Abstract
The acquisition of a protective vertebrate immune system hinges on the efficient generation of a diverse but self-tolerant repertoire of T cells by the thymus through mechanisms that remain incompletely resolved. Here we identified the endosomal-sorting-complex-required-for-transport (ESCRT) protein CHMP5, known to be required for the formation of multivesicular bodies, as a key sensor of thresholds for signaling via the T cell antigen receptor (TCR) that was essential for T cell development. CHMP5 enabled positive selection by promoting post-selection thymocyte survival in part through stabilization of the pro-survival protein Bcl-2. Accordingly, loss of CHMP5 in thymocyte precursor cells abolished T cell development, a phenotype that was 'rescued' by genetic deletion of the pro-apoptotic protein Bim or transgenic expression of Bcl-2. Mechanistically, positive selection resulted in the stabilization of CHMP5 by inducing its interaction with the deubiquitinase USP8. Our results thus identify CHMP5 as an essential component of the post-translational machinery required for T cell development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Starr, T.K., Jameson, S.C. & Hogquist, K.A. Positive and negative selection of T cells. Annu. Rev. Immunol. 21, 139–176 (2003).
Singer, A., Adoro, S. & Park, J.H. Lineage fate and intense debate: myths, models and mechanisms of CD4- versus CD8-lineage choice. Nat. Rev. Immunol. 8, 788–801 (2008).
Bonifacino, J.S. et al. Novel post-translational regulation of TCR expression in CD4+CD8+ thymocytes influenced by CD4. Nature 344, 247–251 (1990).
Grimm, L.M., Goldberg, A.L., Poirier, G.G., Schwartz, L.M. & Osborne, B.A. Proteasomes play an essential role in thymocyte apoptosis. EMBO J. 15, 3835–3844 (1996).
Yang, Y., Fang, S., Jensen, J.P., Weissman, A.M. & Ashwell, J.D. Ubiquitin protein ligase activity of IAPs and their degradation in proteasomes in response to apoptotic stimuli. Science 288, 874–877 (2000).
Maseda, D., Meister, S., Neubert, K., Herrmann, M. & Voll, R.E. Proteasome inhibition drastically but reversibly impairs murine lymphocyte development. Cell Death Differ. 15, 600–612 (2008).
Rusten, T.E., Vaccari, T. & Stenmark, H. Shaping development with ESCRTs. Nat. Cell Biol. 14, 38–45 (2011).
Merrill, S.A. & Hanson, P.I. Activation of human VPS4A by ESCRT-III proteins reveals ability of substrates to relieve enzyme autoinhibition. J. Biol. Chem. 285, 35428–35438 (2010).
Shim, S., Merrill, S.A. & Hanson, P.I. Novel interactions of ESCRT-III with LIP5 and VPS4 and their implications for ESCRT-III disassembly. Mol. Biol. Cell 19, 2661–2672 (2008).
Kranz, A., Kinner, A. & Kölling, R. A family of small coiled-coil-forming proteins functioning at the late endosome in yeast. Mol. Biol. Cell 12, 711–723 (2001).
Shim, J.H. et al. CHMP5 is essential for late endosome function and down-regulation of receptor signaling during mouse embryogenesis. J. Cell Biol. 172, 1045–1056 (2006).
Greenblatt, M.B. et al. CHMP5 controls bone turnover rates by dampening NF-κB activity in osteoclasts. J. Exp. Med. 212, 1283–1301 (2015).
Chang, Y.C. et al. mdm2 and bax, downstream mediators of the p53 response, are degraded by the ubiquitin-proteasome pathway. Cell Growth Differ. 9, 79–84 (1998).
Nencioni, A. et al. Evidence for a protective role of Mcl-1 in proteasome inhibitor-induced apoptosis. Blood 105, 3255–3262 (2005).
van Ewijk, W., Shores, E.W. & Singer, A. Crosstalk in the mouse thymus. Immunol. Today 15, 214–217 (1994).
Bendelac, A., Matzinger, P., Seder, R.A., Paul, W.E. & Schwartz, R.H. Activation events during thymic selection. J. Exp. Med. 175, 731–742 (1992).
Yamashita, I., Nagata, T., Tada, T. & Nakayama, T. CD69 cell surface expression identifies developing thymocytes which audition for T cell antigen receptor-mediated positive selection. Int. Immunol. 5, 1139–1150 (1993).
Barnden, M.J., Allison, J., Heath, W.R. & Carbone, F.R. Defective TCR expression in transgenic mice constructed using cDNA-based α- and β-chain genes under the control of heterologous regulatory elements. Immunol. Cell Biol. 76, 34–40 (1998).
Matusek, T. et al. The ESCRT machinery regulates the secretion and long-range activity of Hedgehog. Nature 516, 99–103 (2014).
Azzam, H.S. et al. CD5 expression is developmentally regulated by T cell receptor (TCR) signals and TCR avidity. J. Exp. Med. 188, 2301–2311 (1998).
Ueno, T. et al. CCR7 signals are essential for cortex-medulla migration of developing thymocytes. J. Exp. Med. 200, 493–505 (2004).
Yu, Q. et al. Cytokine signal transduction is suppressed in preselection double-positive thymocytes and restored by positive selection. J. Exp. Med. 203, 165–175 (2006).
Fu, G. et al. Themis sets the signal threshold for positive and negative selection in T-cell development. Nature 504, 441–445 (2013).
Hogquist, K.A. et al. T cell receptor antagonist peptides induce positive selection. Cell 76, 17–27 (1994).
Daniels, M.A. et al. Thymic selection threshold defined by compartmentalization of Ras/MAPK signalling. Nature 444, 724–729 (2006).
Mingueneau, M. et al. The transcriptional landscape of αβ T cell differentiation. Nat. Immunol. 14, 619–632 (2013).
Carpenter, A.C. & Bosselut, R. Decision checkpoints in the thymus. Nat. Immunol. 11, 666–673 (2010).
Gerondakis, S., Fulford, T.S., Messina, N.L. & Grumont, R.J. NF-κB control of T cell development. Nat. Immunol. 15, 15–25 (2014).
Xing, Y., Wang, X., Jameson, S.C. & Hogquist, K.A. Late stages of T cell maturation in the thymus involve NF-κB and tonic type I interferon signaling. Nat. Immunol. 17, 565–573 (2016).
Hurley, J.H. & Emr, S.D. The ESCRT complexes: structure and mechanism of a membrane-trafficking network. Annu. Rev. Biophys. Biomol. Struct. 35, 277–298 (2006).
Azmi, I.F. et al. ESCRT-III family members stimulate Vps4 ATPase activity directly or via Vta1. Dev. Cell 14, 50–61 (2008).
Rue, S.M., Mattei, S., Saksena, S. & Emr, S.D. Novel Ist1-Did2 complex functions at a late step in multivesicular body sorting. Mol. Biol. Cell 19, 475–484 (2008).
Gallo, E.M. et al. Calcineurin sets the bandwidth for discrimination of signals during thymocyte development. Nature 450, 731–735 (2007).
Staton, T.L. et al. Dampening of death pathways by schnurri-2 is essential for T-cell development. Nature 472, 105–109 (2011).
Czabotar, P.E., Lessene, G., Strasser, A. & Adams, J.M. Control of apoptosis by the BCL-2 protein family: implications for physiology and therapy. Nat. Rev. Mol. Cell Biol. 15, 49–63 (2014).
Azad, N. et al. Role of oxidative/nitrosative stress-mediated Bcl-2 regulation in apoptosis and malignant transformation. Ann. NY Acad. Sci. 1203, 1–6 (2010).
Luanpitpong, S. et al. Regulation of apoptosis by Bcl-2 cysteine oxidation in human lung epithelial cells. Mol. Biol. Cell 24, 858–869 (2013).
Holmström, K.M. & Finkel, T. Cellular mechanisms and physiological consequences of redox-dependent signalling. Nat. Rev. Mol. Cell Biol. 15, 411–421 (2014).
Bouillet, P. et al. BH3-only Bcl-2 family member Bim is required for apoptosis of autoreactive thymocytes. Nature 415, 922–926 (2002).
Pannu, J. et al. Ubiquitin specific protease 21 is dispensable for normal development, hematopoiesis and lymphocyte differentiation. PLoS One 10, e0117304 (2015).
Dufner, A. et al. The ubiquitin-specific protease USP8 is critical for the development and homeostasis of T cells. Nat. Immunol. 16, 950–960 (2015).
Devadas, S., Zaritskaya, L., Rhee, S.G., Oberley, L. & Williams, M.S. Discrete generation of superoxide and hydrogen peroxide by T cell receptor stimulation: selective regulation of mitogen-activated protein kinase activation and fas ligand expression. J. Exp. Med. 195, 59–70 (2002).
Jin, R. et al. Trx1/TrxR1 system regulates post-selected DP thymocytes survival by modulating ASK1-JNK/p38 MAPK activities. Immunol. Cell Biol. 93, 744–752 (2015).
Choudhuri, K. et al. Polarized release of T-cell-receptor-enriched microvesicles at the immunological synapse. Nature 507, 118–123 (2014).
Wi, S.M., Min, Y. & Lee, K.Y. Charged MVB protein 5 is involved in T-cell receptor signaling. Exp. Mol. Med. 48, e206 (2016).
Raiborg, C. & Stenmark, H. The ESCRT machinery in endosomal sorting of ubiquitylated membrane proteins. Nature 458, 445–452 (2009).
Khoury, C.M., Yang, Z., Ismail, S. & Greenwood, M.T. Characterization of a novel alternatively spliced human transcript encoding an N-terminally truncated Vps24 protein that suppresses the effects of Bax in an ESCRT independent manner in yeast. Gene 391, 233–241 (2007).
Mu, R. et al. Two distinct binding modes define the interaction of Brox with the C-terminal tails of CHMP5 and CHMP4B. Structure 20, 887–898 (2012).
Shahmoradgoli, M. et al. Antiapoptotic function of charged multivesicular body protein 5: a potentially relevant gene in acute myeloid leukemia. Int. J. Cancer 128, 2865–2871 (2011).
Lee, P.P. et al. A critical role for Dnmt1 and DNA methylation in T cell development, function, and survival. Immunity 15, 763–774 (2001).
Sentman, C.L., Shutter, J.R., Hockenbery, D., Kanagawa, O. & Korsmeyer, S.J. bcl-2 inhibits multiple forms of apoptosis but not negative selection in thymocytes. Cell 67, 879–888 (1991).
Cibotti, R., Punt, J.A., Dash, K.S., Sharrow, S.O. & Singer, A. Surface molecules that drive T cell development in vitro in the absence of thymic epithelium and in the absence of lineage-specific signals. Immunity 6, 245–255 (1997).
Ohoka, Y. et al. In vitro differentiation and commitment of CD4+ CD8+ thymocytes to the CD4 lineage, without TCR engagement. Int. Immunol. 8, 297–306 (1996).
Rybakin, V. & Gascoigne, N.R. Negative selection assay based on stimulation of T cell receptor transgenic thymocytes with peptide-MHC tetramers. PLoS One 7, e43191 (2012).
Finetti, F. et al. Specific recycling receptors are targeted to the immune synapse by the intraflagellar transport system. J. Cell Sci. 127, 1924–1937 (2014).
Myers, M.D., Dragone, L.L. & Weiss, A. Src-like adaptor protein down-regulates T cell receptor (TCR)-CD3 expression by targeting TCRζ for degradation. J. Cell Biol. 170, 285–294 (2005).
Breitschopf, K., Haendeler, J., Malchow, P., Zeiher, A.M. & Dimmeler, S. Posttranslational modification of Bcl-2 facilitates its proteasome-dependent degradation: molecular characterization of the involved signaling pathway. Mol. Cell. Biol. 20, 1886–1896 (2000).
Nelson, K.J. et al. Use of dimedone-based chemical probes for sulfenic acid detection methods to visualize and identify labeled proteins. Methods Enzymol. 473, 95–115 (2010).
Villén, J. & Gygi, S.P. The SCX/IMAC enrichment approach for global phosphorylation analysis by mass spectrometry. Nat. Protoc. 3, 1630–1638 (2008).
Eng, J.K., McCormack, A.L. & Yates, J.R. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Acknowledgements
We thank A. Singer (US National Institutes of Health) for Bcl2-transgenic mice; S. Dimmeler (J.W. Goethe University, Frankfurt, Germany) for Bcl-2-encoding plasmids; J. Kaplan (University of Utah) for anti-LIP5; M. Greenblatt for critical reading of the manuscript; J. McCormick (Weill Cornell Medicine) for sorting by flow cytometry; L. Cohen-Gould, J. Cohen and J. Jimenez for histology and electron microscopy; and the NIH Tetramer Core Facility (Emory University) for tetramers. Supported by the US National Institutes of Health (R01AR068983 to J.H.S., and R01CA112663 to L.H.G.).
Author information
Authors and Affiliations
Contributions
S.A. designed study, performed experiments, analyzed data and wrote the manuscript; K.H.P., S.E.B., R.L. and H.R.S. assisted with experiments; H.S. and J.H.K. performed bioinformatics; K.-P.K. provided mice with loxP-flanked Usp8 alleles; J.-H.S. designed the study, performed experiments, analyzed data and wrote the manuscript; and L.H.G. designed the study, analyzed data, provided supervision and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
L.H.G. is on the board of directors of and holds equity in Bristol Myers Squibb. L.H.G. is founder of and S.E.B. is a co-founder of Quentis Therapeutics, and S.E.B. and H.R.S. are employed by Quentis Therapeutics.
Integrated supplementary information
Supplementary Figure 1 CHMP5 expression in thymocytes.
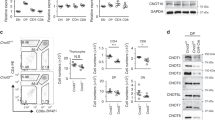
(a) Validation of rabbit anti-CHMP5 antibody in Jurkat T-cells depleted of CHMP5 with two independent shRNA and LacZ control shRNA. (b) Immunoblot of MG132-treated wild-type thymocytes. Protein band intensities relative to untreated samples are indicated. (c) Immunoblot for ESCRT proteins with relative band intensities. (d) Chmp5 mRNA in preselection DP and intermediate (Int) thymocytes from WT, MHCII-selection (B2m–/–) and MHCI-selection (H2Ab1–/–) mice. Error bar, s.d. (e) Chmp5 mRNA levels in the thymocyte populations sorted from WT and CKO mice revealing Chmp5 deletion. Error bar, s.d. (f) mRNA expression of selected ESCRT genes in WT and CKO mice relative to Actin. Each circle represents one mouse. Error bar, s.d. DN, double negative; DP, double positive; Int, intermediate thymocytes; SP, single positive. Data are representative of at least three independent experiments.
Supplementary Figure 2 CHMP5 is essential for thymocyte positive selection.
(a) Representative flow cytometry plot of CD24 versus TCRb expression on CD4 and CD8 single positive (SP) thymocytes. Numbers indicate the proportion of cells within each gate. (b) Immunoblot of CHMP5 protein in residual peripheral CD4 and CD8 T cells from WT and CKO mice. (c) CD44 expression on peripheral CD4 and CD8 T cells. (d,e) Characterization of WT and CKO mice expressing the OT-II TCR transgene. Representative flow cytometry plot of CD4 versus CD8 (top) and CD24 versus Vβ5 thymocytes (bottom) on total thymocytes (d) and Vβ5 expression on splenocytes (e) are shown. Numbers indicate the proportion of cells in each gate. Data are representative of two (b) and five independent experiments (a,c-e).
Supplementary Figure 3 Cell-intrinsic role for CHMP5 in T cell generation.
(a) Mixed bone marrow chimera strategy. (b) Lentiviral vector constructs and strategy for CHMP5 transgenesis rescue experiment. (c, d) Representative flow cytometer plot of thymocyte CD24 versus TCRβ expression (c) and splenic TCRβ versus CD19 expression (d) on GFP– and GFP+ cells within CD45.2+ donor-derived cells from mice reconstituted with control (LV-Ctrl) and CHMP5-transduced (LV-C5) bone marrow cells. Dotted line, untransduced WT thymocytes. Numbers indicate cell proportion within each gate. Data are representative of three independent experiments.
Supplementary Figure 4 Characterization of TCR signaling in CHMP5-deficient thymocytes.
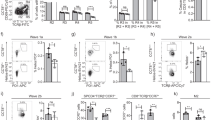
(a) Flow cytometer phenotypic definition and sort gating of intermediate thymocytes. (b) Surface expression of positive selection markers IL-7Rα and CCR7 on preselection DP and intermediate (Int) thymocytes from WT and CKO mice. (c) Proportion of CD69+ thymocytes after overnight (18 hours) stimulation with anti-CD3ɛ alone (top) or with anti-CD2 antibodies (bottom). Error bar, s.e.m., n = 4 mice each. Data are representative of three independent experiments.
Supplementary Figure 5 Expression of thymocyte positive selection genes in wild-type and CKO thymocytes.
(a) Gata3 mRNA (top) and Gata3 protein (bottom) expression. Note immunoblot for Grb2 protein as loading control. Western blot is representative of two independent experiments. (b) mRNA expression of TCR-regulated genes: Nfatc1, Egr1 and Tox. (c) mRNA levels of CD4 and CD8 lineage-specification genes Zbtb7b (Th-POK) and Runx3. (d) Relative NF-κB activity assessed by p65 binding to consensus NF-κB oligonucleotides determined by ELISA. Error bar, s.e.m; four mice each. P values are shown on graph, Students t-Test. Representative of two experiments. (e) NF-κB target gene expression. DP, preselection CD4+CD8+ thymocytes; Int, intermediate thymocytes; CD4SP, CD4 single-positive thymocytes; CD8SP, single-positive thymocytes. All mRNA levels were normalized to Actin. For qPCR data, each circle represents an average of duplicate of individual mice.
Supplementary Figure 6 Regulation of Bcl-2 protein by CHMP5.
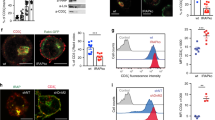
(a) Relative mRNA levels of Bcl-xl, Bim, Nur77 and Bid. (b) Immunoblot of Bcl-2 and Bim proteins in sorted thymocyte subsets. (c, d) Representative histogram (c) and bar graph (d) of mean fluorescence intensity (MFI) of intracellular Bcl-2 protein in gated thymocyte subsets. Error bar, s.e.m; four mice each. Representative of three independent experiments. (e) Intracellular Bcl-2 MFI in intermediate OT-II TCR transgenic thymocytes. Each circle represents one mouse. (f,g) Intracellular Bcl-2 (f) and phosphorylated STAT5 (g) in IL-7-stimulated intermediate thymocytes. Error bar, s.e.m.; four mice each. (h) Intracellular Bcl-2 expression in WT and CKO intermediate thymocytes after overnight (18 hours) culture in cycloheximide (CHX) containing medium. Error bar, s.e.m.; three mice each. (i,j) Co-immunoprecipitation of epitope-tagged CHMP5 and BCL-2 in HEK293 cells (j) and in cell-free assays with GST-tagged BCL-2 and His-tagged CHMP5 (j). Numbers indicate band intensity relative to “GST” lane. Representative of two independent experiments. (k) Total thymocyte ROS levels determined by CellRox Deep Red staining. Error bar, s.e.m.; four mice each. *, P < 0.05; **, P < 0.001; ***, P < 0.0001, Student’s t-tests.
Supplementary Figure 7 Genetic augmentation of prosurvival pathway restores T cell development in CKO mice.
Representative flow cytometry plot of CCR7 versus TCRβ expression and corresponding thymocyte numbers in WT and CKO mice deficient for Bim (a,b) or expressing transgenic Bcl-2 (c,d). Numbers in plots indicate proportion of cells within gates. Each data point circle represents one mouse. Fold-times difference between CKO groups are indicated. **, P < 0.001; ***, P < 0.0001, Student’s t-tests.
Supplementary Figure 8 Differential regulation of CHMP5 by TCR signaling.
(a) Immunoblot (top) and qPCR (bottom) analyses of CHMP5 expression in stimulated pre-selection (CD69–) DP thymocytes. Relative band intensities (normalized to unstimulated (medium) control) are shown. (b,c) Chmp5 mRNA levels (normalized to Actin) in PMA/ionomycin stimulated pre-selection (CD69–) DP thymocytes (b) or kb-peptide tetramer-stimulated OT-I.B2m–/– thymocytes (c). Error bar, s.d. (d) Anti-HA (CHMP5) immunoblot of Nickel-immunoprecipitates of PMA/ionomycin-treated HEK293 cells co-transfected with HA-tagged CHMP5 and His-tagged ubiquitin. (e) Ubiquitin immunoblot of anti-HA immunoprecipitates showing inhibition of Bcl-2 ubiquitination by the antioxidant NAC in PMA+ionomycin-treated HEK293 cells expressing HA-tagged Bcl-2. (f) Ubiquitin blot of anti-CHMP5 immunoprecipitates from unstimulated (none) or Jurkat T-cells activated by CD3e plus CD28 antibody crosslinking or PMA plus ionomycin. Data are representative of two (d–f) and three (a–c) independent experiments.
Supplementary Figure 9 Interaction between CHMP5 and USP8 in thymocytes that have undergone TCR selection.
(a) USP8 expression in thymocyte subsets from WT and CKO mice. (b) Immunoblot of CHMP5 and USP8 in isotype and anti-CHMP5 antibody immunoprecipitates from pre-selection CD4+CD8+ (DP) and intermediate (Int) thymocytes. (c) Anti-USP8 immunoprecipitation to determine the effect of PMA plus ionomycin dose on the interaction between CHMP5 and USP8 in DP thymocytes. *, non-specific band. (d) Usp8 and Bcl2 mRNA levels (normalized to Actin) in USP8-deficient (Usp8CD4cre) or littermate control (Usp8f/f) intermediate thymocytes. All data in a–d are representative of two independent experiments each. (e) Summarized hypothetical model of CHMP5-USP8 axis during thymocyte development.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Table 1.
Rights and permissions
About this article
Cite this article
Adoro, S., Park, K., Bettigole, S. et al. Post-translational control of T cell development by the ESCRT protein CHMP5. Nat Immunol 18, 780–790 (2017). https://doi.org/10.1038/ni.3764
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3764
This article is cited by
-
PSMC6 induces immune cell infiltration and inflammatory response to aggravate primary Sjögren’s syndrome
Journal of Human Genetics (2023)
-
Phospholipase D activation is required for 1-aminocyclopropane 1-carboxylic acid signaling during sexual reproduction in the marine red alga Neopyropia yezoensis (Rhodophyta)
BMC Plant Biology (2022)
-
Molecular basis of ubiquitin-specific protease 8 autoinhibition by the WW-like domain
Communications Biology (2021)
-
Transcriptomic analysis under ethylene precursor treatment uncovers the regulation of gene expression linked to sexual reproduction in the dioecious red alga Pyropia pseudolinearis
Journal of Applied Phycology (2019)
-
The ESCRT protein CHMP5 escorts αβ T cells through positive selection
Cellular & Molecular Immunology (2018)