Abstract
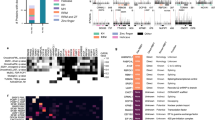
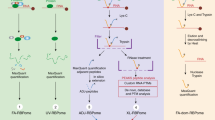
The complexity of transcriptome-wide protein–RNA interaction networks is incompletely understood. While emerging studies are greatly expanding the known universe of RNA-binding proteins, methods for the discovery and characterization of protein–RNA interactions remain resource intensive and technically challenging. Here we introduce a UV-C crosslinking and immunoprecipitation platform, irCLIP, which provides an ultraefficient, fast, and nonisotopic method for the detection of protein–RNA interactions using far less material than standard protocols.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Brimacombe, R., Stiege, W., Kyriatsoulis, A. & Maly, P. Methods Enzymol. 164, 287–309 (1988).
Ule, J., Jensen, K., Mele, A. & Darnell, R.B. Methods 37, 376–386 (2005).
Licatalosi, D.D. et al. Nature 456, 464–469 (2008).
Chi, S.W., Zang, J.B., Mele, A. & Darnell, R.B. Nature 460, 479–486 (2009).
Xue, Y. et al. Mol. Cell 36, 996–1006 (2009).
Hafner, M. et al. Cell 141, 129–141 (2010).
Lebedeva, S. et al. Mol. Cell 43, 340–352 (2011).
König, J. et al. Nat. Struct. Mol. Biol. 17, 909–915 (2010).
Zarnack, K. et al. Cell 152, 453–466 (2013).
Weyn-Vanhentenryck, S.M. et al. Cell Rep. 6, 1139–1152 (2014).
Flynn, R.A. et al. RNA 21, 135–143 (2015).
Castello, A. et al. Cell 149, 1393–1406 (2012).
Baltz, A.G. et al. Mol. Cell 46, 674–690 (2012).
Kwon, S.C. et al. Nat. Struct. Mol. Biol. 20, 1122–1130 (2013).
Moore, M.J. et al. Nat. Protoc. 9, 263–293 (2014).
Huppertz, I. et al. Methods 65, 274–287 (2014).
Kulak, N.A., Pichler, G., Paron, I., Nagaraj, N. & Mann, M. Nat. Methods 11, 319–324 (2014).
Urlaub, H., Hartmuth, K. & Lührmann, R. Methods 26, 170–181 (2002).
Choe, W.S. & Middelberg, A.P. Biotechnol. Prog. 17, 1107–1113 (2001).
Mohr, S. et al. RNA 19, 958–970 (2013).
Acknowledgements
We thank A. Fire, P. Sarnow, and M. Kay for presubmission review. We thank L. Morcom and P. Bernstein for expert administrative assistance and members of the Khavari lab for helpful discussions. This work was supported by the US VA Office of Research and Development, NIH AR49737 and NIH CA142635 (P.A.K.), P50-HG007735 and R01-ES023168 (H.Y.C.), and the Stanford Medical Scientist Program and NIH 1F30CA189514-01 (R.A.F.).
Author information
Authors and Affiliations
Contributions
B.J.Z. designed and executed experiments, analyzed the data, and wrote the manuscript with input from the coauthor R.A.F. R.A.F. and B.T.D. designed the FAST-iCLIP pipeline for analysis of ir/iCLIP data. Y.S. executed statistical analysis of ir/iCLIP data replicates. P.A.K. and H.Y.C. designed experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 irCLIP adapter construction, quantitation and validation
(a) Construction schematic for generation of the pre-adenylated, IR800CW dye conjugated irCLIP DNA adapter. (b) Odyssey CLx infrared scan and (c) quantitation of serial diluted irCLIP adapter showing 5 orders of magnitude sensitivity on nitrocellulose. (d) Autoradiogram and Odyssey CLx infrared scan demonstrating efficient ligation of γ-32P-labeled hnRNP-C-RNA complexes with the irCLIP adapter, evidenced by supershifting of unligated complexes (black vs green box, left panel) and RNase A (in-lysate digestion, 50 ng/ml) dependence for generation of irCLIP adapter signal (right panel). (e) Autoradiograms (left and middle panels) and Odyssey CLx infrared scan (right panel) of a 3-fold serial dilution of γ-32P -labeled, irCLIP-adapter ligated hnRNP-C-RNA complexes.
Supplementary Figure 2 On-bead S1 nuclease digestion improves RNA digestion profile for multiple classes of RBPs
Equivalent amounts of UV-C treated HeLa cell extracts were pre-digested with RNase A at 50, 16.7, or 5.6 ng/mL for 10 min at 37°C or digested post-immunoprecipitation with S1 nuclease at 9 or 3 U/μL for 15 min at 30°C, ligated overnight with the irCLIP-adapter, resolved by SDS-PAGE and transferred to nitrocellulose. (a-d) Odyssey CLx infrared scans of the nitrocellulose bound immunoprecipitates using antibodies against the RRM domain proteins (a) hnRNP-C, 37 kDa (b) HuR, 39 kDa, the RNA-helicase-domain protein DDX5, 73 kDa (c), and the double-stranded-RNA-binding domain protein ILF3, 90 and 110 kDa (d). (e-h) Quantitation of optimal ligations, 40-100 nucleotide RNA fragments (~30-60 kDa window above RBP size) for hnRNP-C, HuR, DDX5, and ILF3, respectively.
Supplementary Figure 3 Comparison of on-bead nuclease digestions for multiple classes of RBPs
Batch immunoprecipitations were equally divided immediately prior to on-bead nuclease digestions. All reactions (see Methods) were incubated for 15 min at 30°C and each nuclease tested at 3 doses along a 4-fold serial dilution (RNase 1 0.4-0.1-0.025 U/μL, RNase A 20-5-1.25 ng/mL, S1 12-4-1 U/μL). Overnight irCLIP-adapter ligated RNP-complexes were resolved by SDS-PAGE. Odyssey CLx infrared scans of the nitrocellulose bound immunoprecipitates for (a) hnRNPC, 37 kDa, (b) HuR 39 kDa, (c) DDX5, 73 kDa and (d) ILF3 90 and 110 kDa. Quantitation of target ligations, 40-100 nucleotide RNA fragments (~30-60 kDa window above RBP size), (e-h) for hnRNP-C, HuR, DDX5, and ILF3, respectively.
Supplementary Figure 4 Optimized recovery of ligated RNA-peptide fragments
irCLIP-adapter ligated hnRNP-C-RNA complexes were diluted in 1xLDS sample buffer and equally divided between 36 lanes for resolution by SDS-PAGE. Identical sections, 65-100 kDa, of nitrocellulose were excised for comparison of proteinase K digestion parameters. (a) Infrared Odyssey CLx scan of representative nitrocellose dot blots and (b) quantitation of precipitated, irCLIP-adapter ligated RNA from proteinase K digestions with the indicated additives at the indicated temperatures. (c) Free irCLIP-adapter and representative RNA samples from (a) were resolved by denaturing PAGE and imaged on an infrared Odyssey CLx.
Supplementary Figure 5 irCLIP enhancement of cDNA synthesis and cDNA purification
irCLIP-ligated RNA was purified from hnRNP-C-RNA complexes. 1 femtomole was used as starting material for quadruplicate cDNA synthesis reactions using Super Script III (Thermo Fisher Scientific) according to the manufacturer’s protocol or TGIRT-III (InGex) reverse transcriptase (Methods). cDNA was purified and amplified according to irCLIP library construction (Methods). (a) Histogram of relative amplification of SuperScript III vs TGIRT-III PCR1 (Methods) reactions. The average Ct values of SuperScript III were used as control. 1 femtomole of IR800CW-cDNA primer was purified in triplicate with the Qiagen Nucleotide Removal Kit, the Zymo Oligo Clean and Concentrator Kit per manufacturer’s instructions or with 2 volumes of AMPure beads and 5 volumes of isopropanol for 15 min. (b) Histogram of quantitated nitrocellulose dot blots of purified IR800CW-cDNA primer. Percent capture is the average of triplicate purifications.
Supplementary Figure 6 Controls for irCLIP experiments
(a) HeLa cells were untreated or subjected to the indicated doses of UVC. RNA was isolated and resolved by denaturing agarose gel electrophoresis showing high quality intact RNA for the highest dose of UV-C, 0.3J/cm2. Odyssey CLx infrared immunoblot analyses of UV-crosslinked HeLa cell extracts showing antibody specificity and IP efficiency for (b) hnRNPC, (c) HuR and (d) PTBP1. Input lanes (b-d) were loaded at 100% of total IP. Specific bands were quantitated using LiCor Image Studio software. For IPs, 1 μg of antibody was incubated with 10μg of lysate for hnRNPC or 30μg of lysate for HuR and PTBP1. For buffer and wash conditions see Methods.
Supplementary Figure 7 irCLIP produces high quality data sets from greatly reduced cellular inputs
Pearson correlations of RT stops between biological replicates of 100,000 cell irCLIP experiments for (a) PTBP1 and (b) HuR. (c) Histogram of the unique read fractions and total clusters of the deeply sequenced biological replicates of irCLIP of PTBP1 and HuR. (d) Pearson correlations of the RT stops within the significant clusters (>100,000 per RBP) identified in the deeply sequenced irCLIP data sets for hnRNP-C, PTBP1, and HuR. Low R2 values indicated highly unique clusters for each RBP.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, and Supplementary Tables 1 and 2. (PDF 1690 kb)
Rights and permissions
About this article
Cite this article
Zarnegar, B., Flynn, R., Shen, Y. et al. irCLIP platform for efficient characterization of protein–RNA interactions. Nat Methods 13, 489–492 (2016). https://doi.org/10.1038/nmeth.3840
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3840
This article is cited by
-
SRSF2 is required for mRNA splicing during spermatogenesis
BMC Biology (2023)
-
Temporal-iCLIP captures co-transcriptional RNA-protein interactions
Nature Communications (2023)
-
The RNA binding proteins TIA1 and TIAL1 promote Mcl1 mRNA translation to protect germinal center responses from apoptosis
Cellular & Molecular Immunology (2023)
-
An mRNA processing pathway suppresses metastasis by governing translational control from the nucleus
Nature Cell Biology (2023)
-
Regulatory non-coding RNAs: everything is possible, but what is important?
Nature Methods (2022)