mass-spectrometry-based draft of the human proteome. Nature 509, 582–587 (2014).
Kim, M.S. et al. A draft map of the human proteome. Nature 509, 575–581 (2014).
Mallick, P. et al. Computational prediction of proteotypic peptides for quantitative proteomics. Nat. Biotechnol. 25, 125–131 (2007).
Fusaro, V.A., Mani, D.R., Mesirov, J.P. & Carr, S.A. Prediction of high-responding peptides for targeted protein assays by mass spectrometry. Nat. Biotechnol. 27, 190–198 (2009).
Karlsson, C., Malmström, L., Aebersold, R. & Malmström, J. Proteome-wide selected reaction monitoring assays for the human pathogen Streptococcus pyogenes. Nat. Commun. 3, 1301 (2012).
Picotti, P. et al. A complete mass-spectrometric map of the yeast proteome applied to quantitative trait analysis. Nature 494, 266–270 (2013).
Schubert, O.T. et al. The Mtb proteome library: a resource of assays to quantify the complete proteome of Mycobacterium tuberculosis. Cell Host Microbe 13, 602–612 (2013).
Kusebauch, U. et al. Human SRMAtlas: a resource of targeted assays to quantify the complete human proteome. Cell 166, 766–778 (2016).
Zeiler, M., Straube, W.L., Lundberg, E., Uhlen, M. & Mann, M. A Protein Epitope Signature Tag (PrEST) library allows SILAC-based absolute quantification and multiplexed determination of protein copy numbers in cell lines. Mol. Cell. Proteomics 11, O111 009613 (2012).
Stergachis, A.B., MacLean, B., Lee, K., Stamatoyannopoulos, J.A. & MacCoss, M.J. Rapid empirical discovery of optimal peptides for targeted proteomics. Nat. Methods 8, 1041–1043 (2011).
Goshima, N. et al. Human protein factory for converting the transcriptome into an in vitro–expressed proteome. Nat. Methods 5, 1011–1017 (2008).
Beck, M. et al. The quantitative proteome of a human cell line. Mol. Syst. Biol. 7, 549 (2011).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Abbatiello, S.E. et al. Large-scale interlaboratory study to develop, analytically validate and apply highly multiplexed, quantitative peptide assays to measure cancer-relevant proteins in plasma. Mol. Cell. Proteomics 14, 2357–2374 (2015).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
DeSouza, L.V. et al. Multiple reaction monitoring of mTRAQ-labeled peptides enables absolute quantification of endogenous levels of a potential cancer marker in cancerous and normal endometrial tissues. J. Proteome Res. 7, 3525–3534 (2008).
Takeda, H. et al. Comparative analysis of human SRC-family kinase substrate specificity in vitro. J. Proteome Res. 9, 5982–5993 (2010).
Nagaraj, N. et al. Deep proteome and transcriptome mapping of a human cancer cell line. Mol. Syst. Biol. 7, 548 (2011).
Marguerat, S. et al. Quantitative analysis of fission yeast transcriptomes and proteomes in proliferating and quiescent cells. Cell 151, 671–683 (2012).
Lawless, C. et al. Direct and absolute quantification of over 1800 yeast proteins via selected reaction monitoring. Mol. Cell. Proteomics 15, 1309–1322 (2016).
Kito, K., Okada, M., Ishibashi, Y., Okada, S. & Ito, T. A strategy for absolute proteome quantification with mass spectrometry by hierarchical use of peptide-concatenated standards. Proteomics 16, 1457–1473 (2016).
Hahn, W.C. et al. Creation of human tumour cells with defined genetic elements. Nature 400, 464–468 (1999).
Akagi, T., Sasai, K. & Hanafusa, H. Refractory nature of normal human diploid fibroblasts with respect to oncogene-mediated transformation. Proc. Natl. Acad. Sci. USA 100, 13567–13572 (2003).
Miller, D.M., Thomas, S.D., Islam, A., Muench, D. & Sedoris, K. c-Myc and cancer metabolism. Clin. Cancer Res. 18, 5546–5553 (2012).
Costenoble, R. et al. Comprehensive quantitative analysis of central carbon and amino-acid metabolism in Saccharomyces cerevisiae under multiple conditions by targeted proteomics. Mol. Syst. Biol. 7, 464 (2011).
Sabidó, E. et al. Targeted proteomics reveals strain-specific changes in the mouse insulin and central metabolic pathways after a sustained high-fat diet. Mol. Syst. Biol. 9, 681 (2013).
Soste, M. et al. A sentinel protein assay for simultaneously quantifying cellular processes. Nat. Methods 11, 1045–1048 (2014).
Maruyama, Y. et al. Human Gene and Protein Database (HGPD): a novel database presenting a large quantity of experiment-based results in human proteomics. Nucleic Acids Res. 37, D762–D766 (2009).
Maruyama, Y. et al. HGPD: Human Gene and Protein Database, 2012 update. Nucleic Acids Res. 40, D924–D929 (2012).
Mori, M., Fukuda, E. & Goshima, N. Protein tag, tagged protein, and protein purification method. PTC patent application WO/2013/150680 (2013).
Wessel, D. & Flügge, U.I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. 138, 141–143 (1984).
Storey, J.D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Acknowledgements
This work was supported by CREST, JST, and JSPS KAKENHI (grant nos. 25290078 and 16H04730) and MEXT KAKENHI (grant nos. 24116518, 26116721, 25134714, 25221303, and 15H01407 to M. Matsumoto and K.I.N.). We thank T. Akagi for the pCX4 system and vectors for hTERT, SV40 ER, and RasG12V; T. Takami for scripting tools and database provision; A. Sakiyama, M. Oda, K. Tsunematsu, and E. Koba for technical assistance; members of our laboratories for comments on the manuscript; and A. Ohta for help in preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
M. Matsumoto, F.M., and K. Oshikawa designed and performed experiments, analyzed data, and prepared the manuscript. M. Mori, K. Ogawa, Y.K., E.F., N.G., and T. Natsume generated recombinant proteins. M. Matsumoto, H.N., K.F., and K.H. contributed to development of the informatics pipeline. T. Nagashima, R.F., and K.N. performed RNA sequencing and data analysis. K.I.N. contributed to supervision of the study and writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Processing and quality control for recombinant proteins.
(a) Schematic representation of processing of a recombinant protein library. Recombinant proteins were generated individually. Portions of each protein preparation were subjected to purification followed by SDS-PAGE in order to check protein quality and to determine protein concentration. The remaining portions of the preparations were mixed and purified. The obtained protein mixture was digested and used for peptide identification or as an internal standard for MRM assays. (b) Recombinant proteins were synthesized in the presence of [13C6,15N2]lysine and [13C6,15N4]arginine, digested, and subjected to determination of protein concentration with an MRM assay for a peptide (LGPLVPR) corresponding to the MAFG tag. The log2 ratio of protein concentration determined by SDS-PAGE to that determined by MRM was calculated.
Supplementary Figure 2 DDA of the in vitro proteome achieves unbiased peptide identification.
The percentage of proteins in the in vitro proteome identified by DDA was determined for each of the indicated pathways (a) and molecular functions (b). Proteins were assigned to pathways and molecular functions with the Kyoto Encyclopedia of Genes and Genomes (KEGG) database, and the coverage was then compared with that for data in PeptideAtlas.
Supplementary Figure 3 Labeling efficiency of mTRAQ.
(a) A recombinant protein digest was labeled (or not) with mTRAQΔ4, and the peptides were then subjected to MRM analysis together with a digest of the same protein containing [13C6, 15N2]lysine and [13C6, 15N4]arginine. The MRM chromatogram of the peptide AGAHLQGGAK is shown as an example. The red and blue lines correspond to the non–mTRAQ-labeled peptide and the internal standard containing a [13C6, 15N2]lysine residue, respectively. (b) A whole HeLa cell digest (100 μg) was incubated for 2 h at room temperature with the indicated amounts of mTRAQ reagent, after which the amount of non–mTRAQ-labeled peptides was quantified by MRM analysis together with a digest of proteins containing [13C6, 15N2]lysine and [13C6, 15N4]arginine. The labeling efficiency was calculated from the percentage of non–mTRAQ-labeled peptides remaining after the reaction. Efficiency values are means ± s.d. for an average of 102 peptides in three independent measurements.
Supplementary Figure 4 Properties of nonlabeled and mTRAQ-labeled peptides in MS.
(a) Nonlabeled and mTRAQΔ4-labeled digests of a mixture of recombinant proteins (108 proteins) were subjected to DDA, and identified unique peptide sequences were compared. (b) The numbers of peptides identified from nonlabeled and mTRAQ-labeled digests were compared for each charge state. (c) Patterns of MS/MS spectra for nonlabeled and mTRAQ-lableled peptides were compared with Spearman’s correlation coefficient on the basis of the intensities of all observed fragment ions (left panel) or only y-ions (right panel). (d) MRM assays developed with nonlabeled and mTRAQ-labeled peptides were compared on the basis of signal intensity. Data for the protein ACADVL are presented as a typical example.
Supplementary Figure 5 Validation of MRM-based absolute quantification.
(a) Reproducible detection rate in repeated MRM analysis of 198 peptides. (b) Coefficient of variation for repeated MRM measurement. (c) Correlation between the absolute abundance of 10 proteins estimated by MRM analysis and the corresponding values obtained by immunoblot analysis. (d) Correlation between the absolute abundance of core metabolic enzymes determined by mTRAQ-based or AQUA protein–based strategies.
Supplementary Figure 6 iMPAQT platform.
The iMPAQT platform includes iMPAQT-knowledge DB and iMPAQT-quant software. iMPAQT-knowledge DB contains all MS/MS spectra assigned to peptides, MRM chromatograms for verification, and a search interface based on functional annotation (such as pathway, biological process, molecular function, and cellular localization) or on ID/accession number. Verified MRM transitions for each protein of interest can be downloaded in csv format. Each assay is specified as iMPAQT probe ID. Acquired data are processed by iMPAQT-quant software, which detects co-eluted peaks and calculates cosine similarity by comparison with corresponding MS/MS spectra. iMPAQT-knowledge DB also contains the concentration of each recombinant protein to assist estimation of the absolute abundance of each protein automatically.
Supplementary Figure 7 Absolute quantification of metabolic enzymes in human diploid fibroblasts.
(a) The absolute values for the abundance of metabolic enzymes in HDFs as determined by MRM in a single experiment were compared between culture lots (A and B) and process replicates (1 and 2). (b) RNA abundance as determined by RNA sequencing for proteins not detected in HDFs by MRM analysis. FPKM, fragments per kilobase of exon model per million mapped fragments.
Supplementary Figure 8 Relationship between protein and RNA amounts for metabolic enzymes in mammalian cells and budding yeast.
(a) Comparison of protein copy number per RNA molecule between human (this study) and mouse (Schwanhäusser et al.20). (b) Protein/RNA ratio for orthologs quantified in human (this study), mouse (Schwanhäusser et al.20), and budding yeast (Lawless et al.26).
Supplementary Figure 9 Establishment of oncogene-induced transformation in HDFs.
(a) Immunoblot analysis of expressed oncogenes and Hsp90 (loading control) in parental HDFs (TIG-3 cells), immortalized cells expressing hTERT and SV40 ER proteins (TS cells), and transformed cells expressing hTERT, SV40 ER proteins, and either c-Myc or RasG12V (TSM and TSR cells, respectively). (b) Phase-contrast microscopy of TIG-3 cell lines. Scale bars, 200 μm. (c) Proliferation curves for parental, TS, TSM, and TSR cells. Data are means ± s.d. for three independent determinations. (d) Colony formation by cells cultured in soft agar for 12 days. Scale bars, 200 μm. (e) Number of colonies formed by parental (P) HDFs and derived cell lines in experiments similar to that in d. Data are means ± s.d. for three independent determinations. ND, none detected. (f, g) Lactate production (f) and glucose consumption (g) in parental HDFs and derived cell lines maintained under normoxic or hypoxic conditions. Data are means ± s.d. for three independent experiments. (h, i) Rates of nucleic acid (h) and fatty acid (i) synthesis as determined by measurement of the incorporation of 14C from 14C-labeled glucose into DNA or RNA or into fatty acids, respectively. Data are means ± s.d. for three independent experiments.
Supplementary Figure 10 Significance of transformation-related changes in the abundance of metabolic enzymes.
Volcano plots of fold change in protein abundance and accompanying q-value are shown for TSM cells (a) or TSR cells (b) relative to parental cells. Proteins are color-coded according to metabolic pathway. Red: glycolysis, TCA cycle, and oxidative phosphorylation. Purple: pentose phosphate pathway. Orange: nucleic acid metabolism. Pink: fatty acid metabolism. Blue: serine, glycine, and one-carbon metabolism.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 (PDF 1687 kb)
Supplementary Table 1
Determination of proteotypic peptides using in vitro proteome. (XLSX 29424 kb)
Supplementary Table 2
Protein identification from natural proteome. (XLSX 4388 kb)
Supplementary Table 3
Verification of MRM assays using recombinant proteins. (XLSX 20510 kb)
Supplementary Table 4
MRM transitions for metabolic enzymes. (XLSX 2716 kb)
Supplementary Table 5
Absolute abundance of human metabolic enzyme in HDFs (TIG-3). (XLSX 884 kb)
Supplementary Table 6
Absolute quantification of metabolic enzymes in parental and oncogene expressing cells. (XLSX 1162 kb)
Rights and permissions
About this article
Cite this article
Matsumoto, M., Matsuzaki, F., Oshikawa, K. et al. A large-scale targeted proteomics assay resource based on an in vitro human proteome. Nat Methods 14, 251–258 (2017). https://doi.org/10.1038/nmeth.4116
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4116
This article is cited by
-
Integrative transcriptome and proteome analyses of clear cell renal cell carcinoma develop a prognostic classifier associated with thrombus
Scientific Reports (2023)
-
Convergent genomic diversity and novel BCAA metabolism in intrahepatic cholangiocarcinoma
British Journal of Cancer (2023)
-
Metabolic remodeling of pyrimidine synthesis pathway and serine synthesis pathway in human glioblastoma
Scientific Reports (2022)
-
Proteomics-based target identification of natural products affecting cancer metabolism
The Journal of Antibiotics (2021)
-
BEX2 suppresses mitochondrial activity and is required for dormant cancer stem cell maintenance in intrahepatic cholangiocarcinoma
Scientific Reports (2020)