Abstract
Rapid activation of memory CD4+ T helper 2 (TH2) cells during allergic inflammation requires their recruitment into the affected tissue. Here we demonstrate that group 2 innate lymphoid (ILC2) cells have a crucial role in memory TH2 cell responses, with targeted depletion of ILC2 cells profoundly impairing TH2 cell localization to the lungs and skin of sensitized mice after allergen re-challenge. ILC2-derived interleukin 13 (IL-13) is critical for eliciting production of the TH2 cell–attracting chemokine CCL17 by IRF4+CD11b+CD103− dendritic cells (DCs). Consequently, the sentinel function of DCs is contingent on ILC2 cells for the generation of an efficient memory TH2 cell response. These results elucidate a key innate mechanism in the regulation of the immune memory response to allergens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Kim, H.Y., DeKruyff, R.H. & Umetsu, D.T. The many paths to asthma: phenotype shaped by innate and adaptive immunity. Nat. Immunol. 11, 577–584 (2010).
Lambrecht, B.N. & Hammad, H. The immunology of asthma. Nat. Immunol. 16, 45–56 (2015).
Islam, S.A. & Luster, A.D. T cell homing to epithelial barriers in allergic disease. Nat. Med. 18, 705–715 (2012).
McKenzie, A.N., Spits, H. & Eberl, G. Innate lymphoid cells in inflammation and immunity. Immunity 41, 366–374 (2014).
Moro, K. et al. Innate production of T(H)2 cytokines by adipose tissue-associated c-Kit(+)Sca-1(+) lymphoid cells. Nature 463, 540–544 (2010).
Neill, D.R. et al. Nuocytes represent a new innate effector leukocyte that mediates type-2 immunity. Nature 464, 1367–1370 (2010).
Price, A.E. et al. Systemically dispersed innate IL-13-expressing cells in type 2 immunity. Proc. Natl. Acad. Sci. USA 107, 11489–11494 (2010).
Chang, Y.J. et al. Innate lymphoid cells mediate influenza-induced airway hyper-reactivity independently of adaptive immunity. Nat. Immunol. 12, 631–638 (2011).
Halim, T.Y., Krauss, R.H., Sun, A.C. & Takei, F. Lung natural helper cells are a critical source of Th2 cell-type cytokines in protease allergen-induced airway inflammation. Immunity 36, 451–463 (2012).
Monticelli, L.A. et al. Innate lymphoid cells promote lung-tissue homeostasis after infection with influenza virus. Nat. Immunol. 12, 1045–1054 (2011).
Drake, L.Y., Iijima, K. & Kita, H. Group 2 innate lymphoid cells and CD4+ T cells cooperate to mediate type 2 immune response in mice. Allergy 69, 1300–1307 (2014).
Gold, M.J. et al. Group 2 innate lymphoid cells facilitate sensitization to local, but not systemic, TH2-inducing allergen exposures. J. Allergy Clin. Immunol. 133, 1142–1148 (2014).
Halim, T.Y. et al. Group 2 innate lymphoid cells are critical for the initiation of adaptive T helper 2 cell-mediated allergic lung inflammation. Immunity 40, 425–435 (2014).
Mirchandani, A.S. et al. Type 2 innate lymphoid cells drive CD4+ Th2 cell responses. J. Immunol. 192, 2442–2448 (2014).
Oliphant, C.J. et al. MHCII-mediated dialog between group 2 innate lymphoid cells and CD4(+) T cells potentiates type 2 immunity and promotes parasitic helminth expulsion. Immunity 41, 283–295 (2014).
van Rijt, L.S. et al. In vivo depletion of lung CD11c+ dendritic cells during allergen challenge abrogates the characteristic features of asthma. J. Exp. Med. 201, 981–991 (2005).
Crapster-Pregont, M., Yeo, J., Sanchez, R.L. & Kuperman, D.A. Dendritic cells and alveolar macrophages mediate IL-13-induced airway inflammation and chemokine production. J. Allergy Clin. Immunol. 129, 1621–1627.e3 (2012).
Mikhak, Z. et al. Contribution of CCR4 and CCR8 to antigen-specific TH2 cell trafficking in allergic pulmonary inflammation. J. Allergy Clin. Immunol. 123, 67–73.e3 (2009).
Imai, T. et al. Selective recruitment of CCR4-bearing Th2 cells toward antigen-presenting cells by the CC chemokines thymus and activation-regulated chemokine and macrophage-derived chemokine. Int. Immunol. 11, 81–88 (1999).
Novey, H.S., Marchioli, L.E., Sokol, W.N. & Wells, I.D. Papain-induced asthma–physiological and immunological features. J. Allergy Clin. Immunol. 63, 98–103 (1979).
Mottram, J.C., Helms, M.J., Coombs, G.H. & Sajid, M. Clan CD cysteine peptidases of parasitic protozoa. Trends Parasitol. 19, 182–187 (2003).
Kamijo, S. et al. IL-33-mediated innate response and adaptive immune cells contribute to maximum responses of protease allergen-induced allergic airway inflammation. J. Immunol. 190, 4489–4499 (2013).
Zheng, W. & Flavell, R.A. The transcription factor GATA-3 is necessary and sufficient for Th2 cytokine gene expression in CD4 T cells. Cell 89, 587–596 (1997).
Moon, J.J. et al. Naive CD4+ T cell frequency varies for different epitopes and predicts repertoire diversity and response magnitude. Immunity 27, 203–213 (2007).
Medoff, B.D. et al. CD11b+ myeloid cells are the key mediators of Th2 cell homing into the airway in allergic inflammation. J. Immunol. 182, 623–635 (2009).
Plantinga, M. et al. Conventional and monocyte-derived CD11b+ dendritic cells initiate and maintain T helper 2 cell–mediated immunity to house dust mite allergen. Immunity 38, 322–335 (2013).
Gao, Y. et al. Control of T helper 2 responses by transcription factor IRF4-dependent dendritic cells. Immunity 39, 722–732 (2013).
Williams, J.W. et al. Transcription factor IRF4 drives dendritic cells to promote Th2 differentiation. Nat. Commun. 4, 2990 (2013).
Halim, T.Y. et al. Retinoic-acid-receptor-related orphan nuclear receptor-α is required for natural helper cell development and allergic inflammation. Immunity 37, 463–474 (2012).
Wong, S.H. et al. Transcription factor RORα is critical for nuocyte development. Nat. Immunol. 13, 229–236 (2012).
Kim, B.S. et al. TSLP elicits IL-33-independent innate lymphoid cell responses to promote skin inflammation. Sci. Transl. Med. 5, 170ra116 (2013).
Roediger, B. et al. Cutaneous immunosurveillance and regulation of inflammation by group 2 innate lymphoid cells. Nat. Immunol. 14, 564–573 (2013).
Salimi, M. et al. A role for IL-25 and IL-33-driven type-2 innate lymphoid cells in atopic dermatitis. J. Exp. Med. 210, 2939–2950 (2013).
Maizels, R.M., Hewitson, J.P. & Smith, K.A. Susceptibility and immunity to helminth parasites. Curr. Opin. Immunol. 24, 459–466 (2012).
Kumar, R.K., Herbert, C. & Foster, P.S. The “classical” ovalbumin challenge model of asthma in mice. Curr. Drug Targets 9, 485–494 (2008).
Stephens, R. & Chaplin, D.D. IgE cross-linking or lipopolysaccharide treatment induces recruitment of Th2 cells to the lung in the absence of specific antigen. J. Immunol. 169, 5468–5476 (2002).
Hammad, H. et al. Inflammatory dendritic cells–not basophils–are necessary and sufficient for induction of Th2 immunity to inhaled house dust mite allergen. J. Exp. Med. 207, 2097–2111 (2010).
MacDonald, A.S., Straw, A.D., Dalton, N.M. & Pearce, E.J. Cutting edge: Th2 response induction by dendritic cells: a role for CD40. J. Immunol. 168, 537–540 (2002).
Ito, T. et al. TSLP-activated dendritic cells induce an inflammatory T helper type 2 cell response through OX40 ligand. J. Exp. Med. 202, 1213–1223 (2005).
Eiwegger, T. & Akdis, C.A. IL-33 links tissue cells, dendritic cells and Th2 cell development in a mouse model of asthma. Eur. J. Immunol. 41, 1535–1538 (2011).
Wills-Karp, M. Interleukin-13 in asthma pathogenesis. Immunol. Rev. 202, 175–190 (2004).
Fahy, J.V. Type 2 inflammation in asthma—present in most, absent in many. Nat. Rev. Immunol. 15, 57–65 (2015).
Tittel, A.P. et al. Functionally relevant neutrophilia in CD11c diphtheria toxin receptor transgenic mice. Nat. Methods 9, 385–390 (2012).
Haymaker, C.L. et al. Bone marrow–derived IL-13Rα1-positive thymic progenitors are restricted to the myeloid lineage. J. Immunol. 188, 3208–3216 (2012).
Schlenner, S.M. et al. Fate mapping reveals separate origins of T cells and myeloid lineages in the thymus. Immunity 32, 426–436 (2010).
Heng, T.S. & Painter, M.W. Immunological Genome Project C. The Immunological Genome Project: networks of gene expression in immune cells. Nat. Immunol. 9, 1091–1094 (2008).
Acknowledgements
We thank the MRC LMB flow core for help with flow cytometry; H.R. Rodewald (Division of Cellular Immunology, German Cancer Research Center) for Il7RaCre mice; and D. Withers (MRC Centre for Immune Regulation, Institute for Biomedical Research, College of Medical and Dental Sciences, University of Birmingham) for advice with 2W1S peptide experiments. Supported by the UK Medical Research Council (UI05178805), the Wellcome Trust (100963) (A.N.J.M.), Science Foundation Ireland and National Children's Research Centre (P.G.F), A-STAR (Y.Y.H.), and the Canadian Institutes of Health Research (T.Y.F.H.). N.G. is a member of the DFG-funded ImmunoSensation Cluster of Excellence.
Author information
Authors and Affiliations
Contributions
T.Y.F.H. designed and performed experiments, and wrote the paper. Y.Y.H. and S.T.S. designed and performed experiments. H.Z. provided the IL13ra1−/− mice. N.G. provided the CD11c-LuciDTR and CD11c-DOG mice. P.G.F. provided the N.b. reagent. A.N.J.M. supervised the project, designed the experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
A.N.J.M. received support from Janssen.
Integrated supplementary information
Supplementary Figure 1 Time-course analysis of papain protease allergen–treated mice.
(a–b) Myeloid cells were identified in the bronchoalveolar lavage (BAL) and lung tissue by first gating for live (DAPI–), CD45+ leukocytes. Additional stains include CD11c, Siglec-F, Gr-1, 7/4 (Ly-6B), FcɛR1α, F4/80, DX5 (CD49b), and CD11b. Eosinophils were identified as CD11c–Siglec-F+ and by their FSC/SSC profile; neutrophils were identified as Siglec-F–Gr-1+7/4+ and by their FSC/SSC profile; alveolar macrophages were identified as Siglec-F+CD11c+F4/80+CD11b– and by their FSC/SSC profile; and dendritic cells were identified as Siglec-F–Gr-1–/loCD11c+F4/80–CD11b± (a). The FSC/SSC profiles of BAL eosinophils, neutrophils and alveolar macrophages are shown (b).
(c–d) Myeloid cells were identified by flow cytometry in the lungs of papain-injected mice at the indicated time points (see Fig. 1a). Total numbers of eosinophils (black), neutrophils (red) and macrophages (MΦ) (blue) were quantified (c). Total numbers of CD11b+ dendritic cells (green) CD11b– dendritic cells (black) and alveolar macrophages (blue) were quantified (d).
(e–f) TH2 cells and ILC2s were identified in the lungs and mediastinal LNs (mLNs) of papain-treated mice at the indicated time points (Fig. 1a) as described in Fig. 1c. Total numbers of ILC2s were quantified in the lungs (e). Total numbers of TH2 cells were identified in the mLN (red) and lung (blue) (f).
Data are representative of at least two independent experiments per group, each containing at least three animals. Mean values ± SEM are indicated in c–f.
Supplementary Figure 2 Time-course analysis of A. alternata–treated mice.
(a–e) Mice were sensitized intranasally with Alternaria alternata extract (10 μg) + 2W1S peptide (50 μg) on days 0 and 1, followed by a single intranasal re-challenge on day 13. Myeloid cells in the lung tissue were identified on the days indicated by flow cytometry (as described in Supplementary Fig. 1a). Total lung eosinophils (black) and neutrophils (red) were quantified (a), as were total lung dendritic cells (black) and macrophages (MΦ) (blue) (b). ILC2s and TH2 cells were identified (as described in Fig. 1c). Total lung IL-13+ ILC2s (c), tetramer– (d) and tetramer+ (e) TH2 cells were quantified.
Data are a compilation of two independent experiments, each containing at least three animals. Mean values ± SEM are indicated in a–e.
Supplementary Figure 3 Analysis of inflammation with different combinations of antigens.
(a–d) Mice were sensitized intranasally with 10 μg of papain (PAP) or heat-inactivated papain (HP) + 2W1S peptide (50 μg) on days 0 and 1, followed by a single intranasal re-challenge on day 132. Myeloid cells in the lung tissue were identified on day 135 by flow cytometry (as described in Supplementary Fig. 1a). Total BAL eosinophils (black) were quantified (b). TH2 cells were identified on day 135 (as described in Fig. 1c). Total lung tetramer– (c) and tetramer+ (d) TH2 cells were quantified.
(e–j) Mice were sensitized intranasally with 10 μg of papain (PAP) + 2W1S peptide (50 μg) on days 0 and 1. Sensitized mice were subsequently re-challenged intranasally with PAP, 2W1S peptide (2W), and Alternaria alternata extract (A.alt) (10 μg) combinations as indicated on the x-axes on day 15. Myeloid cells in the lung tissue were identified on day 18 by flow cytometry (as described in Supplementary Fig. 1a). Total eosinophils in the lung (f) and BAL (g) were quantified. ILC2s and TH2 cells were identified as described (Fig. 1c). Total lung tetramer+ (h), tetramer+ TH2 cells (i) cells, and lung ILC2s (j) were quantified.
Data are representative of two independent experiments, each containing at least five animals. Mean values ± SEM are indicated in b–d, f–j. p ≤ 0.05 =*, p ≤ 0.01 = **, p ≤ 0.001 = ***, p ≤ 0.0001 = ****.
Supplementary Figure 4 ILC2 cell–depletion strategies and experimental design in iCOS-T mice.
(a–d) Mice were sensitized intranasally with papain (10 μg) + 2W1S peptide (50 μg) on days 0 and 1, followed by a single intranasal re-challenge (or HP + 2W1S control as indicated in e) on day 15. Mice were administered via intraperitoneal injection with diphtheria toxin (DTx) or PBS control as indicated. Mice were sacrificed on days 16 (a) or 20 (b), and BAL, lung and mediastinal LNs (mLNs) were obtained for analysis. ILC2s were identified as described (Fig. 1c). On day 20, total lung ILC2s were quantified (c). Lung Tetramer–CD4+GATA3+ TH2 cells in the lungs of mice on day 20 were quantified (d).
(e–f) iCOS-T mice were sensitized intranasally with papain (10 μg) + 2W1S peptide (50 μg) on days 0 and 1, followed by a single intranasal re-challenge on day 132. Lung tetramer+ TH2 cells were identified by flow-cytometry as in Fig. 1c and quantified on day 135 (f). One group of iCOS-T mice was treated with DTx (grey bar).
(g–h) Lung cells from the previous experiment (e–f) were also analyzed for other ILC populations based on GATA3 and RORγt staining of Live CD45+B220–Lineage–CD127+ cells (g). ILC2s, ILC3s and non-ILC2/3s were further tested for intracellular IL-13 production. The total number of non-ILC2/3s in the lung of iCOS-T treated with PBS (black) or DTx (blue), and allergen (as indicated in e) (h).
(i) WT or iCOS-T mice were sensitized intranasally with Alternaria alternata extract (10 μg) + 2W1S-peptide (50 μg) on days 0 and 1, followed by a single intranasal re-challenge on day 15, or DTx administration, as indicated. Lung tetramer- TH2 cells were identified by flow cytometry as in Fig. 1c and quantified.
(j) Mice were intranasally sensitized on days 0 and 1, and re-challenged on day 15 with papain as indicated. All groups of animals were analyzed on day 16 for IL-4 production by intracellular staining in ILC2s or TH2 cells of the lung or mLN.
Data are representative of three independent experiments in c, and one experiment with groups containing 3-6 animals in f–h. Mean values ± SEM are indicated in c–d, f, h–j. ns = not significant, p ≤ 0.05 =*, p ≤ 0.01 = **, p ≤ 0.001 = ***, p ≤ 0.0001 = ****.
Supplementary Figure 5 CCL17 production depends on ILC2 cell activation in allergen-sensitized mice.
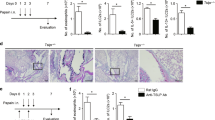
(a) Mice were sensitized intranasally with papain (10 μg) + 2W1S-peptide (50 μg) on days 0 and 1, followed by a single intranasal re-challenge on day 15. Mice were administered via intraperitoneal injection with 300 µg of anti-IL-13 mAb, anti-CCL17 mAb, or IgG isotype control in PBS as indicated. Mice were analyzed on day 16.
(b) WT or Il33–/– mice were injected intranasally with papain or control heat-inactivated papain (HP) on days 0 and 1, followed by analysis on day 2 for CCL17 concentration in the BAL.
(c) WT or iCOS-T mice were sensitized intranasally with Alternaria alternata extract (10 μg) + 2W1S peptide (50 μg) on days 0 and 1, followed by a single intranasal re-challenge on day 15 and DTx administration as indicated. Lung CD11b+CCL17+ DC were identified by flow cytometry as in Fig. 4a and quantified.
Data are representative of one experiment, with each group containing at least three animals. Mean values ± SEM are indicated in b–c. p ≤ 0.05 =*, p ≤ 0.01 = **.
Supplementary Figure 6 DC depletion and mixed bone-marrow chimera strategy.
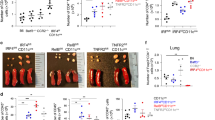
(a–e) CD11c-DTR or WT littermates were sensitized intranasally with papain (10 μg) + 2W1S peptide (50 μg) on days 0 and 1, followed by a single intranasal re-challenge on day 15. Mice were administered via intraperitoneal injection with diphtheria toxin (DTx) as indicated. Mice were sacrificed on day 16. CD11b± DCs (b) and macrophages (c) were identified by flow-cytometry (Supplementary Fig. 1a) and quantified in the lung. ILC2 and TH2 cells were identified as described (Fig. 1c). Total lung tetramer± TH2 cells were quantified (d). Total interferon-gamma positive CD4+ T cells were identified by intracellular staining and quantified in the lung (e).
(f) Mixed bone-marrow chimera mice were generated by reconstituting lethally irradiated WT recipients with 100% CD11c-LuciDTR, 50% CD11c-LuciDTR + 50% Il13ra1–/–, or 50% CD11c-LuciDTR + 50% WT bone marrow (100% = 107 whole bone marrow cells). Recipients were left for 24–32 weeks post transplant, which was required for efficient ILC2 reconstitution (unpublished results).
(g–h) Recipients from (f) were sensitized intranasally with papain (10 μg) + 2W1S peptide (50 μg) on days 0 and 1, followed by a single intranasal re-challenge on day 15. Animals were administered with DTx (500 ng) by intraperitoneal injection on days 13, 14 and 15 (g). Animals were analyzed on day 16 for total numbers of lung macrophages (h) (as described in Supplementary Fig. 1a).
Data are representative of two independent experiments, each containing at least three animals in b-e, and one experiment containing at least five animals per group in h. Mean values ± SEM are indicated in b–e, h. ns = not significant, p ≤ 0.05 =*, p ≤ 0.01 = **, p ≤ 0.001 = ***.
Supplementary Figure 7 Analysis of interactions among ILC2 cells, DCs and TH2 cells in the skin and gut of mice.
(a–b) WT, Il13−/− or Rora−/flIl7ra-Cre mice were injected with 10 μg of papain or HP (into WT mice) intradermally into each ear on days 0 and 1, followed by the isolation of skin cells on day 2. Cutaneous live CD45+B220–Lineage–CD127+GATA3+ ILC2s were detected by flow cytometry (a). Eosinophils were detected similar to described in Supplementary Fig. 1a and quantified in each ear of treated animals (b).
(c–f) iCOS-T or WT mice were sensitized intradermally in the left ear with papain (10 μg) on day 0 and 1 followed by re-challenge in the right ear on day 15 (c).
Right ears from WT mice, treated as indicated, were harvested on day 16. Skin TH2 cells (Live CD45+CD3+CD4+GATA3+Foxp3–) were quantified (d).
iCOS-T mice, treated as in c, were administered with DTx (blue) or PBS (black), followed by quantification of ILC2 (Live CD45+lineage−CD127+GATA3+) and TH2 cells in the right ears on day 16 after papain re-challenge (e). iCOS-T and WT mice were sensitized as in c, but treated with DTx and re-challenged intradermally as indicated. The total number of IL-13+ TH2 cells was quantified in the right ears (f).
Data are representative of two independent experiments, each containing at least three animals, e and f represent the combined results of 2 independent experiments. Mean values ± SEM are indicated in b, d, f, Pearson’s test was performed in (e). p ≤ 0.05 =*, p ≤ 0.01 = **, p ≤ 0.001 = ***.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 1755 kb)
Rights and permissions
About this article
Cite this article
Halim, T., Hwang, Y., Scanlon, S. et al. Group 2 innate lymphoid cells license dendritic cells to potentiate memory TH2 cell responses. Nat Immunol 17, 57–64 (2016). https://doi.org/10.1038/ni.3294
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3294
This article is cited by
-
Group 2 innate lymphoid cells and their surrounding environment
Inflammation and Regeneration (2023)
-
An αvβ3 integrin checkpoint is critical for efficient TH2 cell cytokine polarization and potentiation of antigen-specific immunity
Nature Immunology (2023)
-
The kinase p38α functions in dendritic cells to regulate Th2-cell differentiation and allergic inflammation
Cellular & Molecular Immunology (2022)
-
Physiological microbial exposure transiently inhibits mouse lung ILC2 responses to allergens
Nature Immunology (2022)
-
Early-life EV-A71 infection augments allergen-induced airway inflammation in asthma through trained macrophage immunity
Cellular & Molecular Immunology (2021)