Abstract
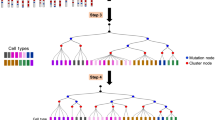
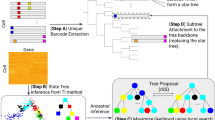
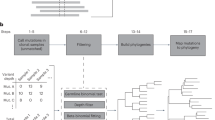
Because mutations are inevitable, the genome of each cell in a multicellular organism becomes unique and therefore encodes a record of its ancestry. Here we coupled arbitrary single primer PCR with next-generation DNA sequencing to catalog mutations and deconvolve the phylogeny of cultured mouse cells. This study helps pave the way toward construction of retrospective cell-fate maps based on mutations accumulating in genomes of somatic cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Frumkin, D., Wasserstrom, A., Kaplan, S., Feige, U. & Shapiro, E. PLoS Comput. Biol. 1, e50 (2005).
Salipante, S.J. & Horwitz, M.S. Proc. Natl. Acad. Sci. USA 103, 5448–5453 (2006).
Frumkin, D. et al. Cancer Res. 68, 5924–5931 (2008).
Wasserstrom, A. et al. PLoS ONE 3, e1939 (2008).
Wasserstrom, A. et al. PLoS Comput. Biol. 4, e1000058 (2008).
Salipante, S.J., Thompson, J.M. & Horwitz, M.S. Genetics 178, 967–977 (2008).
Salipante, S.J., Kas, A., McMonagle, E. & Horwitz, M.S. Evol. Dev. 12, 84–94 (2010).
Shibata, D. & Tavare, S. Stem Cell Rev. 3, 94–103 (2007).
Dasgupta, S. et al. PLoS ONE 4, e6533 (2009).
Fellous, T.G. et al. Stem Cells 27, 1410–1420 (2009).
Navin, N. et al. Nature 472, 90–94 (2011).
Sulston, J.E. ChemBioChem 4, 688–696 (2003).
Preston, B.D., Albertson, T.M. & Herr, A.J. Semin. Cancer Biol. 20, 281–293 (2010).
McClelland, M. & Welsh, J. PCR Methods Appl. 4, S59–S65 (1994).
Li, H., Ruan, J. & Durbin, R. Genome Res. 18, 1851–1858 (2008).
Nye, T.M., Lio, P. & Gilks, W.R. Bioinformatics 22, 117–119 (2006).
Petit, A.C., Legue, E. & Nicolas, J.F. Reprod. Nutr. Dev. 45, 321–339 (2005).
Baker, S.M. et al. Nat. Genet. 13, 336–342 10.1038/ng0796-336 (1996).
Albertson, T.M. et al. Proc. Natl. Acad. Sci. USA 106, 17101–17104 (2009).
Zheng, Q. Math. Biosci. 196, 198–214 (2005).
Zheng, Q. Math. Biosci. 216, 150–153 (2008).
Croyle, M.L., Woo, A.L. & Lingrel, J.B. Eur. J. Biochem. 248, 488–495 (1997).
Fallows, D. et al. Mol. Cell. Biol. 7, 2985–2987 (1987).
Price, E.M. & Lingrel, J.B. Biochemistry 27, 8400–8408 (1988).
Cantley, L.G., Cunha, M.J. & Zhou, X.M. J. Biol. Chem. 269, 15358–15361 (1994).
Li, H. et al. Bioinformatics 25, 2078–2079 (2009).
Felsenstein, J. Inferring Phylogenies (Sinauer Associates, 2004).
Ronquist, F. & Huelsenbeck, J.P. Bioinformatics 19, 1572–1574 (2003).
Acknowledgements
We thank D. Anderson, J. Salk and L. Loeb for discussion and comments. This work was supported by US National Institutes of Health grants DP1OD003278 and R01DK078340 (to M.S.H.), R01CA111582 and R01CA098243 (to B.D.P.), T32HL007093 (for C.A.C.), F30AG030316 (to S.J.S.) and T32GM007266 (to M.S.H.), and Achievement Rewards for College Scientists Foundation fellowship grants to the University of Washington Medical Scientist Training Program (for S.J.S.).
Author information
Authors and Affiliations
Contributions
C.A.C., B.D.P., L.E.H., S.J.S. and M.S.H. designed the experiments. C.A.C., S.J.S. and L.E.H. carried out the experiments. C.A.C., A.K., R.K. and M.S.H. contributed to analyzing the data. M.S.H., with input from other authors, wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1,3,4 (PDF 3453 kb)
Supplementary Table 2
Compilation of identified mutations. (XLS 167 kb)
Supplementary Software
PrimerDesign.pl is a Perl program for evaluation of arbitrary primers; ChromosomePrep.pl is a Perl program to prepare mouse reference sequence for PrimerDesign.pl; and Geno.pl is a Perl program to perform mutational analysis. (ZIP 9 kb)
Rights and permissions
About this article
Cite this article
Carlson, C., Kas, A., Kirkwood, R. et al. Decoding cell lineage from acquired mutations using arbitrary deep sequencing. Nat Methods 9, 78–80 (2012). https://doi.org/10.1038/nmeth.1781
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1781
This article is cited by
-
The use of technical replication for detection of low-level somatic mutations in next-generation sequencing
Nature Communications (2019)
-
Sample path properties of the average generation of a Bellman–Harris process
Journal of Mathematical Biology (2019)
-
UbC-StarTrack, a clonal method to target the entire progeny of individual progenitors
Scientific Reports (2016)
-
Inferring average generation via division-linked labeling
Journal of Mathematical Biology (2016)
-
Barrett oesophagus: lessons on its origins from the lesion itself
Nature Reviews Gastroenterology & Hepatology (2015)