Abstract
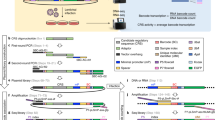
We developed a high-content reporter system that allows quantitative assessment of activities of multiple transcription factors (TFs) in a eukaryotic cell. The system comprises a library of reporter constructs that are evaluated according to their transcription rates. All reporters produce essentially identical messages that are subjected to 'processing', which generates a spectrum of distinguishable fragments that are analyzed quantitatively. The homogeneity of the reporter library afforded inherently uniform detection conditions for all reporters and provided repeatability, accuracy and robustness of assessment. We showed that this technology can be used to identify pathways transmitting cell responses to inducers, and that the profile of TF activities generated using this system represents a stable and sustained cell signature that clearly distinguishes different cell types and pathological conditions. This technology provides a framework for functional characterization of signal transduction networks through profiling activities of multiple TFs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Babu, M.M., Luscombe, N.M., Aravind, L., Gerstein, M. & Teichmann, S.A. Structure and evolution of transcriptional regulatory networks. Curr. Opin. Struct. Biol. 14, 283–291 (2004).
Stegmaier, P., Kel, A.E. & Wingender, E. Systematic DNA-binding domain classification of transcription factors. Genome Inform. 15, 276–286 (2004).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34 Database issue, D108–D110 (2006).
Benotmane, A.M., Hoylaerts, M.F., Collen, D. & Belayew, A. Nonisotopic quantitative analysis of protein-DNA interactions at equilibrium. Anal. Biochem. 250, 181–185 (1997).
Rosenau, C., Emery, D., Kaboord, B. & Qoronfleh, M.W. Development of a high-throughput plate-based chemiluminescent transcription factor assay. J. Biomol. Screen. 9, 334–342 (2004).
Hermanson, O., Glass, C.K. & Rosenfeld, M.G. Nuclear receptor coregulators: multiple modes of modification. Trends Endocrinol. Metab. 13, 55–60 (2002).
Bronstein, I., Fortin, J., Stanley, P.E., Stewart, G.S. & Kricka, L.J. Chemiluminescent and bioluminescent reporter gene assays. Anal. Biochem. 219, 169–181 (1994).
Kovacs, D.M. & Kaplan, B.B. Discordant estimates of heterologous promoter activity as determined by reporter gene mRNA levels and enzyme activity. Biochem. Biophys. Res. Commun. 189, 912–918 (1992).
Li, X., Jiang, X. & Yaoi, T. High throughput assays for analyzing transcription factors. Assay Drug Dev. Technol. 4, 333–341 (2006).
Guhaniyogi, J. & Brewer, G. Regulation of mRNA stability in mammalian cells. Gene 265, 11–23 (2001).
Shim, J. & Karin, M. The control of mRNA stability in response to extracellular stimuli. Mol. Cells 14, 323–331 (2002).
Levitt, N., Briggs, D., Gil, A. & Proudfoot, N.J. Definition of an efficient synthetic poly(A) site. Genes Dev. 3, 1019–1025 (1989).
Enriquez-Harris, P., Levitt, N., Briggs, D. & Proudfoot, N.J. A pause site for RNA polymerase II is associated with termination of transcription. EMBO J. 10, 1833–1842 (1991).
MAQC Consortium. The MicroArray Quality Control (MAQC) project shows inter- and intraplatform reproducibility of gene expression measurements. Nat. Biotechnol. 24, 1151–1161 (2006).
Oberg, M., Bergander, L., Hakansson, H., Rannug, U. & Rannug A. Identification of the tryptophan photoproduct 6-formylindolo[3,2-b]carbazole, in cell culture medium, as a factor that controls the background aryl hydrocarbon receptor activity. Toxicol. Sci. 85, 935–943 (2005).
Montminy, M.R. & Bilezikjian, L.M. Binding of a nuclear protein to the cyclic-AMP response element of the somatostatin gene. Nature 328, 175–178 (1987).
Dennler, S. et al. Direct binding of Smad3 and Smad4 to critical TGF beta-inducible elements in the promoter of human plasminogen activator inhibitor-type 1 gene. EMBO J. 17, 3091–3100 (1998).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Zhang, L. et al. Gene expression profiles in normal and cancer cells. Science 276, 1268–1272 (1997).
Perou, C.M. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000).
Okabe, H. et al. Genome-wide analysis of gene expression in human hepatocellular carcinomas using cDNA microarray: identification of genes involved in viral carcinogenesis and tumor progression. Cancer Res. 61, 2129–2137 (2001).
Ross, D.T. et al. Systematic variation in gene expression patterns in human cancer cell lines. Nat. Genet. 24, 227–235 (2000).
Korinek, V. et al. Constitutive transcriptional activation by a beta-catenin-Tcf complex in APC−/− colon carcinoma. Science 275, 1784–1787 (1997).
Morin, P.J. et al. Activation of beta-catenin-Tcf signaling in colon cancer by mutations in beta-catenin or APC. Science 275, 1787–1790 (1997).
Nakshatri, H., Bhat-Nakshatri, P., Martin, D.A., Goulet, R.J. Jr. & Sledge, G.W. Jr. Constitutive activation of NF-kappaB during progression of breast cancer to hormone-independent growth. Mol. Cell. Biol. 17, 3629–3639 (1997).
Troester, M.A. et al. Cell-type-specific responses to chemotherapeutics in breast cancer. Cancer Res. 64, 4218–4226 (2004).
Waldman, T., Lengauer, C., Kinzler, K.W. & Vogelstein, B. Uncoupling of S phase and mitosis induced by anticancer agents in cells lacking p21. Nature 381, 713–716 (1996).
Goyette, M.C. et al. Progression of colorectal cancer is associated with multiple tumor suppressor gene defects but inhibition of tumorigenicity is accomplished by correction of any single defect via chromosome transfer. Mol. Cell. Biol. 12, 1387–1395 (1992).
Shao, Z.M. et al. p53 independent G0/G1 arrest and apoptosis induced by a novel retinoid in human breast cancer cells. Oncogene 11, 493–504 (1995).
Sugikawa, E. et al. Mutant p53 mediated induction of cell cycle arrest and apoptosis at G1 phase by 9-hydroxyellipticine. Anticancer Res. 19, 3099–3108 (1999).
Tamaru, N. et al. Estrogen receptor-associated expression of keratinocyte growth factor and its possible role in the inhibition of apoptosis in human breast cancer. Lab. Invest. 84, 1460–1471 (2004).
Nuwaysir, E.F., Bittner, M., Trent, J., Barrett, J.C. & Afshari, C.A. Microarrays and toxicology: the advent of toxicogenomics. Mol. Carcinog. 24, 153–159 (1999).
Acknowledgements
We thank S. Langdon for invaluable help with fragment analysis, and A.S. Baldwin and R. Gaynor for discussions. The work was supported by US National Institutes of Health grants 1R43CA101636, 1R43CA101271 and UO1 AI061360.
Author information
Authors and Affiliations
Contributions
S.R., A.M. and S.M. are authors of the concept. A.M. and M.M. constructed the RTU library; A.M. and S.R. designed the experiments; Ma.G., M.M., N.P., L.M. and L.D. performed the experiments; Mi.G. designed the software; S.R. and S.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.R., A.M., Ma.G., N.P., M.M., L.M. and S.M. are current employees of Attagene Inc. S.R., L.D. and S.M. have significant financial interest in company.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Figures 1–4, Supplementary Tables 1–5 (PDF 338 kb)
Rights and permissions
About this article
Cite this article
Romanov, S., Medvedev, A., Gambarian, M. et al. Homogeneous reporter system enables quantitative functional assessment of multiple transcription factors. Nat Methods 5, 253–260 (2008). https://doi.org/10.1038/nmeth.1186
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1186
This article is cited by
-
Comprehensive assessment of NR ligand polypharmacology by a multiplex reporter NR assay
Scientific Reports (2022)
-
Quantitative, solution-phase profiling of multiple transcription factors in parallel
Analytical and Bioanalytical Chemistry (2013)
-
The living microarray: a high-throughput platform for measuring transcription dynamics in single cells
BMC Genomics (2011)
-
Integrated analysis of receptor activation and downstream signaling with EXTassays
Nature Methods (2010)
-
Cloning-free regulated monitoring of reporter and gene expression
BMC Molecular Biology (2009)