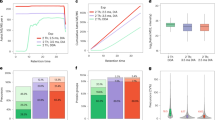
Abstract
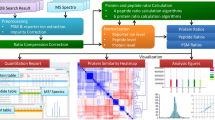
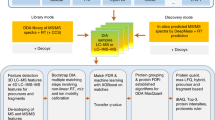
Next-generation mass spectrometric (MS) techniques such as SWATH-MS have substantially increased the throughput and reproducibility of proteomic analysis, but ensuring consistent quantification of thousands of peptide analytes across multiple liquid chromatography–tandem MS (LC-MS/MS) runs remains a challenging and laborious manual process. To produce highly consistent and quantitatively accurate proteomics data matrices in an automated fashion, we developed TRIC (http://proteomics.ethz.ch/tric/), a software tool that utilizes fragment-ion data to perform cross-run alignment, consistent peak-picking and quantification for high-throughput targeted proteomics. TRIC reduced the identification error compared to a state-of-the-art SWATH-MS analysis without alignment by more than threefold at constant recall while correcting for highly nonlinear chromatographic effects. On a pulsed-SILAC experiment performed on human induced pluripotent stem cells, TRIC was able to automatically align and quantify thousands of light and heavy isotopic peak groups. Thus, TRIC fills a gap in the pipeline for automated analysis of massively parallel targeted proteomics data sets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Hudson, T.J. et al. International network of cancer genome projects. Nature 464, 993–998 (2010).
McLendon, R. et al. Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 455, 1061–1068 (2008).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Haines, J.L. et al. Complement factor H variant increases the risk of age-related macular degeneration. Science 308, 419–421 (2005).
International Consortium for Blood Pressure Genome-Wide Association Studies. Genetic variants in novel pathways influence blood pressure and cardiovascular disease risk. Nature 478, 103–109 (2011).
Röst, H.L., Malmström, L. & Aebersold, R. Reproducible quantitative proteotype data matrices for systems biology. Mol. Biol. Cell 26, 3926–3931 (2015).
de Godoy, L.M.F. et al. Comprehensive mass-spectrometry-based proteome quantification of haploid versus diploid yeast. Nature 455, 1251–1254 (2008).
Hebert, A.S. et al. The one hour yeast proteome. Mol. Cell. Proteomics 13, 339–347 (2014).
Beck, M. et al. The quantitative proteome of a human cell line. Mol. Syst. Biol. 7, 549 (2011).
Nagaraj, N. et al. Deep proteome and transcriptome mapping of a human cancer cell line. Mol. Syst. Biol. 7, 548 (2011).
Desiere, F. et al. The PeptideAtlas project. Nucleic Acids Res. 34, D655–D658 (2006).
Omenn, G.S. et al. Metrics for the Human Proteome Project 2015: progress on the human proteome and guidelines for high-confidence protein identification. J. Proteome Res. 14, 3452–3460 (2015).
Picotti, P. et al. A complete mass-spectrometric map of the yeast proteome applied to quantitative trait analysis. Nature 494, 266–270 (2013).
Li, X.J. et al. A blood-based proteomic classifier for the molecular characterization of pulmonary nodules. Sci. Transl. Med. 5, 207ra142 (2013).
Drabovich, A.P. et al. Differential diagnosis of azoospermia with proteomic biomarkers ECM1 and TEX101 quantified in seminal plasma. Sci. Transl. Med. 5, 212ra160 (2013).
Surinova, S. et al. Prediction of colorectal cancer diagnosis based on circulating plasma proteins. EMBO Mol. Med. 7, 1166–1178 (2015).
Surinova, S. et al. Non-invasive prognostic protein biomarker signatures associated with colorectal cancer. EMBO Mol. Med. 7, 1153–1165 (2015).
Gillet, L.C. et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol. Cell. Proteomics 11, O111.016717 (2012).
Röst, H.L. et al. OpenSWATH enables automated, targeted analysis of data-independent acquisition MS data. Nat. Biotechnol. 32, 219–223 (2014).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Martin, D.B. et al. MRMer, an interactive open source and cross-platform system for data extraction and visualization of multiple reaction monitoring experiments. Mol. Cell. Proteomics 7, 2270–2278 (2008).
Mead, J.A. et al. MRMaid, the web-based tool for designing multiple reaction monitoring (MRM) transitions. Mol. Cell. Proteomics 8, 696–705 (2009).
Prakash, A. et al. Expediting the development of targeted SRM assays: using data from shotgun proteomics to automate method development. J. Proteome Res. 8, 2733–2739 (2009).
Walsh, G.M. et al. Implementation of a data repository-driven approach for targeted proteomics experiments by multiple reaction monitoring. J. Proteomics 72, 838–852 (2009).
Sherwood, C.A. et al. MaRiMba: a software application for spectral library-based MRM transition list assembly. J. Proteome Res. 8, 4396–4405 (2009).
Bertsch, A. et al. Optimal de novo design of MRM experiments for rapid assay development in targeted proteomics. J. Proteome Res. 9, 2696–2704 (2010).
Reiter, L. et al. mProphet: automated data processing and statistical validation for large-scale SRM experiments. Nat. Methods 8, 430–435 (2011).
Teleman, J. et al. Automated selected reaction monitoring software for accurate label-free protein quantification. J. Proteome Res. 11, 3766–3773 (2012).
Prakash, A. et al. Signal maps for mass spectrometry-based comparative proteomics. Mol. Cell. Proteomics 5, 423–432 (2006).
Mueller, L.N. et al. SuperHirn—a novel tool for high resolution LC-MS-based peptide/protein profiling. Proteomics 7, 3470–3480 (2007).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Reiter, L. et al. Protein identification false discovery rates for very large proteomics data sets generated by tandem mass spectrometry. Mol. Cell. Proteomics 8, 2405–2417 (2009).
Doherty, M.K., Hammond, D.E., Clague, M.J., Gaskell, S.J. & Beynon, R.J. Turnover of the human proteome: determination of protein intracellular stability by dynamic SILAC. J. Proteome Res. 8, 104–112 (2009).
Schwanhäusser, B. et al. Global quantification of mammalian gene expression control. Nature 473, 337–342 (2011).
Reinstein, E. & Ciechanover, A. Narrative review: protein degradation and human diseases: the ubiquitin connection. Ann. Intern. Med. 145, 676–684 (2006).
Pratt, J.M. et al. Dynamics of protein turnover, a missing dimension in proteomics. Mol. Cell. Proteomics 1, 579–591 (2002).
Eden, E., Navon, R., Steinfeld, I., Lipson, D. & Yakhini, Z. GOrilla: a tool for discovery and visualization of enriched GO terms in ranked gene lists. BMC Bioinformatics 10, 48 (2009).
Teleman, J. et al. DIANA—algorithmic improvements for analysis of data-independent acquisition MS data. Bioinformatics 31, 555–562 (2015).
Prince, J.T. & Marcotte, E.M. Chromatographic alignment of ESI-LC-MS proteomics data sets by ordered bijective interpolated warping. Anal. Chem. 78, 6140–6152 (2006).
Kohlbacher, O. et al. TOPP–the OpenMS proteomics pipeline. Bioinformatics 23, e191–e197 (2007).
Sturm, M. et al. OpenMS—an open-source software framework for mass spectrometry. BMC Bioinformatics 9, 163 (2008).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Liu, Y. et al. Quantitative variability of 342 plasma proteins in a human twin population. Mol. Syst. Biol. 11, 786 (2015).
Tsou, C.-C. et al. DIA-Umpire: comprehensive computational framework for data-independent acquisition proteomics. Nat. Methods 12, 258–264 (2015).
Adamo, A. et al. 7q11.23 dosage-dependent dysregulation in human pluripotent stem cells affects transcriptional programs in disease-relevant lineages. Nat. Genet. 47, 132–141 (2015).
Kim, S.C. et al. A clean, more efficient method for in-solution digestion of protein mixtures without detergent or urea. J. Proteome Res. 5, 3446–3452 (2006).
Escher, C. et al. Using iRT, a normalized retention time for more targeted measurement of peptides. Proteomics 12, 1111–1121 (2012).
Liu, Y. et al. Glycoproteomic analysis of prostate cancer tissues by SWATH mass spectrometry discovers N-acylethanolamine acid amidase and protein tyrosine kinase 7 as signatures for tumor aggressiveness. Mol. Cell. Proteomics 13, 1753–1768 (2014).
Collins, B.C. et al. Quantifying protein interaction dynamics by SWATH mass spectrometry: application to the 14-3-3 system. Nat. Methods 10, 1246–1253 (2013).
Kessner, D., Chambers, M., Burke, R., Agus, D. & Mallick, P. ProteoWizard: open source software for rapid proteomics tools development. Bioinformatics 24, 2534–2536 (2008).
Craig, R. & Beavis, R.C. TANDEM: matching proteins with tandem mass spectra. Bioinformatics 20, 1466–1467 (2004).
Geer, L.Y. et al. Open mass spectrometry search algorithm. J. Proteome Res. 3, 958–964 (2004).
Kunszt, P. et al. iPortal: the swiss grid proteomics portal: requirements and new features based on experience and usability considerations. Concurr. Comput. 27, 433–445 (2015).
Keller, A., Eng, J., Zhang, N., Li, X.J. & Aebersold, R. A uniform proteomics MS/MS analysis platform utilizing open XML file formats. Mol. Syst. Biol. 1, 2005.0017 (2005).
Keller, A., Nesvizhskii, A.I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Shteynberg, D. et al. iProphet: multi-level integrative analysis of shotgun proteomic data improves peptide and protein identification rates and error estimates. Mol. Cell. Proteomics 10, M111.007690 (2011).
Schubert, O.T. et al. Building high-quality assay libraries for targeted analysis of SWATH MS data. Nat. Protoc. 10, 426–441 (2015).
Lam, H. et al. Development and validation of a spectral library searching method for peptide identification from MS/MS. Proteomics 7, 655–667 (2007).
Röst, H.L., Aebersold, R. & Schubert, O. Automated SWATH data analysis using targeted extraction of ion chromatograms. Preprint at. bioRxiv http://dx.doi.org/10.1101/044552 (2016).
Acknowledgements
We would like to thank the SyBIT project of SystemsX.ch for support and maintenance of the lab-internal computing infrastructure. L. Blum (SyBIT) contributed substantially to making this software available to the lab and provided critical feedback. This work was supported by ETH Zurich (grant ETH-30 11-2 to H.R. and R.A.), the Swiss National Science Foundation (SNSF grants P2EZP3_162268 to H.R. and 31003A_166435 to R.A.), ERC Proteomics v3.0 (AdvG grant 233226 to R.A.), ERC Proteomics4D (AdvG grant 670821 to R.A.), the PhosphonetX project of SystemsX.ch, the ERC DISEASEAVATARS (grant 616441 to R.A. and G.T.), the Telethon Foundation (GGP14265 to G.T.), the Regione Lombardia (Grant Ricerca Indipendente 2012 to G.T.) and the Fondazione Umberto Veronesi (G.D.).
Author information
Authors and Affiliations
Contributions
H.L.R. designed and wrote code, performed data analysis and produced the figures. Y.L., G.D. and M.Z. performed iPSC experiments and acquired MS data. P.N. and G.R. contributed to code and provided an initial prototype of the implementation. B.C.C. and L.G. acquired MS data and gave critical input. G.T., L.M. and R.A. designed and supervised the study. All authors contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Notes 1–7 (PDF 2736 kb)
Supplementary Software
msproteomicstools 0.4.3 (ZIP 999 kb)
Supplementary Table 1
Result table describing the manually picked peptides with retention times and peak boundaries. (CSV 5750 kb)
Supplementary Table 2
Significant proteins from the S. pyogenes analysis without any alignment. (CSV 137 kb)
Supplementary Table 3
Significant proteins from the S. pyogenes analysis with TRIC alignment. (CSV 162 kb)
Supplementary Table 4
Degradation rates for the iPSC as determined by SWATH-MS analysis on the peptide level. (CSV 556 kb)
Supplementary Table 5
Degradation rates for the iPSCs as determined by SWATH-MS analysis on the protein level. (CSV 87 kb)
Supplementary Table 6
GO-enrichment analysis using Gorilla. (XLS 32 kb)
Rights and permissions
About this article
Cite this article
Röst, H., Liu, Y., D'Agostino, G. et al. TRIC: an automated alignment strategy for reproducible protein quantification in targeted proteomics. Nat Methods 13, 777–783 (2016). https://doi.org/10.1038/nmeth.3954
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3954
This article is cited by
-
SeFilter-DIA: Squeeze-and-Excitation Network for Filtering High-Confidence Peptides of Data-Independent Acquisition Proteomics
Interdisciplinary Sciences: Computational Life Sciences (2024)
-
Achieving quantitative reproducibility in label-free multisite DIA experiments through multirun alignment
Communications Biology (2023)
-
Introducing untargeted data-independent acquisition for metaproteomics of complex microbial samples
ISME Communications (2022)
-
Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity
Nature Communications (2022)
-
Implementing the reuse of public DIA proteomics datasets: from the PRIDE database to Expression Atlas
Scientific Data (2022)