Abstract
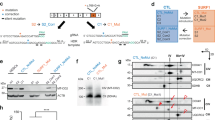
Smith-Lemli-Opitz syndrome (SLOS) is a malformation disorder caused by mutations in DHCR7, which impair the reduction of 7-dehydrocholesterol (7DHC) to cholesterol. SLOS results in cognitive impairment, behavioral abnormalities and nervous system defects, though neither affected cell types nor impaired signaling pathways are fully understood. Whether 7DHC accumulation or cholesterol loss is primarily responsible for disease pathogenesis is also unclear. Using induced pluripotent stem cells (iPSCs) from subjects with SLOS, we identified cellular defects that lead to precocious neuronal specification within SLOS derived neural progenitors. We also demonstrated that 7DHC accumulation, not cholesterol deficiency, is critical for SLOS-associated defects. We further identified downregulation of Wnt/β-catenin signaling as a key initiator of aberrant SLOS iPSC differentiation through the direct inhibitory effects of 7DHC on the formation of an active Wnt receptor complex. Activation of canonical Wnt signaling prevented the neural phenotypes observed in SLOS iPSCs, suggesting that Wnt signaling may be a promising therapeutic target for SLOS.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Porter, F.D. & Herman, G.E. Malformation syndromes caused by disorders of cholesterol synthesis. J. Lipid Res. 52, 6–34 (2011).
Wassif, C.A. et al. Mutations in the human sterol Δ7-reductase gene at 11q12–13 cause Smith-Lemli-Opitz syndrome. Am. J. Hum. Genet. 63, 55–62 (1998).
Spector, A.A. & Yorek, M.A. Membrane lipid composition and cellular function. J. Lipid Res. 26, 1015–1035 (1985).
Lingwood, D. & Simons, K. Lipid rafts as a membrane-organizing principle. Science 327, 46–50 (2010).
Jurevics, H. & Morell, P. Cholesterol for synthesis of myelin is made locally, not imported into brain. J. Neurochem. 64, 895–901 (1995).
Mellon, S.H. Neurosteroid regulation of central nervous system development. Pharmacol. Ther. 116, 107–124 (2007).
Lee, R.W., Conley, S.K., Gropman, A., Porter, F.D. & Baker, E.H. Brain magnetic resonance imaging findings in Smith-Lemli-Opitz syndrome. Am. J. Med. Genet. A. 161A, 2407–2419 (2013).
Tierney, E. et al. Behavior phenotype in the RSH/Smith-Lemli-Opitz syndrome. Am. J. Med. Genet. 98, 191–200 (2001).
Gaoua, W., Chevy, F., Roux, C. & Wolf, C. Oxidized derivatives of 7-dehydrocholesterol induce growth retardation in cultured rat embryos: a model for antenatal growth retardation in the Smith-Lemli-Opitz syndrome. J. Lipid Res. 40, 456–463 (1999).
Xu, L., Korade, Z., Rosado, D.A. Jr., Mirnics, K. & Porter, N.A. Metabolism of oxysterols derived from nonenzymatic oxidation of 7-dehydrocholesterol in cells. J. Lipid Res. 54, 1135–1143 (2013).
Somers, A. et al. Generation of transgene-free lung disease-specific human induced pluripotent stem cells using a single excisable lentiviral stem cell cassette. Stem Cells 28, 1728–1740 (2010).
Hu, B.Y . et al. Neural differentiation of human induced pluripotent stem cells follows developmental principles but with variable potency. Proc. Natl. Acad. Sci. USA 107, 4335–4340 (2010).
Ludwig, T.E. et al. Derivation of human embryonic stem cells in defined conditions. Nat. Biotechnol. 24, 185–187 (2006).
Jiang, X.S. et al. Activation of Rho GTPases in Smith-Lemli-Opitz syndrome: pathophysiological and clinical implications. Hum. Mol. Genet. 19, 1347–1357 (2010).
Elkabetz, Y. et al. Human ES cell-derived neural rosettes reveal a functionally distinct early neural stem cell stage. Genes Dev. 22, 152–165 (2008).
Cenedella, R.J. & Bierkamper, G.G. Mechanism of cataract production by 3-β(2-diethylaminoethoxy) androst-5-en-17-one hydrochloride, U18666A: an inhibitor of cholesterol biosynthesis. Exp. Eye Res. 28, 673–688 (1979).
Gofflot, F., Kolf-Clauw, M., Clotman, F., Roux, C. & Picard, J.J. Absence of ventral cell populations in the developing brain in a rat model of the Smith-Lemli-Opitz syndrome. Am. J. Med. Genet. 87, 207–216 (1999).
Frolov, A. et al. Cholesterol overload promotes morphogenesis of a Niemann-Pick C (NPC)-like compartment independent of inhibition of NPC1 or HE1/NPC2 function. J. Biol. Chem. 276, 46414–46421 (2001).
Wassif, C.A. et al. Cholesterol storage defect in RSH/Smith-Lemli-Opitz syndrome fibroblasts. Mol. Genet. Metab. 75, 325–334 (2002).
Krakowiak, P.A. et al. Lathosterolosis: an inborn error of human and murine cholesterol synthesis due to lathosterol 5-desaturase deficiency. Hum. Mol. Genet. 12, 1631–1641 (2003).
Sato, N., Meijer, L., Skaltsounis, L., Greengard, P. & Brivanlou, A.H. Maintenance of pluripotency in human and mouse embryonic stem cells through activation of Wnt signaling by a pharmacological GSK-3–specific inhibitor. Nat. Med. 10, 55–63 (2004).
Fernandez, A. et al. The WNT receptor FZD7 is required for maintenance of the pluripotent state in human embryonic stem cells. Proc. Natl. Acad. Sci. USA 111, 1409–1414 (2014).
Lie, D.C. et al. Wnt signalling regulates adult hippocampal neurogenesis. Nature 437, 1370–1375 (2005).
Hirabayashi, Y. et al. The Wnt/β-catenin pathway directs neuronal differentiation of cortical neural precursor cells. Development 131, 2791–2801 (2004).
Blauwkamp, T.A., Nigam, S., Ardehali, R., Weissman, I.L. & Nusse, R. Endogenous Wnt signalling in human embryonic stem cells generates an equilibrium of distinct lineage-specified progenitors. Nat. Commun. 3, 1070 (2012).
Sheng, R. et al. Cholesterol selectively activates canonical Wnt signalling over non-canonical Wnt signalling. Nat. Commun. 5, 4393 (2014).
Sheng, R. et al. Cholesterol modulates cell signaling and protein networking by specifically interacting with PDZ domain-containing scaffold proteins. Nat. Commun. 3, 1249 (2012).
Munji, R.N., Choe, Y., Li, G., Siegenthaler, J.A. & Pleasure, S.J. Wnt signaling regulates neuronal differentiation of cortical intermediate progenitors. J. Neurosci. 31, 1676–1687 (2011).
Chenn, A. & Walsh, C.A. Regulation of cerebral cortical size by control of cell cycle exit in neural precursors. Science 297, 365–369 (2002).
Mutch, C.A., Schulte, J.D., Olson, E. & Chenn, A. β-catenin signaling negatively regulates intermediate progenitor population numbers in the developing cortex. PLoS ONE 5, e12376 (2010).
Inestrosa, N.C. & Arenas, E. Emerging roles of Wnts in the adult nervous system. Nat. Rev. Neurosci. 11, 77–86 (2010).
Sowers, L.P. et al. Disruption of the non-canonical Wnt gene PRICKLE2 leads to autism-like behaviors with evidence for hippocampal synaptic dysfunction. Mol. Psychiatry 18, 1077–1089 (2013).
Mohn, J.L. et al. Adenomatous polyposis coli protein deletion leads to cognitive and autism-like disabilities. Mol. Psychiatry 19, 1133–1142 (2014).
Mermelstein, C.S., Portilho, D.M., Mendes, F.A., Costa, M.L. & Abreu, J.G. Wnt/β-catenin pathway activation and myogenic differentiation are induced by cholesterol depletion. Differentiation 75, 184–192 (2007).
Yamaguchi, T.P., Bradley, A., McMahon, A.P. & Jones, S. A Wnt5a pathway underlies outgrowth of multiple structures in the vertebrate embryo. Development 126, 1211–1223 (1999).
Amar, M.J. et al. 5A apolipoprotein mimetic peptide promotes cholesterol efflux and reduces atherosclerosis in mice. J. Pharmacol. Exp. Ther. 334, 634–641 (2010).
Lowry, W.E. et al. Generation of human induced pluripotent stem cells from dermal fibroblasts. Proc. Natl. Acad. Sci. USA 105, 2883–2888 (2008).
Krakowiak, P.A. et al. Mutation analysis and description of sixteen RSH/Smith-Lemli-Opitz syndrome patients: polymerase chain reaction-based assays to simplify genotyping. Am. J. Med. Genet. 94, 214–227 (2000).
Davisson, M.T. & Akeson, E.C. An improved method for preparing G-banded chromosomes from mouse peripheral blood. Cytogenet. Cell Genet. 45, 70–74 (1987).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Xue, W. et al. CRISPR-mediated direct mutation of cancer genes in the mouse liver. Nature 514, 380–384 (2014).
Stahelin, R.V. & Cho, W. Differential roles of ionic, aliphatic, and aromatic residues in membrane-protein interactions: a surface plasmon resonance study on phospholipases A2. Biochemistry 40, 4672–4678 (2001).
Stahelin, R.V. & Cho, W. Roles of calcium ions in the membrane binding of C2 domains. Biochem. J. 359, 679–685 (2001).
Ananthanarayanan, B., Stahelin, R.V., Digman, M.A. & Cho, W. Activation mechanisms of conventional protein kinase C isoforms are determined by the ligand affinity and conformational flexibility of their C1 domains. J. Biol. Chem. 278, 46886–46894 (2003).
Cho, W., Bittova, L. & Stahelin, R.V. Membrane binding assays for peripheral proteins. Anal. Biochem. 296, 153–161 (2001).
Hsieh, J.C. et al. Mesd encodes an LRP5/6 chaperone essential for specification of mouse embryonic polarity. Cell 112, 355–367 (2003).
Koyama-Honda, I. et al. Fluorescence imaging for monitoring the colocalization of two single molecules in living cells. Biophys. J. 88, 2126–2136 (2005).
Zidovetzki, R. & Levitan, I. Use of cyclodextrins to manipulate plasma membrane cholesterol content: evidence, misconceptions and control strategies. Biochim. Biophys. Acta 1768, 1311–1324 (2007).
Wassif, C.A. et al. Biochemical, phenotypic and neurophysiological characterization of a genetic mouse model of RSH/Smith-Lemli-Opitz syndrome. Hum. Mol. Genet. 10, 555–564 (2001).
Ahn, S. & Joyner, A.L. In vivo analysis of quiescent adult neural stem cells responding to Sonic hedgehog. Nature 437, 894–897 (2005).
Acknowledgements
This work was supported by the intramural research programs of the Eunice Kennedy Shriver National Institute of Child Health and Human Development (NICHD) and the National Human Genome Research Institute (NHGRI), a pilot award for iPSC research from the NIH Center for Regenerative Medicine, and NIH grant GM110128 (W.C.). We thank K. Perez, N. Khezri and G.F. Rodriguez for assistance with hESC and iPSC culture. We also thank C. Toth and A. Ely for assistance with mice and sequencing. We thank A. Dutra, E. Pak and the NHGRI Cytogenetics Core for assistance with hESC and iPSC karyotyping. We thank L. (Chip) Dye and the NICHD Microscopy and Imaging Core for electron microscopy assistance. We thank B.S. Mallon (National Institute of Neurological Disorders and Stroke, NIH Stem Cell Unit), S. Kuznetsov (National Institute of Dental and Craniofacial Research, NIDCR), and P.G. Robey (NIDCR, NIH Stem Cell Unit) for experimental advice and the i19 and i21 human iPSC lines. We thank G. Mostoslavksy (Boston University) for the kind gift of the STEMCCA plasmid. We thank D. Kotton (Boston University) for the kind gift of the Cre-IRES-PuroR plasmid. We thank F. Zhang (Massachusetts Institute of Technology) for the gift of the pX330-U6-Chimeric_BB-CBh-hSpCas9 plasmid. We thank J. Mills (Children's Hospital of Philadelphia) for experimental advice, the National Heart, Lung and Blood Institute iPSC Core Facility for assistance with lathosterolosis iPSC generation, and M. Rao and K. Pfeifer for critical discussions. Finally, we would like to thank the patients and guardians who participated in the NIH clinical program and contributed to this study.
Author information
Authors and Affiliations
Contributions
K.R.F. and F.D.P. designed and directed the study, analyzed data and wrote the manuscript. K.R.F., A.N.T., P.E.O'H., N.M., I.M.W. and C.V.C. performed experiments and analyzed data. H.W. contributed to study design. C.A.W. provided assistance with GC/MS and study design. N.S.T. and W.J.P. provided support and performed statistical analysis of microarray experiments. Y.X. and W.C. designed, carried out and analyzed experiments on lipid-protein interactions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 1–3 (PDF 5069 kb)
Supplementary Table 4
Gene network analysis of microarray data identify highly correlated signaling pathways associated with sterol perturbation and pluripotent signaling. (XLSX 86 kb)
41591_2016_BFnm4067_MOESM50_ESM.mov
Beating areas generated from the CWI 4F-2 SLOS iPS line. Areas of cardiomyocyte formation and rhythmic beating were observed during germ layer differentiation assays. Beating area corresponds to immunocytochemical staining positive for NKX2.5 (immature cardiomyocyte marker) in Supplementary Figure 1h. (MOV 393 kb)
Rights and permissions
About this article
Cite this article
Francis, K., Ton, A., Xin, Y. et al. Modeling Smith-Lemli-Opitz syndrome with induced pluripotent stem cells reveals a causal role for Wnt/β-catenin defects in neuronal cholesterol synthesis phenotypes. Nat Med 22, 388–396 (2016). https://doi.org/10.1038/nm.4067
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.4067
This article is cited by
-
7-Dehydrocholesterol is an endogenous suppressor of ferroptosis
Nature (2024)
-
Integrated multi-dimensional analysis highlights DHCR7 mutations involving in cholesterol biosynthesis and contributing therapy of gastric cancer
Journal of Experimental & Clinical Cancer Research (2023)
-
Nanoscale fluorescence imaging of biological ultrastructure via molecular anchoring and physical expansion
Nano Convergence (2022)
-
Wnt signaling mediates oncogenic synergy between Akt and Dlx5 in T-cell lymphomagenesis by enhancing cholesterol synthesis
Scientific Reports (2020)
-
NIPBL+/− haploinsufficiency reveals a constellation of transcriptome disruptions in the pluripotent and cardiac states
Scientific Reports (2018)