Abstract
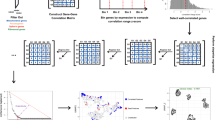
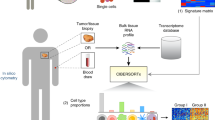
We introduce CIBERSORT, a method for characterizing cell composition of complex tissues from their gene expression profiles. When applied to enumeration of hematopoietic subsets in RNA mixtures from fresh, frozen and fixed tissues, including solid tumors, CIBERSORT outperformed other methods with respect to noise, unknown mixture content and closely related cell types. CIBERSORT should enable large-scale analysis of RNA mixtures for cellular biomarkers and therapeutic targets (http://cibersort.stanford.edu/).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Hanahan, D. & Weinberg, R.A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Coussens, L.M., Zitvogel, L. & Palucka, A.K. Neutralizing tumor-promoting chronic inflammation: a magic bullet? Science 339, 286–291 (2013).
Shen-Orr, S.S. & Gaujoux, R. Computational deconvolution: extracting cell type-specific information from heterogeneous samples.. Curr. Opin. Immunol. 25, 571–578 (2013).
Abbas, A.R., Wolslegel, K., Seshasayee, D., Modrusan, Z. & Clark, H.F. Deconvolution of blood microarray data identifies cellular activation patterns in systemic lupus erythematosus. PLoS ONE 4, e6098 (2009).
Gong, T. et al. Optimal deconvolution of transcriptional profiling data using quadratic programming with application to complex clinical blood samples. PLoS ONE 6, e27156 (2011).
Qiao, W. et al. PERT: a method for expression deconvolution of human blood samples from varied microenvironmental and developmental conditions. PLoS Comput. Biol. 8, e1002838 (2012).
Liebner, D.A., Huang, K. & Parvin, J.D. MMAD: microarray microdissection with analysis of differences is a computational tool for deconvoluting cell type-specific contributions from tissue samples. Bioinformatics 30, 682–689 (2014).
Zhong, Y., Wan, Y.-W., Pang, K., Chow, L. & Liu, Z. Digital sorting of complex tissues for cell type-specific gene expression profiles. BMC Bioinformatics 14, 89 (2013).
Zuckerman, N.S., Noam, Y., Goldsmith, A.J. & Lee, P.P. A self-directed method for cell-type identification and separation of gene expression microarrays. PLoS Comput. Biol. 9, e1003189 (2013).
Schölkopf, B., Smola, A.J., Williamson, R.C. & Bartlett, P.L. New support vector algorithms. Neural Comput. 12, 1207–1245 (2000).
Lukk, M. et al. A global map of human gene expression.. Nat. Biotechnol. 28, 322–324 (2010).
Shen-Orr, S.S. et al. Cell type–specific gene expression differences in complex tissues. Nat. Methods 7, 287–289 (2010).
Kuhn, A., Thu, D., Waldvogel, H.J., Faull, R.L.M. & Luthi-Carter, R. Population-specific expression analysis (PSEA) reveals molecular changes in diseased brain. Nat. Methods 8, 945–947 (2011).
Yoshihara, K. et al. Inferring tumour purity and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Farrar, D.E. & Glauber, R.R. Multicollinearity in regression analysis: the problem revisited. Rev. Econ. Stat. 49, 92–107 (1967).
Burington, B. et al. CD40 pathway activation status predicts response to CD40 therapy in diffuse large B cell lymphoma. Sci. Transl. Med. 3, 74ra22 (2011).
Gong, T. & Szustakowski, J.D. DeconRNASeq: a statistical framework for deconvolution of heterogeneous tissue samples based on mRNA-Seq data.. Bioinformatics 29, 1083–1085 (2013).
Levy, R. et al. Active idiotypic vaccination versus control immunotherapy for follicular lymphoma. J. Clin. Oncol. 32, 1797–1803 (2014).
Lu, P., Nakorchevskiy, A. & Marcotte, E.M. Expression deconvolution: a reinterpretation of DNA microarray data reveals dynamic changes in cell populations. Proc. Natl. Acad. Sci. USA 100, 10370–10375 (2003).
Zhong, Y. & Liu, Z. Gene expression deconvolution in linear space. Nat. Methods 9, 8–9 (2012).
Guyon, I. & Elisseeff, A. An introduction to variable and feature selection. J. Mach. Learn. Res. 3, 1157–1182 (2003).
Cherkassky, V. & Ma, Y. Practical selection of SVM parameters and noise estimation for SVM regression. Neural Netw. 17, 113–126 (2004).
Hoerl, A.E. & Kennard, R.W. Ridge regression: biased estimation for nonorthogonal problems. Technometrics 12, 55–67 (1970).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning 2nd edn. (Springer, 2009).
Wang, L., Zhu, J. & Zou, H. The doubly regularized support vector machine. Statist. Sinica 16, 589–615 (2006).
Drucker, H., Burges, C.J.C., Kaufman, L., Smola, A. & Vapnik, V. in Adv. Neural Inf. Process. Syst. (eds. Mozer, M.C., Jordan, M.I. & Petsche, T.) 9, 155–161 (MIT Press, 1997).
Su, L.J. et al. Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC Genomics 8, 140 (2007).
Landi, M.T. et al. Gene expression signature of cigarette smoking and its role in lung adenocarcinoma development and survival. PLoS One 3, e1651 (2008).
Storey, J.D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 100, 9440–9445 (2003).
Benita, Y. et al. Gene enrichment profiles reveal T-cell development, differentiation, and lineage-specific transcription factors including ZBTB25 as a novel NF-AT repressor. Blood 115, 5376–5384 (2010).
Watkins, N.A. et al. A HaemAtlas: characterizing gene expression in differentiated human blood cells. Blood 113, e1–e9 (2009).
Abbas, A.R. et al. Immune response in silico (IRIS): immune-specific genes identified from a compendium of microarray expression data. Genes Immun. 6, 319–331 (2005).
Acknowledgements
We are grateful to H. Maecker, M. Davis, R. Levy and the Stanford Human Immune Monitoring Center for assistance with this study. This work was supported by grants from the Doris Duke Charitable Foundation (A.A.A.), the Damon Runyon Cancer Research Foundation (A.A.A.), the B&J Cardan Oncology Research Fund (A.A.A.), the Ludwig Institute for Cancer Research (A.A.A. and M.D.), US National Institutes of Health (NIH) grant U01 CA154969 (A.J.G., W.F., Y.X., C.D.H. and M.D.), NIH grant U19 AI090019, NIH grant PHS NRSA 5T32 CA09302-35 (A.M.N.), US Department of Defense grant W81XWH-12-1-0498 (A.M.N.) and a grant from the Siebel Stem Cell Institute and the Thomas and Stacey Siebel Foundation (A.M.N.).
Author information
Authors and Affiliations
Contributions
A.M.N. and A.A.A. conceived of CIBERSORT, developed strategies for related experiments, analyzed the data and wrote the paper. A.M.N. developed and implemented CIBERSORT. C.L.L. implemented web infrastructure and wrote the paper. M.R.G. performed flow cytometry and gene expression profiling of leukocytes from human tonsils and peripheral blood. A.J.G. assisted in the conceptual development of CIBERSORT. W.F., Y.X., C.D.H. and M.D. assisted in the collection and analysis of lung tissue. All authors discussed the results and implications and commented on the manuscript at all stages.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Illustrative example of support vector regression
A simple two-dimensional dataset analyzed with linear ν-SVR, with results shown for two values of ν (note that both panels show the same data points). As detailed in Online Methods, linear SVR identifies a hyperplane (which, in this two-dimensional example, is a line) that fits as many data points as possible (given its objective function) within a constant distance, ɛ (open circles). Data points lying outside of this ‘ɛ-tube’ are termed ‘support vectors’ (red circles), and are penalized according to their distance from the ɛ-tube by linear slack variables (ξi). Importantly, the support vectors alone are sufficient to completely specify the linear function, and provide a sparse solution to the regression that reduces the chance of overfitting. In ν-SVR, the ν parameter determines both the lower bound of support vectors and upper bound of training errors. As such, higher values of ν result in a smaller ɛ-tube and a greater number of support vectors (right panel). For CIBERSORT, the support vectors represent genes selected from the signature matrix for analysis of a given mixture sample, and the orientation of the regression hyperplane determines the estimated cell type proportions in the mixture. For further details, including the key technical advantages of ν-SVR for GEP deconvolution, see Online Methods.
Reference (Main Text)
10. Schölkopf, B., Smola, A.J., Williamson, R.C. & Bartlett, P.L. Neural Comput. 12, 1207-1245 (2000).
Supplementary Figure 2 LM22 signature matrix and comparison to Abbas et al.
(a) Heat map of the LM22 signature matrix (Supplementary Table 1) depicting the relative expression of each gene across 22 leukocyte subsets. Gene expression levels were unit variance normalized, and cell subsets and genes were clustered hierarchically using Euclidean distance (higher expression, red; lower expression, blue). (b) Overlap between LM22 and a previously published signature matrix (GSE22886) with respect to genes, cell subsets, and expression arrays used. For gene overlap between Abbas et al. and LM22, we considered all Affymetrix probesets as ‘genes’, including those not resolvable to HUGO gene symbols (n = 36). For cell subset overlap, we only considered cell types profiled in GSE22886; this dataset contains 22 distinct phenotypes, and six were pooled into three phenotypes for incorporation into LM22, resulting in an overlap of 13 (for details, see Supplementary Table 1). (c) All-versus-all heat map of correlation coefficients (Pearson) comparing the reference profiles of each cell subset in LM22 (genes were normalized as described in Methods; same as Supplementary Table 1).
Reference (Main Text)
4. Abbas, A.R., Wolslegel, K., Seshasayee, D., Modrusan, Z. & Clark, H.F. PLoS One 4, e6098 (2009).
Supplementary Figure 3 Validation of LM22 by analysis of purified leukocyte subsets
(a) Fractions of each LM22 cell subset called by CIBERSORT in validation arrays containing purified/enriched leukocytes profiled in LM22 (related to Fig. 1b; also see Supplementary Table 2). Results for arrays of a given cell subset are summarized as median fractions. Cell subset abbreviations in the color key are defined in Supplementary Table 1. (b) Left: B and T lymphocytes were flow-sorted from five human tonsils to mean purity levels exceeding 95% and 98%, respectively, and then profiled by microarray. Right: The fractional representations of these B/T cells, along with any remaining leukocyte content, as inferred by CIBERSORT.
Supplementary Figure 4 Resolution of well-defined mixtures with CIBERSORT
Analysis of CIBERSORT performance using different signature matrices (top) applied to different mixtures (bottom). Top: Cell population reference expression signatures for (a) purified blood cancer cell line expression profiles in GSE11103, (b) neural gene expression profiles in GSE19380, and (c) LM22 (Supplementary Table 1). Bottom: Comparison of known and inferred fractions for defined mixtures of (a) blood cancer cell lines (GSE11103) and (b) neural cell types (GSE19380). Concordance was assessed by Pearson correlation (R) and linear regression (solid lines). (c) CIBERSORT analysis of pre- and post-Rituximab therapy PBMC samples, including one paired sample, from four Non-Hodgkin’s lymphoma patients using LM22 (pooled into 11 leukocyte types for clarity; see Supplementary Table 1).
References (Main Text)
4. Abbas, A.R., Wolslegel, K., Seshasayee, D., Modrusan, Z. & Clark, H.F. PLoS One 4, e6098 (2009).
13. Kuhn, A., Thu, D., Waldvogel, H.J., Faull, R.L.M. & Luthi-Carter, R. Nat. Methods 8, 945-947 (2011).
Supplementary Figure 5 Comparative analysis of deconvolution methods on simulated tumors with added noise
(a, b) Performance of each method with respect to added tumor content (x-axis) and non-log linear noise (y-axis) (see Online Methods for details). Each parameter was analyzed in evenly spaced intervals for a total of 900 mixtures (30 x 30). (a) Concordance between known and estimated cell type proportions as determined by Pearson correlation coefficient (R), with a floor of zero. (b) Performance evaluated as a function of the deviation of each mixture from its original, unmodified values (represented on the x-axis as 1 – R). Differences between known and predicted cell type proportions (represented as percentages) in b were assessed by root mean squared error (RMSE), with a ceiling of 40.
Supplementary Figure 6 Comparison of deconvolution methods with respect to detection limit in simulated mixtures with unknown content
Each color represents a defined input concentration for a given cell type (here, Jurkat), and each line represents its concentration as predicted by GEP deconvolution. Known Jurkat concentrations were measured across a range of added tumor content in five simulated mixtures of four blood cell lines, with different concentrations of a colon cancer line (see Online Methods). Data are presented as medians (n = 5 mixtures) ± 95% confidence intervals.
Supplementary Figure 7 Analysis of detection limit for each cell subset in LM22
(a) Same as in Supplementary Fig. 6, except here detection limit was assessed using defined inputs of naïve B-cells added to simulated mixtures of the remaining 21 cell types from LM22 (Supplementary Table 1). The impact of unknown content on detection limit was evaluated by adding simulated GEPs created by randomly permuting naïve B-cell genes. Data are presented as medians (n = 4 mixtures) ± 95% confidence intervals. (b) Same as in a, but for all cell types in LM22. To prevent higher magnitude spike-ins from driving the correlation, we summarized performance using the non-parametric Spearman rank correlation, and compared known and predicted fractions over all spike-ins and levels of unknown content tested. Considering these results in aggregate, CIBERSORT significantly outperformed other methods tested (P < 0.0001; paired two-sided Wilcoxon signed rank test; n = 22 cell subsets). Of note, CIBERSORT also outperformed other methods in relation to linear fit, as measured by Pearson correlation. For further details, see Online Methods.
Supplementary Figure 8 Impact of marker genes on deconvolution
(a) Heat map of signature matrix 1 (SM1), which contains only cell type specific marker genes. (b) Heat map of signature matrix 2 (SM2), which contains only non-cell type specific marker genes. (c) CIBERSORT and DSA deconvolution performance on ten mixtures created using SM1. (d,e) Deconvolution performance on ten mixtures created using SM2: (d) CIBERSORT and RLR, (e) QP, LLSR, and PERT. For details, see Online Methods. Statistical concordance between known and observed cell type proportions was determined by linear regression (dashed lines) and Pearson correlation (R).
Supplementary Figure 9 Analysis of feature (gene) selection in defined mixtures
(a) Results from applying CIBERSORT to a spike series, in which the LM22 reference profile for CD8 T cells was spiked into the corresponding reference profile for resting mast cells (MCs–) in even increments (n = 21). (Of note, both cell types have highly distinct expression vectors in LM22; see Supplementary Fig. 2c.) (b) Comparison between genes selected by support vector regression (SVR) to deconvolve 100% resting mast cells, but not CD8 T cells, and vice versa. For each unique subset of genes, expression levels in the LM22 signature matrix are further compared between resting mast cells and CD8 T cells. A paired two-sided Wilcoxon signed rank test was used for within group comparison and an unpaired two-sided Wilcoxon rank sum test was used for between group comparisons. Data are presented as medians ± interquartile range. While genes uniquely selected for the 100% CD8 T cell sample are significantly more expressed in CD8 T cells than resting mast cells, the magnitude is small. Moreover, the opposite scenario is not observed for resting mast cell genes in the 100% resting mast cell sample, suggesting SVR gene selection is not strongly correlated with the presence or absence of a particular cell subset in the mixture. (c) Comparison between gene expression levels in LM22 (n = 547 genes) and the frequency that each gene was selected, if at all, by SVR from the set of 19 mixtures with >0% CD8 T cells and >0% resting mast cells (see panel a). Top: Comparison with expression levels of (left) CD8 T cells or (right) resting mast cells. Bottom: Comparison with mean expression levels of (left) CD8 T cells and resting mast cells or (right) all cell subsets in LM22. Regardless of spike-in composition, the highest correlation between expression and gene selection frequency was observed when considering all cell types in LM22. Statistical concordance in c was determined by linear regression (red lines).
Supplementary Figure 10 Impact of multicollinearity on signature matrix–based methods
(a–d) The effect of multicollinearity on deconvolution performance is shown for mixtures with unknown content (a-c) or noise added to the signature matrix (d). Each panel is organized as follows: Top: The mean cross-correlation coefficient (left y-axis) and corresponding mean condition number kappa (right y-axis) of signature matrix GEPs over a broad range of multicollinearity values (x-axis; Online Methods). ‘Mean cross correlation’ indicates the average value of an all-versus-all correlation comparison (Pearson) of signature matrix reference profiles, whereas ‘kappa’ is a measure of signature matrix stability (Online Methods). Both metrics capture multicollinearity (or the degree of similarity among reference profiles) in the signature matrix. Bottom left: The relative performance of four deconvolution methods on simulated mixtures, comparing known and predicted cellular fractions (y-axis). Results from 20 levels of multicollinearity are shown ordered by increasing multicollinearity (left to right). Each level of multicollinearity was simulated ten times, and summarized values are presented as means ± s.e.m. Bottom right: Summarization of the performance of each method as a box plot, with interquartile range contained in the box and minimum and maximum points denoted by whiskers. Group comparisons between CIBERSORT and other methods were performed using a paired two-sided Wilcoxon signed rank test. All signature matrices and mixture vectors were unit variance normalized prior to analysis. For additional details, see Online Methods.
Supplementary Figure 11 Comparison of leukocyte deconvolution results between frozen and FFPE samples in 18 individual DLBCL tumors
(a) Results are shown for the 22 leukocyte subsets resolved in each tumor, related to Fig. 2g. Data points (circles) are colored as in Fig. 2g and indicate cell type. Deconvolution results for sample IDs 11 and 14 were not significantly correlated between FFPE and frozen conditions (NS). (b) Scatter plots for representative cell types across all 18 tumors. Expression data were obtained from GSE18377. Linear regression (solid lines) was used to compare all paired data in a, b. R, Pearson correlation.
Reference (Main Text)
16. Burington, B. et al. Sci. Transl. Med. 3, 74ra22 (2011).
Supplementary Figure 12 Comparison of deconvolution methods for enumeration of nine leukocyte subsets in PBMCs
(a) Scatter plots comparing flow cytometry with five deconvolution methods for the enumeration of eight leukocyte subsets in 20 PBMC samples. (b) Same as in a but for Tregs, which were exclusively profiled in a separate cohort of seven PBMC samples. Of ten total phenotypic subsets analyzed in PBMCs (Online Methods), the nine subsets shown here were deconvolved by at least one method with a correlation coefficient of at least 0.5 (only gamma delta T cells are not shown). Detailed performance metrics for all ten subsets are provided in Supplementary Table 4. Linear regression (solid lines) was used to statistically compare all paired data in a, b. R, Pearson correlation.
Supplementary Figure 13 Comparison of deconvolution methods for enumeration of three leukocyte subsets in FL tumor biopsies
Scatter plots comparing flow cytometry with five deconvolution methods for the enumeration of three leukocyte subsets, including malignant B cells, in disaggregated FL lymph node biopsies. For RMSE values of individual cell subsets, see Supplementary Table 4. Concordance between flow cytometry and GEP deconvolution was assessed by linear regression (solid lines). R, Pearson correlation.
Supplementary Figure 14 Summary of benchmarking results for complex mixtures
Using two measures of performance (R and RMSE), CIBERSORT significantly outperformed other gene expression-based methods (paired two-sided Wilcoxon signed rank test), and generally performed better than all other methods on complex mixtures (Fig. 2d). Raw data are provided in ‘Complex Mixtures’ in Supplementary Table 4. For details of deconvolution methods, see Online Methods (also see Supplementary Table 3 and Supplementary Discussion).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14, Supplementary Note, Supplementary Results and Supplementary Discussion (PDF 13078 kb)
Supplementary Table 1
Leukocyte signature matrix (LM22). Details of LM22, including gene expression matrix and source data. (XLS 424 kb)
Supplementary Table 2
Validation of LM22 on external datasets of purified leukocyte subsets. Analysis of external GEP datasets consisting of distinct leukocyte subsets. (XLS 39 kb)
Supplementary Table 3
Feature comparison of GEP deconvolution methods analyzed in this work. Table comparing key features of GEP deconvolution approaches. (XLS 31 kb)
Supplementary Table 4
Comparative analysis of GEP deconvolution methods. Performance comparison of CIBERSORT, RLR, PERT, LLSR, and QP on both complex and idealized mixtures. (XLS 64 kb)
Rights and permissions
About this article
Cite this article
Newman, A., Liu, C., Green, M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat Methods 12, 453–457 (2015). https://doi.org/10.1038/nmeth.3337
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3337
This article is cited by
-
Targeted deletion of CD244 on monocytes promotes differentiation into anti-tumorigenic macrophages and potentiates PD-L1 blockade in melanoma
Molecular Cancer (2024)
-
Clustering of HR + /HER2− breast cancer in an Asian cohort is driven by immune phenotypes
Breast Cancer Research (2024)
-
Quantifying the proportion of different cell types in the human cortex using DNA methylation profiles
BMC Biology (2024)
-
The innovative checkpoint inhibitors of lung adenocarcinoma, cg09897064 methylation and ZBP1 expression reduction, have implications for macrophage polarization and tumor growth in lung cancer
Journal of Translational Medicine (2024)
-
Single-cell and transcriptomic analyses reveal the influence of diabetes on ovarian cancer
BMC Genomics (2024)