Abstract
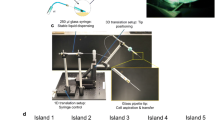
The profound metabolic reprogramming that occurs in cancer cells has been investigated primarily in two-dimensional cell cultures, which fail to recapitulate spatial aspects of cell-to-cell interactions as well as tissue gradients present in three-dimensional tumours. Here, we describe an engineered model to assemble three-dimensional tumours by rolling a scaffold–tumour composite strip. By unrolling the strip, the model can be rapidly disassembled for snapshot analysis, allowing spatial mapping of cell metabolism in concert with cell phenotype. We also show that the establishment of oxygen gradients within samples that are shaped by oxygen-dependent signalling pathways, as well as the consequential variations in cell growth, response to hypoxic gradients extending from normoxia to severe hypoxia, and therapy responsiveness, are consistent with those of tumours in vivo. Moreover, by using liquid chromatography tandem mass spectrometry, we mapped cellular metabolism and identified spatially defined metabolic signatures of cancer cells to reveal both known and novel metabolic responses to hypoxia.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
01 December 2015
Corrected online 1 December 2015. In the version of the Article originally published online, in Fig. 1 there were some image display errors in panels a and b and the label 'GFP SK-OV-3' should have been green in panels c and e. In Fig. 2, panels a and b, layer numbers should have been defined. These errors have now been corrected in all versions of the Article.
References
Cairns, R. A., Harris, I. S. & Mak, T. W. Regulation of cancer cell metabolism. Nature Rev. Cancer 11, 85–95 (2011).
Carmeliet, P. & Jain, R. K. Principles and mechanisms of vessel normalization for cancer and other angiogenic diseases. Nature Rev. Drug Discov. 10, 417–427 (2011).
Li, X. F. et al. Visualization of hypoxia in microscopic tumors by immunofluorescent microscopy. Cancer Res. 67, 7646–7653 (2007).
Wouters, B. G. et al. Modulation of cell death in the tumor microenvironment. Semin. Radiat. Oncol. 13, 31–41 (2003).
Semenza, G. L. Regulation of metabolism by hypoxia-inducible factor 1. Cold Spring Harb. Symp. Quant. Biol. 76, 347–353 (2011).
Wouters, B. G. & Koritzinsky, M. Hypoxia signalling through mTOR and the unfolded protein response in cancer. Nature Rev. Cancer 8, 851–864 (2008).
Ron, D. & Walter, P. Signal integration in the endoplasmic reticulum unfolded protein response. Nature Rev. Mol. Cell Biol. 8, 519–529 (2007).
Bi, M. et al. ER stress-regulated translation increases tolerance to extreme hypoxia and promotes tumor growth. EMBO J. 24, 3470–3481 (2005).
Guillaumond, F. et al. Strengthened glycolysis under hypoxia supports tumor symbiosis and hexosamine biosynthesis in pancreatic adenocarcinoma. Proc. Natl Acad. Sci. USA 110, 3919–3924.
Mueller-Klieser, W. Multicellular spheroids. A review on cellular aggregates in cancer research. J. Cancer Res. Clin. Oncol. 113, 101–122 (1987).
Fischbach, C. et al. Engineering tumors with 3D scaffolds. Nature Methods 4, 855–860 (2007).
Infanger, D. W., Lynch, M. E. & Fischbach, C. Engineered culture models for studies of tumor-microenvironment interactions. Annu. Rev. Biomed. Eng. 15, 29–53 (2013).
McGuigan, A. P. & Sefton, M. V. Vascularized organoid engineered by modular assembly enables blood perfusion. Proc. Natl Acad. Sci. USA 103, 11461–11466 (2006).
Derda, R. et al. Paper-supported 3D cell culture for tissue-based bioassays. Proc. Natl Acad. Sci. USA 106, 18457–18462 (2009).
Carmeliet, P. & Jain, R. K. Angiogenesis in cancer and other diseases. Nature 407, 249–257 (2000).
Paszek, M. J. et al. Tensional homeostasis and the malignant phenotype. Cancer Cell 8, 241–254 (2005).
Minchinton, A. I. & Tannock, I. F. Drug penetration in solid tumours. Nature Rev. Cancer 6, 583–592 (2006).
Primeau, A. J., Rendon, A., Hedley, D., Lilge, L. & Tannock, I. F. The distribution of the anticancer drug Doxorubicin in relation to blood vessels in solid tumors. Clin. Cancer Res. 11, 8782–8788 (2005).
Horsman, M. R., Mortensen, L. S., Petersen, J. B., Busk, M. & Overgaard, J. Imaging hypoxia to improve radiotherapy outcome. Nature Rev. Clin. Oncol. 9, 674–687 (2012).
Waleh, N. S. et al. Mapping of the vascular endothelial growth factor-producing hypoxic cells in multicellular tumor spheroids using a hypoxia-specific marker. Cancer Res. 55, 6222–6226 (1995).
Koch, C. J. Measurement of absolute oxygen levels in cells and tissues using oxygen sensors and 2-nitroimidazole EF5. Methods Enzymol. 352, 3–31 (2002).
Dang, C. V. Links between metabolism and cancer. Genes Dev. 26, 877–890 (2012).
Latasa, M. U. et al. Identification of argininosuccinate lyase as a hypoxia-responsive gene in rat hepatocytes. J. Hepatol. 33, 709–715 (2000).
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009).
Wise, D. R. et al. Hypoxia promotes isocitrate dehydrogenase-dependent carboxylation of α-ketoglutarate to citrate to support cell growth and viability. Proc. Natl Acad. Sci. USA 108, 19611–19616 (2011).
Tribble, D. L. & Jones, D. P. Oxygen dependence of oxidative stress. Rate of NADPH supply for maintaining the GSH pool during hypoxia. Biochem. Pharmacol. 39, 729–736 (1990).
Chouchani, E. T. et al. Ischaemic accumulation of succinate controls reperfusion injury through mitochondrial ROS. Nature 515, 431–435 (2014).
Schmidt, S. K. et al. Regulation of IDO activity by oxygen supply: Inhibitory effects on antimicrobial and immunoregulatory functions. PLoS ONE 8, e63301 (2013).
Platten, M., Wick, W. & Van den Eynde, B. J. Tryptophan catabolism in cancer: beyond IDO and tryptophan depletion. Cancer Res. 72, 5435–5440 (2012).
Evans, S. M., Hahn, S. M., Magarelli, D. P. & Koch, C. J. Hypoxic heterogeneity in human tumors: EF5 binding, vasculature, necrosis, and proliferation. Am. J. Clin. Oncol. 24, 467–472 (2001).
Acknowledgements
The authors acknowledge V. Bindokas (University of Chicago), M. Macasaet-Peralta (Pathology Research Program, UHN), J. Cathcart (AOMF), J. Stewart and D. Scollard (STTARR), B. Calvieri, S. Doyle, J. Soleas, S. Javaherian, C. Londono, C. Crossman, R. Vellanki, J. Han, M. Young and S. Lakhani (University of Toronto) for technical assistance. This work was funded by a Natural Science and Engineering Council Discovery Accelerator Supplement (RGPIN-314056) to A.P.M., a YSF NSERC fellowship to D.R., MRC Cancer Unit Core Funding to C.F. and E.G., Ontario Ministry of Health and Long Term Care (OMOHLTC), the Terry Fox New Frontiers Research Program (PPG09-020005) and the Canadian Institute for Health Research (CIHR grant 201592) grants to B.G.W., and by a Ontario Graduate Scholarship to D.C.
Author information
Authors and Affiliations
Contributions
D.R., E.G., D.C., R.M., B.G.W., C.F. and A.P.M. designed experiments, analysed the data and wrote the manuscript. D.R., E.G. and D.C. conducted experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 69958 kb)
Rights and permissions
About this article
Cite this article
Rodenhizer, D., Gaude, E., Cojocari, D. et al. A three-dimensional engineered tumour for spatial snapshot analysis of cell metabolism and phenotype in hypoxic gradients. Nature Mater 15, 227–234 (2016). https://doi.org/10.1038/nmat4482
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmat4482
This article is cited by
-
Middle-out methods for spatiotemporal tissue engineering of organoids
Nature Reviews Bioengineering (2023)
-
PACER: a novel 3D plant cell wall model for the analysis of non-catalytic and enzymatic responses
Biotechnology for Biofuels and Bioproducts (2022)
-
Integrating a dynamic central metabolism model of cancer cells with a hybrid 3D multiscale model for vascular hepatocellular carcinoma growth
Scientific Reports (2022)
-
Crosstalk between mechanotransduction and metabolism
Nature Reviews Molecular Cell Biology (2021)
-
Oncometabolites in renal cancer
Nature Reviews Nephrology (2020)