Abstract
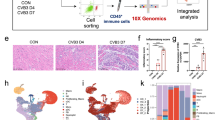
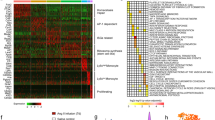
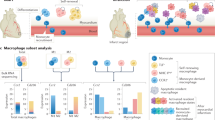
Interferon regulatory factor 3 (IRF3) and type I interferons (IFNs) protect against infections1 and cancer2, but excessive IRF3 activation and type I IFN production cause autoinflammatory conditions such as Aicardi–Goutières syndrome3,4 and STING-associated vasculopathy of infancy (SAVI)3. Myocardial infarction (MI) elicits inflammation5, but the dominant molecular drivers of MI-associated inflammation remain unclear. Here we show that ischemic cell death and uptake of cell debris by macrophages in the heart fuel a fatal response to MI by activating IRF3 and type I IFN production. In mice, single-cell RNA-seq analysis of 4,215 leukocytes isolated from infarcted and non-infarcted hearts showed that MI provokes activation of an IRF3–interferon axis in a distinct population of interferon-inducible cells (IFNICs) that were classified as cardiac macrophages. Mice genetically deficient in cyclic GMP-AMP synthase (cGAS), its adaptor STING, IRF3, or the type I IFN receptor IFNAR exhibited impaired interferon-stimulated gene (ISG) expression and, in the case of mice deficient in IRF3 or IFNAR, improved survival after MI as compared to controls. Interruption of IRF3-dependent signaling resulted in decreased cardiac expression of inflammatory cytokines and chemokines and decreased inflammatory cell infiltration of the heart, as well as in attenuated ventricular dilation and improved cardiac function. Similarly, treatment of mice with an IFNAR-neutralizing antibody after MI ablated the interferon response and improved left ventricular dysfunction and survival. These results identify IRF3 and the type I IFN response as a potential therapeutic target for post-MI cardioprotection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Goubau, D., Deddouche, S. & Reis e Sousa, C. Cytosolic sensing of viruses. Immunity 38, 855–869 (2013).
Woo, S.R. et al. STING-dependent cytosolic DNA sensing mediates innate immune recognition of immunogenic tumors. Immunity 41, 830–842 (2014).
Liu, Y. et al. Activated STING in a vascular and pulmonary syndrome. N. Engl. J. Med. 371, 507–518 (2014).
Gall, A. et al. Autoimmunity initiates in nonhematopoietic cells and progresses via lymphocytes in an interferon-dependent autoimmune disease. Immunity 36, 120–131 (2012).
Swirski, F.K. & Nahrendorf, M. Leukocyte behavior in atherosclerosis, myocardial infarction, and heart failure. Science 339, 161–166 (2013).
Frangogiannis, N.G. The inflammatory response in myocardial injury, repair, and remodelling. Nat. Rev. Cardiol. 11, 255–265 (2014).
Mann, D.L. Innate immunity and the failing heart: the cytokine hypothesis revisited. Circ. Res. 116, 1254–1268 (2015).
Heusch, G. Cardioprotection: chances and challenges of its translation to the clinic. Lancet 381, 166–175 (2013).
Cai, X., Chiu, Y.H. & Chen, Z.J. The cGAS–cGAMP–STING pathway of cytosolic DNA sensing and signaling. Mol. Cell 54, 289–296 (2014).
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z.J. Cyclic GMP–AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013).
Wu, J. et al. Cyclic GMP–AMP is an endogenous second messenger in innate immune signaling by cytosolic DNA. Science 339, 826–830 (2013).
Ishikawa, H., Ma, Z. & Barber, G.N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Fitzgerald, K.A. et al. IKKɛ and TBK1 are essential components of the IRF3 signaling pathway. Nat. Immunol. 4, 491–496 (2003).
West, A.P. et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature 520, 553–557 (2015).
Gao, D. et al. Activation of cyclic GMP–AMP synthase by self-DNA causes autoimmune diseases. Proc. Natl. Acad. Sci. USA 112, E5699–E5705 (2015).
Yoshida, H., Okabe, Y., Kawane, K., Fukuyama, H. & Nagata, S. Lethal anemia caused by interferon-β produced in mouse embryos carrying undigested DNA. Nat. Immunol. 6, 49–56 (2005).
Andersen, J., VanScoy, S., Cheng, T.F., Gomez, D. & Reich, N.C. IRF-3-dependent and augmented target genes during viral infection. Genes Immun. 9, 168–175 (2008).
Grandvaux, N. et al. Transcriptional profiling of interferon regulatory factor 3 target genes: direct involvement in the regulation of interferon-stimulated genes. J. Virol. 76, 5532–5539 (2002).
Kühn, R., Schwenk, F., Aguet, M. & Rajewsky, K. Inducible gene targeting in mice. Science 269, 1427–1429 (1995).
Epelman, S. et al. Embryonic and adult-derived resident cardiac macrophages are maintained through distinct mechanisms at steady state and during inflammation. Immunity 40, 91–104 (2014).
Liu, S. et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347, aaa2630 (2015).
Takeda, K. & Akira, S. TLR signaling pathways. Semin. Immunol. 16, 3–9 (2004).
Yoneyama, M. & Fujita, T. RNA recognition and signal transduction by RIG-I-like receptors. Immunol. Rev. 227, 54–65 (2009).
Sauer, J.D. et al. The N-ethyl-N-nitrosourea-induced Goldenticket mouse mutant reveals an essential function of Sting in the in vivo interferon response to Listeriamonocytogenes and cyclic dinucleotides. Infect. Immun. 79, 688–694 (2011).
Manzanillo, P.S., Shiloh, M.U., Portnoy, D.A. & Cox, J.S. Mycobacteriumtuberculosis activates the DNA-dependent cytosolic surveillance pathway within macrophages. Cell Host Microbe 11, 469–480 (2012).
Hasan, M. et al. Trex1 regulates lysosomal biogenesis and interferon-independent activation of antiviral genes. Nat. Immunol. 14, 61–71 (2013).
Essers, M.A. et al. IFNα activates dormant haematopoietic stem cells in vivo. Nature 458, 904–908 (2009).
Heymans, S. et al. Inhibition of plasminogen activators or matrix metalloproteinases prevents cardiac rupture but impairs therapeutic angiogenesis and causes cardiac failure. Nat. Med. 5, 1135–1142 (1999).
Corrales, L. & Gajewski, T.F. Molecular pathways: targeting the stimulator of interferon genes (STING) in the immunotherapy of cancer. Clin. Cancer Res. 21, 4774–4779 (2015).
Fu, J. et al. STING agonist formulated cancer vaccines can cure established tumors resistant to PD-1 blockade. Sci. Transl. Med. 7, 283ra52 (2015).
Hanson, M.C. et al. Nanoparticulate STING agonists are potent lymph node–targeted vaccine adjuvants. J. Clin. Invest. 125, 2532–2546 (2015).
Furie, R. et al. Anifrolumab, an anti-interferon-α receptor monoclonal antibody, in moderate-to-severe systemic lupus erythematosus. Arthritis Rheumatol. 69, 376–386 (2017).
Khamashta, M. et al. Sifalimumab, an anti-interferon-α monoclonal antibody, in moderate to severe systemic lupus erythematosus: a randomised, double-blind, placebo-controlled study. Ann. Rheum. Dis. 75, 1909–1916 (2016).
Goldberg, A. et al. Dose-escalation of human anti-interferon-α receptor monoclonal antibody MEDI-546 in subjects with systemic sclerosis: a phase 1, multicenter, open label study. Arthritis Res. Ther. 16, R57 (2014).
Muzumdar, M.D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Sun, Q. et al. The specific and essential role of MAVS in antiviral innate immune responses. Immunity 24, 633–642 (2006).
Hoebe, K. et al. Identification of Lps2 as a key transducer of MyD88-independent TIR signalling. Nature 424, 743–748 (2003).
Sato, M. et al. Distinct and essential roles of transcription factors IRF-3 and IRF-7 in response to viruses for IFN-α/β gene induction. Immunity 13, 539–548 (2000).
Kamran, P. et al. Parabiosis in mice: a detailed protocol. J. Vis. Exp. (80), e50556 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P.T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Anders, S. et al. Count-based differential expression analysis of RNA sequencing data using R and Bioconductor. Nat. Protoc. 8, 1765–1786 (2013).
Huang, W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang, W., Sherman, B.T. & Lempicki, R.A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Ahn, J., Gutman, D., Saijo, S. & Barber, G.N. STING manifests self DNA–dependent inflammatory disease. Proc. Natl. Acad. Sci. USA 109, 19386–19391 (2012).
Schoggins, J.W. et al. Pan-viral specificity of IFN-induced genes reveals new roles for cGAS in innate immunity. Nature 505, 691–695 (2014).
Zilionis, R. et al. Single-cell barcoding and sequencing using droplet microfluidics. Nat. Protoc. 12, 44–73 (2017).
Klein, A.M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Satija, R., Farrell, J.A., Gennert, D., Schier, A.F. & Regev, A. Spatial reconstruction of single-cell gene expression data. Nat. Biotechnol. 33, 495–502 (2015).
Aguirre, A.D., Vinegoni, C., Sebas, M. & Weissleder, R. Intravital imaging of cardiac function at the single-cell level. Proc. Natl. Acad. Sci. USA 111, 11257–11262 (2014).
Lee, S. et al. Real-time in vivo imaging of the beating mouse heart at microscopic resolution. Nat. Commun. 3, 1054 (2012).
Acknowledgements
We thank T. Taniguchi (University of Tokyo) and M. Diamond (Washington University School of Medicine) for their gift of the Irf3−/− mice; P. Dutta, H. Sager, M. Hulsman, G. Courties, R. Giedt, and K. Yang for helpful discussions and technical assistance; B. Tricot for technical assistance with cardiac MRI as part of the Molecular Imaging Program core; and the Harvard Stem Cell Institute Flow Cytometry Core, the Massachusetts General Hospital NextGen Sequencing Core, the National Mouse Metabolic Phenotyping Center at the University of Massachusetts funded by US NIH grant 2U2C-DK093000, and the Single-Cell Core at Harvard Medical School. The work was funded in part by NIH-NHLBI grant T32HL007604, the Harvard Medical School LaDue Fellowship, and AHA17IRG33410543 and by NIH-NHLBI grants 1K99HL129168 and R00HL129168 (K.R.K.); AHA14FTF20380185 (A.D.A.); AHA16FTF29630016 and the Yeatts Fund for Innovative Research (J.D.R.); AHA16POST27030088 and Deutsche Herzstiftung S/05/12 (Y.-X.Y.); grant R01HL117829 (M.N.); grant NIH-R01 HL080472 and the RRM Charitable Fund (P.L.); and grant 5R01HL122208 (R.W.).
Author information
Authors and Affiliations
Contributions
K.R.K. and A.D.A. designed and performed the experiments, analyzed the data, and wrote the manuscript; Y.-X.Y. designed and performed experiments and analyzed data; M.K. performed MI on WT, Ifnar−/−, and cGAS−/− mice with T.P.F.; Y.S. performed MI on WT mice and all other mouse strains; A.S. and R.I.S. performed bioinformatics analysis; Y.I. performed histological analysis; R.P.N. performed biomolecular analysis; J.D.R. performed echocardiography and data analysis, R.H.K. performed confocal imaging, and S.P.A. performed bone marrow–derived macrophage experiments; T.M., K.A.F., and P.L. provided guidance on experimental design; and M.N. and R.W. designed experiments, analyzed data, and revised the manuscript. All authors reviewed results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures and Tables
Supplementary Figures 1–12 and Supplementary Tables 1–7 (PDF 43710 kb)
Rights and permissions
About this article
Cite this article
King, K., Aguirre, A., Ye, YX. et al. IRF3 and type I interferons fuel a fatal response to myocardial infarction. Nat Med 23, 1481–1487 (2017). https://doi.org/10.1038/nm.4428
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4428
This article is cited by
-
The combination of Tanshinone IIA and Astragaloside IV attenuates myocardial ischemia–reperfusion injury by inhibiting the STING pathway
Chinese Medicine (2024)
-
TREM2hi resident macrophages protect the septic heart by maintaining cardiomyocyte homeostasis
Nature Metabolism (2023)
-
Mitochondrial DNA-triggered innate immune response: mechanisms and diseases
Cellular & Molecular Immunology (2023)
-
CRAT links cholesterol metabolism to innate immune responses in the heart
Nature Metabolism (2023)
-
Hallmarks of cardiovascular ageing
Nature Reviews Cardiology (2023)