Abstract
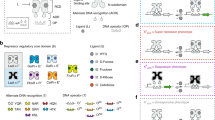
Bacterial regulators of transcriptional elongation are versatile units for building custom genetic switches, as they control the expression of both coding and noncoding RNAs, act on multigene operons and can be predictably tethered into higher-order regulatory functions (a property called composability). Yet the less versatile bacterial regulators of translational initiation are substantially easier to engineer. To bypass this tradeoff, we have developed an adaptor that converts regulators of translational initiation into regulators of transcriptional elongation in Escherichia coli. We applied this adaptor to the construction of several transcriptional attenuators and activators, including a small molecule–triggered attenuator and a group of five mutually orthogonal riboregulators that we assembled into NOR gates of two, three or four RNA inputs. Continued application of our adaptor should produce large collections of transcriptional regulators whose inherent composability can facilitate the predictable engineering of complex synthetic circuits.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Isaacs, F.J., Dwyer, D.J. & Collins, J.J. RNA synthetic biology. Nat. Biotechnol. 24, 545–554 (2006).
Lucks, J.B., Qi, L., Mutalik, V.K., Wang, D. & Arkin, A.P. Versatile RNA-sensing transcriptional regulators for engineering genetic networks. Proc. Natl. Acad. Sci. USA 108, 8617–8622 (2011).
Liu, C.C., Qi, L., Yanofsky, C. & Arkin, A.P. Regulation of transcription by unnatural amino acids. Nat. Biotechnol. 29, 164–168 (2011).
Werstuck, G. & Green, M.R. Controlling gene expression in living cells through small molecule-RNA interactions. Science 282, 296–298 (1998).
Bayer, T.S. & Smolke, C.D. Programmable ligand-controlled riboregulators of eukaryotic gene expression. Nat. Biotechnol. 23, 337–343 (2005).
Carothers, J.M., Goler, J.A., Juminaga, D. & Keasling, J.D. Model-driven engineering of RNA devices to quantitatively program gene expression. Science 334, 1716–1719 (2011).
Topp, S. & Gallivan, J.P. Emerging applications of riboswitches in chemical biology. ACS Chem. Biol. 5, 139–148 (2010).
Lucks, J.B., Qi, L., Whitaker, W.R. & Arkin, A.P. Toward scalable parts families for predictable design of biological circuits. Curr. Opin. Microbiol. 11, 567–573 (2008).
Roth, A. & Breaker, R.R. The structural and functional diversity of metabolite-binding riboswitches. Annu. Rev. Biochem. 78, 305–334 (2009).
Link, K.H. & Breaker, R.R. Engineering ligand-responsive gene-control elements: lessons learned from natural riboswitches. Gene Ther. 16, 1189–1201 (2009).
Romby, P. & Springer, M. Bacterial translational control at atomic resolution. Trends Genet. 19, 155–161 (2003).
Narberhaus, F., Waldminghaus, T. & Chowdhury, S. RNA thermometers. FEMS Microbiol. Rev. 30, 3–16 (2006).
Grundy, F.J. & Henkin, T.M. From ribosome to riboswitch: control of gene expression in bacteria by RNA structural rearrangements. Crit. Rev. Biochem. Mol. Biol. 41, 329–338 (2006).
Reuter, J.S. & Mathews, D.H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics 11, 129 (2010).
Salis, H.M., Mirsky, E.A. & Voigt, C.A. Automated design of synthetic ribosome binding sites to control protein expression. Nat. Biotechnol. 27, 946–950 (2009).
Mutalik, V.K., Qi, L., Guimaraes, J., Lucks, J.B. & Arkin, A.P. Rationally designed families of orthogonal RNA regulators of translation. Nat. Chem. Biol. 8, 447–454 (2012).
Naville, M. & Gautheret, D. Transcription attenuation in bacteria: theme and variations. Brief. Funct. Genomic. Proteomic. 8, 482–492 (2009).
Sudarsan, N. et al. Tandem riboswitch architectures exhibit complex gene control functions. Science 314, 300–304 (2006).
Gutiérrez-Preciado, A., Henkin, T.M., Grundy, F.J., Yanofsky, C. & Merino, E. Biochemical features and functional implications of RNA-based T-box regulatory mechanism. Microbiol. Mol. Biol. Rev. 73, 36–61 (2009).
Gong, F. & Yanofsky, C. Instruction of translating ribosome by nascent peptide. Science 297, 1864–1867 (2002).
Gong, F., Ito, K., Nakamura, Y. & Yanofsky, C. The mechanism of tryptophan induction of tryptophanase operon expression: tryptophan inhibits release factor-mediated cleavage of TnaC-peptidyl-tRNA(Pro). Proc. Natl. Acad. Sci. USA 98, 8997–9001 (2001).
Seidelt, B. et al. Structural insight into nascent polypeptide chain-mediated translational stalling. Science 326, 1412–1415 (2009).
Simons, R.W. & Kleckner, N. Translation control of IS10 transposition. Cell 34, 683–691 (1983).
Pédelacq, J.-D. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Isaacs, F.J. et al. Engineered riboregulators enable post-transcriptional control of gene expression. Nat. Biotechnol. 22, 841–847 (2004).
Kittle, J.D., Simons, R.W., Lee, J. & Kleckner, N. Insertion sequences IS10 anti-sense pairing initiates by an interaction between the 5′ end of the target RNA and a loop in the anti-sense RNA. J. Mol. Biol. 210, 561–572 (1989).
Qi, L., Haurwitz, R.E., Shao, W., Doudna, J.A. & Arkin, A.P. RNA processing enables predictable programming of gene expression. Nat. Biotechnol. doi:10.1038/nbt.2355. (Advanced online publication 16 September 2012).
Qi, L., Lucks, J.B., Liu, C.C., Mutalik, V.K. & Arkin, A.P. Engineering naturally occurring trans-acting non-coding RNAs to sense molecular signals. Nucleic Acids Res. 40, 5775–5786 (2012).
Cruz-Vera, L.R., Yang, R. & Yanofsky, C. Tryptophan inhibits Proteus vulgaris TnaC leader peptide elongation, activating tna operon expression. J. Bacteriol. 191, 7001–7006 (2009).
Low, J.T. & Weeks, K.M. SHAPE-directed RNA secondary structure prediction. Methods 52, 150–158 (2010).
Merino, E.J., Wilkinson, K.A., Coughlan, J.L. & Weeks, K.M. RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005).
Mortimer, S.A. & Weeks, K.M. A fast-acting reagent for accurate analysis of RNA secondary and tertiary structure by SHAPE chemistry. J. Am. Chem. Soc. 129, 4144–4145 (2007).
Mortimer, S.A. & Weeks, K.M. Time-resolved RNA SHAPE chemistry: quantitative RNA structure analysis in one-second snapshots and at single-nucleotide resolution. Nat. Protoc. 4, 1413–1421 (2009).
Tamsir, A., Tabor, J.J. & Voigt, C.A. Robust multicellular computing using genetically encoded NOR gates and chemical 'wires'. Nature 469, 212–215 (2011).
Acknowledgements
We thank C. Yanofsky and M. Samoilov for thoughtful discussions, D. Chen for experimental assistance and S. Meyer for assistance with SHAPE experiments. This work was funded by the US National Science Foundation as part of the Synthetic Biology Engineering Research Center (A.P.A.), the Miller Institute for Basic Scientific Research (C.C.L. and J.B.L.) and the US Department of Defense through the National Defense Science and Engineering Graduate Fellowship (T.H.S.-S.).
Author information
Authors and Affiliations
Contributions
C.C.L. conceived of the study, and A.P.A. advised in all aspects of the study. All authors were involved in designing the experiments. C.C.L., L.Q., J.B.L., T.H.S.-S. and V.K.M. performed experiments and interpreted the data. L.Q. and D.W. conducted the NOR gate experiments and the aptamer fusion experiments. J.B.L. conducted and analyzed the SHAPE experiments. C.C.L., L.Q. and A.P.A. wrote the manuscript. All authors discussed results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
US Provisional Patent number 61/540,413 is pending on this work.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1 and 2. (PDF 1981 kb)
Rights and permissions
About this article
Cite this article
Liu, C., Qi, L., Lucks, J. et al. An adaptor from translational to transcriptional control enables predictable assembly of complex regulation. Nat Methods 9, 1088–1094 (2012). https://doi.org/10.1038/nmeth.2184
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2184
This article is cited by
-
Programmable RNA-based systems for sensing and diagnostic applications
Analytical and Bioanalytical Chemistry (2019)
-
Advances in engineered trans-acting regulatory RNAs and their application in bacterial genome engineering
Journal of Industrial Microbiology and Biotechnology (2019)
-
High-throughput in vivo mapping of RNA accessible interfaces to identify functional sRNA binding sites
Nature Communications (2018)
-
Creating small transcription activating RNAs
Nature Chemical Biology (2015)
-
A versatile framework for microbial engineering using synthetic non-coding RNAs
Nature Reviews Microbiology (2014)