Abstract
A primary goal of systems biology is to understand the design principles of the transcription networks that govern the timing of gene expression1,2,3,4,5. Here we measured promoter activity for ∼100 genes in parallel from living cells at a resolution of minutes and accuracy of 10%, based on GFP and Lux reporter libraries3. Focusing on the amino-acid biosynthesis systems of Escherichia coli4, we identified a previously unknown temporal expression program and expression hierarchy that matches the enzyme order in unbranched pathways. We identified two design principles: the closer the enzyme is to the beginning of the pathway, the shorter the response time of the activation of its promoter and the higher its maximal promoter activity. Mathematical analysis suggests that this 'just-in-time' (ref. 5) transcription program is optimal under constraints of rapidly reaching a production goal with minimal total enzyme production6,7. Our findings suggest that metabolic regulation networks are designed to generate precision promoter timing and activity programs that can be understood using the engineering principles of production pipelines.
Similar content being viewed by others
Main
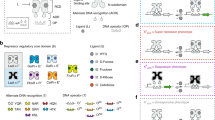
Amino-acid biosynthesis (AAB) in E. coli is carried out by well-characterized enzymatic pathways4,6,7,8,9,10,11. The genes encoding these enzymes are governed by a transcriptional regulatory network12,13, which is an excellent model system for studying the design principles of metabolic regulation. To study the dynamics of transcription of AAB genes at high temporal resolution and accuracy, we constructed a library of 52 reporter strains that represent ∼50% of known AAB genes. We designed each reporter strain by cloning one of the promoter regions of E. coli K-12 MG1655 upstream of a Lux or a fast-folding GFP reporter gene (Fig. 1a). We measured promoter activity with a high temporal resolution by measuring fluorescence, luminescence and absorbance from 96 cultures in parallel in a multiwell fluorimeter3,14. The fluorimeter allowed automated readings at 4-min or 8-min intervals during growth with shaking at a constant temperature. Day-to-day reproducibility was ∼10% (Fig. 1b,c).
(a) Reporter plasmids: pUA66 contains the gene GFPmut2; pUAL94 contains the operon LuxCDABE. Both vectors include a BamHI and XhoI cloning site for the promoter region, a low-copy origin (SC101 origin) and a kanamycin resistance gene. Two additional vectors (pUA139 with GFPmut2 and pUAL306 with luxCDABE) contain a reversed (BamHI-XhoI) cloning site for promoters in the opposite direction (not shown). Day-to-day reproducibility of the high-throughput measurement of absorbance (b) and GFP (c). The mean relative error for the absorbance and fluorescence measurements was 11% and 10%, respectively.
We first studied the dynamics of the AAB promoter activity after a shift from defined medium with no amino acids to defined medium supplemented with one amino acid (Fig. 2). We measured expression every 8 min for 8 h of growth and plotted data as a ratio relative to the expression at the same absorbance in the absence of amino acids. Virtually all promoters were active in the absence of amino acids, whereas promoter activity was repressed in the presence of the complete set of amino acids (Fig. 2). Transferring the cells to medium that included one amino acid generally resulted in the downregulation of the corresponding biosynthesis pathway (Fig. 2); arginine biosynthesis is a prominent example. Addition of glutamate, a precursor of the arginine biosynthesis pathway, also resulted in a mild downregulation of the arginine biosynthesis genes. We found many such cross-activations and -repressions15, in which a given amino acid affected the promoter activity of genes belonging to other amino acid pathways. For example, addition of leucine downregulated the activity of many of the promoters, probably owing to the activity of the global transcription factor Lrp16,17. These results suggest that there is an intricate interplay between amino acids and the transcriptional regulation of the various biosynthesis genes.
Reporter strains were grown on defined medium (M9 with 0.5% glucose and 25 μg ml−1 of kanamycin) supplemented with one amino acid (as indicated at the top), with all 20 amino acids (last column) or with no amino acids. Rows represent promoters, arranged by amino acid system, and columns represent experimental conditions, with 31 absorbance points between A = 0.01 and A = 0.15 per condition, corresponding to early- to mid-logarithmic growth. Shown is the logarithm of the ratio of the promoter activity to the promoter activity in the absence of all amino acids at the same absorbance. Blue indicates downregulation; red indicates upregulation; green-yellow indicates no substantial change in the expression levels. Ilv, leucine/isoleucine/valine; Aro, aromatic amino acids.
We then focused on the detailed activation dynamics of AAB systems after removing the amino acid. We studied the dynamics of selected systems using fewer reporter strains per plate, which allowed for higher temporal resolution. We analyzed the promoter activity of the nine arginine biosynthesis operons using GFP and Lux reporters, at a 4-min resolution. Eight of the operons encode a single enzyme, and one, argCBH, encodes three. To activate the arginine system, we diluted the cells into a defined medium supplemented with all amino acids except arginine. To study the relative timing, we normalized the activity profile of each of the promoters to its maximal level (Fig. 3). We found a detailed temporal order of expression with delays on the order of 10 min between the different promoters.
Lux reporter strains were grown in defined medium containing all amino acids and diluted into the same medium lacking arginine. Shown is the ratio of Lux activity to absorbance at each time point divided by the highest ratio for each strain. Blue and red indicate low and high expression, respectively. The pathway diagram illustrates the metabolic steps of the arginine biosynthesis pathway and the dashed arrows indicate the stages at which each gene product participates. Similar results were obtained using GFP reporter strains.
The main path for arginine production uses glutamate as a precursor metabolite. Two other amino acids, glutamine and aspartate, are also necessary for arginine production, and their products converge into the main pathway from two additional linear pathways. We found that the temporal order matched the functional enzyme order in each unbranched pathway (Fig. 3). In the main pathway, argA was the first promoter to be upregulated, followed by argCBH, argD and argE. This is the order in which the ArgA, ArgBC, ArgD and ArgE gene products convert glutamate to ornithine. Temporal expression also matched gene order in the other two unbranched pathways in the arginine system. carAB preceded argF and argI, which function in a complex18, in a pathway that converts glutamine to citrulline, and argG preceded argH in the pathway that converts citrulline to arginine. The timing of the ArgG-ArgH pathway was similar to that of the early promoters in the ArgA-ArgE pathway, even though the ArgG-ArgH pathway acts on the end product of the ArgA-ArgE pathway. Therefore, gene functional order seems to match the timing order only within an unbranched pathway and not between different pathways.
Two other complete unbranched pathways are available in our strain library: those of serine and methionine biosynthesis. We found temporal order in these systems also, after the systems were induced by a shift to medium with no serine or methionine but containing all other amino acids (Fig. 4). In the serine biosynthesis pathway, the expression order (serA, serC, serB) matched the functional order of the enzymes in the pathway (Fig. 4a). In the methionine biosynthesis system, the expression order (metA, metB, metC) also matched the pathway order (Fig. 4b).
GFP reporter strains were grown on defined medium containing all 20 amino acids and then transferred into the same medium lacking serine (a) or methionine (b). Shown is the ratio of GFP activity to absorbance at each time point divided by the highest ratio for each strain. Blue and red indicate low and high expression, respectively. The pathway diagram illustrates the metabolic steps in the pathways and the dashed arrows indicate the stages at which each gene product participates.
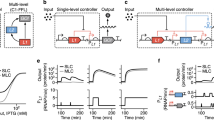
To detect the temporal order (Figs. 3 and 4), we considered the normalized activity for each promoter. When considering the un-normalized promoter activity in the arginine biosynthesis pathway from argA through argE, we observed a hierarchy in the activity levels (Fig. 5a): argA reached the highest promoter activity, followed by argCBH, argD and argE. Thus, the closer an enzyme is to the beginning of the pathway, the higher its maximal level of promoter activity. We found that the same principle applied in the methionine and serine biosynthesis pathways (Fig. 5d,e). In the methionine biosynthesis pathway, the promoter activity hierarchy was metA > metB > metC, and in the serine biosynthesis pathway, it was serA > serC > serB.
(a) Un-normalized expression kinetics of arginine biosynthesis promoters for genes in the unbranched pathway from glutamate to ornithine. The reporter strains argA, argCBH, argD and argE were diluted into a medium that lacks arginine (as in Fig. 3). (b) Normalized expression kinetics for argA, argCBH, argD and argE. Dashed lines represent response time, the time to reach 50% of the maximal expression. (c) Maximal level of expression versus response time for argA, argCBH, argD and argE. The mean ± s.d. of two experiments is shown. (d) Un-normalized expression kinetics of methionine biosynthesis promoters. The reporter strains metA, metB and metC were diluted into a minimal medium that lacks methionine (as in Fig. 4b). (e) Un-normalized expression kinetics of serine biosynthesis promoters. The reporter strains serA, serC and serB were diluted into a minimal medium that lacks serine (as in Fig. 4a). (f) Maximal level of promoter activity versus response time for methionine biosynthesis promoters. The mean ± s.d. of two experiments is shown. (g) Maximal level of promoter activity versus response time for serine biosynthesis promoters. The mean ± s.d. of two experiments is shown. (h) Maximal level of promoter activity versus response time for the lacZYA promoter induced with various levels of IPTG ranging from 1 μM to 1 mM. Similar results were obtained with the corresponding Lux reporter.
The temporal order and the activity hierarchy can be represented in a single plot by considering the response time, defined as the time it takes to reach half of the maximal promoter activity level. In the arginine, methionine and serine systems, the earlier the enzyme is involved in a pathway, the shorter is the response time and the higher the maximal promoter activity of its gene (Fig. 5c,f,g).
In control experiments, we induced the lacZYA promoter to different levels using the inducer isopropyl-β-D-thiogalactopyranoside (IPTG) and measured its promoter activity dynamics (Fig. 5h). The promoter activity level increased with IPTG, as expected, but the response time was nearly independent of IPTG levels. Hence, in this control system, expression strength did not correlate significantly with response time.
Unbranched biosynthesis pathways can be viewed as 'production pipelines', in which the end product is produced from an initial substrate through several intermediate steps. We analyzed an enzyme pathway, using a theoretical approach that is an extension of classical theoretical studies6,7,19,20. Important extensions include treatment of transcriptional regulation, as well as dilution effects due to cell growth. We modeled a pathway consisting of three consecutive steps, representing a biosynthetic pathway composed of three enzymes. The enzymes E1, E2 and E3 produce products according to standard Michaelis-Menten equations, in which the rate of production, utilization and dilution of metabolite Si is described by equation 1 in Box 1 in which α is the cell division rate and one can assume for simplicity that the enzymes have the same velocities (Vi = V) and Km values (Kmi = Km; ref. 19). The initial substrate S0 is assumed to be present at saturation. The amino acid product is P = S3. The equation for S3 has an additional sink term representing the cell's use of the amino acid, represented by α′ S3. The concentration of each enzyme is a balance of its expression rate and dilution by cell growth21,22 (equation 2 in Box 1).
The parameters βi and ki are the maximal promoter activity and repression coefficient3 of gene i. ki , the concentration of repressor needed for 50% repression, represents complex combinations of the binding affinity of the repressor to its cis-regulatory site, the strength of the RNA polymerase binding and the relative positions of these sites23. The active repressor level (repressor bound to P) is given by equation 3 in Box 1 in which RT is the total repressor concentration and Kr is the dissociation constant. The flux of the product is given by equation 4 in Box 1.
To find the optimal gene expression program, we optimized the parameters βi , ki and Kr to minimize a cost function given by equation 5 in Box 1. This cost function is composed of two factors: the cost to produce the enzymes and the rate and precision at which F approaches its goal, Fgoal. The cost function has two parameters: a represents the relative cost of producing the enzymes, and the integration time T represents a typical time scale for the duration of activation of the system in the environment1.
The optimized solution that minimizes this cost function shows a hierarchy in promoter strengths as well as temporal order (Fig. 6): the earlier the enzyme acts in the pathway, the higher its maximal promoter activity and the earlier it is expressed. These programs are generated in the optimal model solution by differential βi and ki values. We find β1 > β2 > β3, representing a hierarchy in the maximal promoter activity, and k1 < k2 < k3, representing feedback strength by the repressor that is stronger the earlier the enzyme in the pathway.
(a) Enzyme concentrations as a function of time after induction of the pathway. (b) Enzyme concentrations normalized by their steady-state values. (c) Maximal level of expression versus response time (time to 50% of maximal concentration) for the three enzymes. The enzyme kinetic parameters (equation 1) are V = 1, Km = 1, and the dilution and utilization rates are α = 1, α′ = 2. The cost function in equation 5 (Box 1) used T = 10, Fgoal = 0.006, a = 0.001. The optimal coefficients of the regulatory system are β1 = 3.4, β2 = 0.67, β3 = 0.10, k1 = 0.23, k2 = 0.68, k3 = 2.1, Kr = 1 × 10−4. (d) Properties of the optimal solution for various values of the cost function parameters T (integration time) and a (relative cost of enzyme production). In the blue region, the optimal solutions show the following two properties: (i) τ1 < τ2 < τ3 and (ii) max (E1) > max (E2) > max (E3). τi is the response time (time to reach 50% of maximal concentration) and max (Ei) is the maximal concentration of the enzyme.
These qualitative features of the optimized solution apply to a broad region of the cost function parameters a and T (a < 0.01 and T > 5; Fig. 6d). As T increases, the differences between the response times of the enzymes in the pathway become smaller, reflecting optimization with greater emphasis on steady-state production than on the transient phase of the response. We analytically show in Supplementary Note online that when T → ∞, the optimal solution has a hierarchy of expression levels6.
To understand intuitively this optimized solution and the role of the delays between the AAB genes, it is useful to estimate how long it takes an enzyme to produce enough product to reach the Km of the next enzyme in the pathway. At full production, each stage of a linear AAB pathway needs to makes about 107 product molecules per cell cycle4. Typical AAB enzymes have a Km on the order of 1 mM, which corresponds to about 106 substrate molecules per cell4. It therefore typically takes about 0.1 cell cycles for enzyme i to produce enough molecules to reach the Km of enzyme i+1. In our experiment, the cell cycle is about 100 min. The estimated delay, Tdelay = 0.1 cell cycles ∼10 min, is of the same order as the observed delays between enzyme promoter activations in our experimental data. The present mathematical treatment also helps us to understand the role of the observed hierarchy of promoter activity: higher promoter activity for the early enzymes makes the response faster by providing a boost of initial substrates. In addition, increased production rate of the early enzymes and substrates reduces the effects of their dilution by cell growth over the time it takes to express the full pathway24.
Some of the AAB pathways, such as the arginine pathway, are controlled by a single repressor that also represses its own transcription21,25. We added such autoregulation to the mathematical pathway model (Supplementary Note online) using equation 6 in Box 1 in which βo and ko describe the interaction of the repressor with its own promoter. The optimal solutions show an increase in the regulator level from its basal level, an overshoot and finally a relaxation to a steady-state level that is modestly higher than the basal level. The optimal response time of the repressor is faster than that of any of the enzymes in the pathway (Supplementary Fig. 1 online). Our experimental data for the arginine system show a similar behavior: the promoter of the gene encoding the master repressor, argR, is the earliest activated (Fig. 3 and Supplementary Fig. 1 online).
Genes that participate in multi-step differentiation processes5,26,27 or in building protein machines such as flagella14,27 are often expressed in a temporal order that matches the functional order of the gene products3,13. Here, we found such a temporal order in a new type of gene system: metabolic pathways. There are two design principles in the unbranched pathways that we analyzed: the earlier the enzyme acts in the pathway, the shorter the response time of its promoter activity and the higher its maximal promoter activity. We also found that these two design principles characterize optimal solutions of a simple mathematical model, optimized to rapidly reach a desired product goal with minimal enzyme production. More generally, these two design principles may characterize gene systems regulated by the commonly occurring single-input-module network motif 3,13, in which a single regulator controls a set of target genes.
The present findings suggest that regulation of metabolic pathways has an unexpectedly beautiful precision design, and that this design can be understood by means of optimization for engineering criteria1,6,7,8,9,10,11,19,20 such as response time and resource economy. The present approach can be used to study experimentally the design principles of other metabolic pathways.
Methods
Materials.
We obtained enzymes from Roche, amino acids from Calbiochem, primers from Sigma, 96-well PCR plates from Symport and 96-well plates for overnight growth from Nunc.
Primer design.
We designed primers to flank regions between the two open reading frames (ORFs) that border each promoter region as described3. The primers amplify the regions between two adjacent ORFs with an extension of 50–100 bp into each ORF. If no suitable primers were found, we extended the amplified region to 300 bp into each ORF. For AAB promoters known to contain leader peptides28, the promoter region included the leader peptide coding region. We designed each of the two primers with either XhoI or BamHI restriction sites. If the target genomic promoter region contained XhoI or BamHI sites, we used other restriction enzymes that leave the same overhang (SalI or BglII, respectively). All the primers were designed with Tm = 60 °C to allow high-throughput multiwell PCR.
Vector preparation.
The vectors used for the reporter strain library construction contain a reporter gene, GFPmut2 (ref. 29) or luxCDABE (ref. 30; Fig. 1a), under the control of the promoter. We used two versions of each type of the vectors, one for promoters in the positive direction (pUA66 for GFP and pUA94 for Lux) and the others for promoters in the negative direction (pUA139 for GFP and pUA306 for Lux). We purified the plasmids by Maxi-Prep (Qiagen) from MG1655 cells, digested them with XhoI for 7 h at 37 °C and then purified and digested them with BamHI for 5 h at 37 °C. We checked digested vectors for background by self-ligation experiments followed by transformation.
PCR and digestion of the promoter regions.
Primers were arranged in 96-well block plates and carried out PCR in 96-well plates (Symport). We used E. coli K-12 MG1655 genomic DNA as template and the Expand High-Fidelity PCR System (Roche). PCR conditions were 95 °C for 10 min, followed by 24 cycles of 95 °C for 30 s, 60 °C for 1 min and 72 °C for 1 min, and a final step of 72 °C for 5 min. We checked product size on 96-lane 1% agarose gel. We then purified PCR products using a 96-well PCR purification kit (Qiagen) and digested them for 6 h at 30 °C with XhoI and BamHI restriction enzymes. For primers using SalI or BglII restriction sites, we carried out the digestion in two steps. All digestions took place in 96-well plates (Symport). We cleaned the digested PCR products using a 96-well PCR kit (Qiagen).
Ligation and transformation.
We ligated the digested PCR products (T4 ligase, Roche) overnight at room temperature with the corresponding vectors in 96-well plates and transformed them into CaCl2-competent MG1655 cells in high-brim 96-well plates. We then plated the transformed cells on 9-cm plates with selective media with 25 μg ml−1 of kanamycin. We screened positive colonies by 96-well colony PCR using primers designed for the upstream and downstream regions of the two restriction sites, BamHI and XhoI. Primer sequences are available on request. We loaded products on 96-well agarose gels to check for the correct sizes of the corresponding inserts. We prepared frozen stocks (25% glycerol) of the reporter strains in 96-well plates. We sequenced 25 randomly selected clones and detected no mutations.
Reproducibility experiment.
We inoculated 80 different reporter strains, including 45 for AAB operons, from frozen stocks into Luria broth medium supplemented with 25 μg ml−1 of kanamycin and grew them for 16 h. We then diluted the cells 1:300 into a fresh Luria broth medium with 25 μg ml−1 of kanamycin to a total volume of 150 μl and covered them with 100 μl of mineral oil (Sigma). The dilution was done in a 96-well flat-bottom plate (Nunc), where each well contained a different reporter strain culture. We allowed the cells to grow for 10 h at 37 °C with shaking as described3 in a Wallac Victor2 multiwell fluorimeter and measured the fluorescence (at 535 nm) and the absorbance (at 600 nm) every 8 min (75 time points per promoter) for a total of 6,000 data points (Fig. 1b,c).We normalized the absorbance and fluorescence data to the mean for each gene in each experiment and multiplied by the overall mean of the two experiments.
Experiments for measuring the expression levels under various conditions.
The reporter strains used in these experiments (Supplementary Table 1 online) were directly inoculated from frozen stocks and grown for 20 h in Luria broth with kanamycin (25 μg ml−1) at 37 °C. We then diluted the cultures 1:150 into the specific experimental medium at a final volume of 150 μl per well in flat-bottom 96-well plates and, as previously described, covered them with mineral oil and grew them in a Victor2 multiwell fluorimeter. Time between repeated measurements was 8 min. We subtracted the background fluorescence of cells bearing a promoterless GFP vector.
Amino acid deprivation experiments.
We inoculated frozen stocks of the amino acid reporter strains (Supplementary Table 1 online) into M9 defined medium supplemented with 0.5% glucose, 0.2% casamino and 25 μg ml−1 of kanamycin. We grew the cells for 16 h at 37 °C in deep 96-well plates with shaking. We then diluted the cell cultures 1:150 to a final volume of 150 μl with M9 medium plus 0.5% glucose and 25 μg ml−1 of kanamycin without amino acids, with one amino acid or with the complete set of amino acids in 96-well plates and covered them with 100 μl of mineral oil to avoid evaporation3. The concentration of amino acids added to the medium were as follows: alanine, 0.47 mM; arginine, 0.6 mM; asparagine, 0.32 mM; aspartic acid, 0.3mM; cysteine, 0.3 mM; glutamate, 5 mM; glutamine, 5 mM; glycine, 0.13 mM; histidine, 0.1 mM; isoleucine, 0.3 mM; leucine, 0.3 mM; lysine, 0.3 mM; methionine, 0.3 mM; phenylalanine, 0.3 mM; proline, 2 mM; serine, 4 mM; threonine, 0.3 mM; tryptophan, 0.1 mM; tyrosine, 0.1 mM; valine, 0.3 mM. We inserted the plate into a Victor 2 multiwell fluorimeter (Wallac) and measured the luminescence, fluorescence (at 535 nm) and absorbance (at 600 nm). Experiments were done at 30 °C with orbital shaking before each measurement. The full plate was measured every 8 min (Fig. 2). In experiments with fewer strains, we measured every 4 min (Figs. 3,4,5). We measured parameters for the lacZYA promoter in the same conditions with different concentrations of IPTG (Sigma). We analyzed data using custom Matlab software.
Note: Supplementary information is available on the Nature Genetics website.
References
Savageau, M.A. Biochemical Systems Analysis: A Study of Function and Design in Molecular Biology (Addison-Wesley, Reading, Massachusetts, 1976).
Hartwell, L.H., Hopfield, J.J., Leibler, S. & Murray, A.W. From molecular to modular cell biology. Nature 402, C47–C52 (1999).
Ronen, M., Rosenberg, R., Shraiman, B.I. & Alon, U. Assigning numbers to the arrows: parameterizing a gene regulation network by using accurate expression kinetics. Proc. Natl. Acad. Sci. USA 99, 10555–10560 (2002).
Neidhardt, F.C. (ed.) Escherichia coli and Salmonella Cellular and Molecular Biology (American Society for Microbiology Press, 1996).
McAdams, H.H. & Shapiro, L. A bacterial cell-cycle regulatory network operating in time and space. Science 301, 1874–1877 (2003).
Heinrich, R. & Klipp, E. Control analysis of unbranched enzymatic chains in states of maximal activity. J. Theor. Biol. 182, 243–252 (1996).
Klipp, E., Heinrich, R. & Holzhutter, H.G. Prediction of temporal gene expression. Metabolic optimization by re-distribution of enzyme activities. Eur. J. Biochem. 269, 5406–5413 (2002).
Ibarra, R.U., Edwards, J.S. & Palsson, B.O. Escherichia coli K-12 undergoes adaptive evolution to achieve in silico predicted optimal growth. Nature 420, 186–189 (2002).
Heinrich, R. & Schuster, S. The Regulation of Cellular Systems (Chapman & Hall, New York, 1996).
Gilman, A. & Ross, J. Genetic-algorithm selection of a regulatory structure that directs flux in a simple metabolic model. Biophys. J. 69, 1321–1333 (1995).
Segre, D., Vitkup, D. & Church, G.M. Analysis of optimality in natural and perturbed metabolic networks. Proc. Natl. Acad. Sci. USA 99, 15112–15117 (2002).
Covert, M.W. & Palsson, B.O. Transcriptional regulation in constraints-based metabolic models of Escherichia coli. J. Biol. Chem. 277, 28058–28064 (2002).
Shen-Orr, S.S., Milo, R., Mangan, S. & Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet. 31, 64–68 (2002).
Kalir, S. et al. Ordering genes in a flagella pathway by analysis of expression kinetics from living bacteria. Science 292, 2080–2083 (2001).
Oh, M.K. & Liao, J.C. Gene expression profiling by DNA microarrays and metabolic fluxes in Escherichia coli. Biotechnol. Prog. 16, 278–286 (2000).
Calvo, J.M. & Matthews, R.G. The leucine-responsive regulatory protein, a global regulator of metabolism in Escherichia coli. Microbiol. Rev. 59, 466–490 (1994).
Hung, S.P., Baldi, P. & Hatfield, G.W. Global gene expression profiling in Escherichia coli K12. The effects of leucine-responsive regulatory protein. J. Biol. Chem. 277, 40309–40323 (2002).
Legrain, C., Halleux, P., Stalon, V. & Glansdorff, N. The dual genetic control of ornithine carbamolytransferase in Escherichia coli. A case of bacterial hybrid enzymes. Eur. J. Biochem. 27, 93–102 (1972).
Pettersson, G. Optimal kinetic design of enzymes in a linear metabolic pathway. Biochim. Biophys. Acta 1164, 1–7 (1993).
Heinrich, R., Schuster, S. & Holzhutter, H.G. Mathematical analysis of enzymic reaction systems using optimization principles. Eur. J. Biochem 201, 1–21 (1991).
Rosenfeld, N., Elowitz, M.B. & Alon, U. Negative autoregulation speeds the response times of transcription networks. J. Mol. Biol. 323, 785–793 (2002).
Rosenfeld, N. & Alon, U. Response delays and the structure of transcription networks. J. Mol. Biol. 329, 645–654 (2003).
Charlier, D. et al. Arginine regulon of Escherichia coli K-12. A study of repressor-operator interactions and of in vitro binding affinities versus in vivo repression. J. Mol. Biol. 226, 367–386 (1992).
Aizawa, S.I. & Kubori, T. Bacterial flagellation and cell division. Genes Cells 3, 625–634 (1998).
Savageau, M.A. Comparison of classical and autogenous systems of regulation in inducible operons. Nature 252, 546–549 (1974).
Chu, S. et al. The transcriptional program of sporulation in budding yeast. Science 282, 699–705 (1998).
Laub, M.T., McAdams, H.H., Feldblyum, T., Fraser, C.M. & Shapiro, L. Global analysis of the genetic network controlling a bacterial cell cycle. Science 290, 2144–2148 (2000).
Henkin, T.M. & Yanofsky, C. Regulation by transcription attenuation in bacteria: how RNA provides instructions for transcription termination/antitermination decisions. Bioessays 24, 700–707 (2002).
Cormack, B.P., Valdivia, R.H. & Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173, 33–38 (1996).
Goh, E.B. et al. Transcriptional modulation of bacterial gene expression by subinhibitory concentrations of antibiotics. Proc. Natl. Acad. Sci. USA 99 (26), 17025–17030 (2002).
Acknowledgements
We thank E. Klipp, W. Leibermeister, M. Vidal and all members of our laboratory for discussions. U.A. acknowledges support from the US National Institutes of Health, Israel Science Foundation and Minerva. A.M. acknowledges a Pacific Theatres Foundation postdoctoral grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Zaslaver, A., Mayo, A., Rosenberg, R. et al. Just-in-time transcription program in metabolic pathways. Nat Genet 36, 486–491 (2004). https://doi.org/10.1038/ng1348
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1348
This article is cited by
-
Enzyme expression kinetics by Escherichia coli during transition from rich to minimal media depends on proteome reserves
Nature Microbiology (2023)
-
Dosage sensitivity of JDPs, a valuable tool for understanding their function: a case study on Caj1 overexpression-mediated filamentous growth in budding yeast
Current Genetics (2021)
-
Using optimal control to understand complex metabolic pathways
BMC Bioinformatics (2020)
-
Machine learning-powered antibiotics phenotypic drug discovery
Scientific Reports (2019)
-
Ordering up gene expression by slowing down transcription factor binding kinetics
Current Genetics (2019)