Abstract
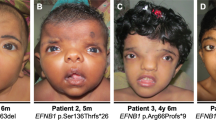
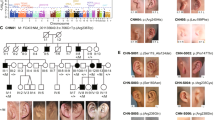
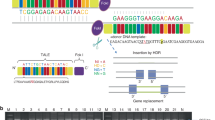
Saethre-Chotzen syndrome (acrocephalo-syndactyly type III, ACS III) is an autosomal dominant craniosynostosis with brachydactyly, soft tissue syndactyly and facial dysmorphism including ptosis, facial asymmetry and prominent ear crura. ACS III has been mapped to chromosome 7p21–22. Of interest, TWIST, the human counterpart of the murine Twist gene, has been localized on chromosome 7p21 as well. The Twist gene product is a transcription factor containing a basic helix–loop–helix (b-HLH) domain, required in head mesenchyme for cranial neural tube morphogenesis in mice. The co-localisation of ACS III and TWIST prompted us to screen ACS III patients for TW/57gene mutations especially as mice heterozygous for Twist null mutations displayed skull defects and duplication of hind leg digits. Here, we report 21-bp insertions and nonsense mutations of the TWIST gene (S127X, E130X) in seven ACS III probands and describe impairment of head mesenchyme induction by TWIST as a novel pathophysiological mechanism in human craniosynostoses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Reardon, W. & Winter, R.M. Saethre-Chotzen syndrome. J. Med. Genet. 31, 393–396 (1994).
Brueton, L.A>, Van Herwerden, L.,, Chotai, K.A. & Winter, R.M. The mapping of a gene for craniosynostosis: evidence for linkage of the Saethre-Chotzen syndrome to distal chromosome 7p. J. Med. Genet. 29, 681–685 (1992).
Van Herwerden, L. et al. Evidence for locus heterogeneity in acrocephalosyndactyly: a refined localization for Saethre-Chotzen syndrome locus on distal chromosome 7p and exclusion of Jackson-Weiss syndrome from craniosynostosis loci on 7p and 5q. Am. J. Med. Genet. 54, 669–674 (1994).
Lewanda, A.F. et al. Genetic heterogeneity among craniosynostosis syndromes: mapping the Saethre-Chotzen syndrome locus between D7S513 and D7S516 and exclusion of Jackson-Weiss and Crouzon syndrome loci from 7p. Genomics 19, 115–119 (1994).
Lewanda, A.F. et al. Evidence that the Saethre-Chotzen syndrome locus lies between D7S664 and D7S507, by genetic analysis and detection of a microdeletion in a patient. Am. J. Hum. Genet. 55, 1195–1201 (1994).
Rose, C.S.P. et al. Localization of the genetic locus for Saethre-Chotzen syndrome to a 6 cM region of chromosome 7 using four cases with apparently balanced translocations at 7p21.2. Hum. Mol. Genet. 3, 1405–1408 (1994).
Tsuji, K. et al. Craniosynostosis and hemizygosity for D7S135 caused by a de novo and apparently balanced t(6;7) translocation. Am. J. Med. Genet. 49, 98–102 (1994).
Readron, W., McManus, S.P., Summers, D. & Winter, R.M. Cytogenetic evidence that the Saethre-Chotzen syndrome maps to 7p21.2. Am. J. Med. Genet. 47, 633–636 (1993).
Reid, C.S. et al. Saethre-Chotzen syndrome with familial translocation at chromosome 7p22. Am. J. Med. Genet. 47, 637–639 (1993).
Ma, H.W. et al. Possible genetic heterogeneity in the Saethre-Chotzen syndrome. Hum. Genet. 98, 228–232 (1996).
Muenke, M. et al. A common mutation in the fibroblast growth factor receptor 1 gene in Pfeiffer syndrome. Nature Genet. 8, 269–274 (1994).
Wilkie, A.O.M. et al. Apert syndrome results from localized mutations of FGFR2 and is allelic with Crouzon syndrome. Nature Genet. 9, 165–172 (1995).
Lajeunie, E. et al. FGFR2 mutations in Pfeiffer syndrome. Nature Genet. 9, 108 (1995).
Rutland, P. et al. Identical mutations in the FGFR2 gene cause both Pfeiffer and Crouzon syndrome phenotypes. Nature Genet. 9, 173–176 (1995).
Meyers, G.A. et al. FGFR2 exon Ilia and Ilk mutations in Crouzon, Jackson-Weiss and Pfeiffer syndromes: evidence for missense changes, insertions and a deletion due toalternative RNA splicing. Am. J. Hum. Genet. 58, 491–498 (1996).
Bourgeois, P., Stoezel, C., Bolcato-Bellemin, A.L.,, Mattel, M.G & Perrin-Schmitt, F. The human TWIST gene is located at 7p21 and encodes a b-HLH protein which is 96% similar to its murine M-twist counterpart. Mamm.Genome 7, 915–917.
Benezra, R., Davis, R.L., Lockshon, D., Turner, D.L. & Weintraub, H. The protein Id: a negative regulator of helix-loop-helix DNA binding proteins. Cell 61, 49–59 (1990).
Chen, Z.F. & Behringer, R.R. twist is required in head mesenchyme for cranial neural tube morphogenesis. Genes & Dev. 9, 686–699 (1995).
Murre, C., McCaw, P.S. & Baltimore, D. A new DNA binding and dimerization motif in immunoglobulin enhancer binding, daughterless, MyoD, and myc proteins. Cell 56, 777–783 (1989).
Thisse, B., Stoetzel, C., Gorostiza-Thisse, C. & Perrin-Schmitt, F. Sequence of the twist gene and nuclear localization of its protein in endomesodermal cells of early Drosophila embryos. Embo J. 7, 2175–2183 (1988).
Wolf, C., Thisse, C., Stoezel, C., Thisse, B., Gerlinger, P. & Perrin-Schmitt, F., M-twistgene of Mus is expressed in subsets of mesodermal cells and is closely related to the Xenopus X-twi and the Drosophila twist genes. Dev. Biol. 143, 363–373 (1991).
Stoetzel, C., Weber, B., Bourgeois, R., Bolcato-Bellemin, A.L. & Perrin-Schmitt, F. Dorso-ventral and rostro-caudal sequential expression of M-twist in the postimplantation murine embryo. Mech. Dev. 51, 251–263 (1995).
Vortkamp, A., Gessber, M. & Grzeschik, K.H. GLI3 zinc finger gene interruped by translocations in Greig syndrome families. Nature 352, 539–540 (1991).
Jabs, E.W. et al. A mutation in the homeodomain of the human MSX2 gene in a family affected with autosomal dominant craniosynostosis. Cell 75, 443–450 (1993).
Shishido, E., Higashijima, S., Emori, Y. & Saigo, K., FGF-receptor homologues of Drosophila: one is expressed in mesodermal primordium in early embryos. Development 117, 751–761 (1993).
Amaya, E., Musci, T.J. & Kirschner, M.W. Expression of a dominant negative mutant of the FGF receptor disrupts mesoderm formation in Xenopus embryos. Cell 66, 257–270 (1991).
Deng, C.X. et al. Murine FGFR1 is required for early postimplantation growth and axial organization. Genes & Dev. 8, 3045–3057 (1994).
Yamaguchi, T.P., Harpal, K., Henkemeyer, M. & Rossant, J. FGFR-1 is required for embryonic growth and mesodermal patterning during mouse gastrulation. Genes & Dev. 8, 3032–3044 (1994).
Inouye, M. Differential staining of cartilage and bone in fetal mouse skeleton by alcian blue and alizarin red. Congenital Anom. 16, 171–173 (1976).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ghouzzi, V., Merrer, M., Perrin-Schmitt, F. et al. Mutations of the TWIST gene in the Saethre-Chotzene syndrome. Nat Genet 15, 42–46 (1997). https://doi.org/10.1038/ng0197-42
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0197-42
This article is cited by
-
CDC20 promotes radioresistance of prostate cancer by activating Twist1 expression
Apoptosis (2023)
-
Twist alters the breast tumor microenvironment via choline kinase to facilitate an aggressive phenotype
Molecular and Cellular Biochemistry (2023)
-
The spontaneous mouse mutant low set ears (Lse) is caused by tandem duplication of Fgf3 and Fgf4
Mammalian Genome (2023)
-
A multi-stem cell basis for craniosynostosis and calvarial mineralization
Nature (2023)
-
The CMS19 disease model specifies a pivotal role for collagen XIII in bone homeostasis
Scientific Reports (2022)