Abstract
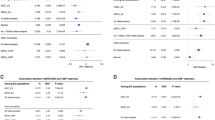
We conducted a meta-analysis of genome-wide association studies of systolic (SBP) and diastolic (DBP) blood pressure in 19,608 subjects of east Asian ancestry from the AGEN-BP consortium followed up with de novo genotyping (n = 10,518) and further replication (n = 20,247) in east Asian samples. We identified genome-wide significant (P < 5 × 10−8) associations with SBP or DBP, which included variants at four new loci (ST7L-CAPZA1, FIGN-GRB14, ENPEP and NPR3) and a newly discovered variant near TBX3. Among the five newly discovered variants, we obtained significant replication in the independent samples for all of these loci except NPR3. We also confirmed seven loci previously identified in populations of European descent. Moreover, at 12q24.13 near ALDH2, we observed strong association signals (P = 7.9 × 10−31 and P = 1.3 × 10−35 for SBP and DBP, respectively) with ethnic specificity. These findings provide new insights into blood pressure regulation and potential targets for intervention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ezzati, M. et al. Selected major risk factors and global and regional burden of disease. Lancet 360, 1347–1360 (2002).
Lopez, A.D. et al. Global and regional burden of disease and risk factors, 2001: systematic analysis of population health data. Lancet 367, 1747–1757 (2006).
Lawes, C.M.M., Vander Hoorn, S. & Rodgers, A. International Society of Hypertension. Global burden of blood-pressure-related disease, 2001. Lancet 371, 1513–1518 (2008).
Kearney, P.M. et al. Global burden of hypertension: analysis of worldwide data. Lancet 365, 217–223 (2005).
He, J. et al. Premature deaths attributable to blood pressure in China: a prospective cohort study. Lancet 374, 1765–1772 (2009).
Eastern Stroke and Coronary Heart Disease Collaborative Research Group. Blood pressure, cholesterol, and stroke in eastern Asia. Lancet 352, 1801–1807 (1998).
Cowley, A.W. Jr. The genetic dissection of essential hypertension. Nat. Rev. Genet. 7, 829–840 (2006).
Kurtz, T.W. Genome-wide association studies will unlock the genetic basis of hypertension: con side of the argument. Hypertension 56, 1021–1025 (2010).
Newton-Cheh, C. et al. Genome-wide association study identifies eight loci associated with blood pressure. Nat. Genet. 41, 666–676 (2009).
Levy, D. et al. Genome-wide association study of blood pressure and hypertension. Nat. Genet. 41, 677–687 (2009).
Dominiczak, A.F. & Munroe, P.B. Genome-wide association studies will unlock the genetic basis of hypertension: pro side of the argument. Hypertension 56, 1017–1020 (2010).
Anand-Srivastava, M.B. Natriuretic peptide receptor-C signaling and regulation. Peptides 26, 1044–1059 (2005).
Takeuchi, F. et al. Blood pressure and hypertension are associated with 7 loci in the Japanese population. Circulation 121, 2302–2309 (2010).
Chen, L., Davey Smith, G., Harbord, R.M. & Lewis, S.J. Alcohol intake and blood pressure: a systematic review implementing a Mendelian randomization approach. PLoS Med. 5, e52 (2008).
Tsuchihashi-Makaya, M. et al. Gene-environmental interaction regarding alcohol-metabolizing enzymes in the Japanese general population. Hypertens. Res. 32, 207–213 (2009).
Li, H. et al. Refined geographic distribution of the oriental ALDH2*504Lys (nee 487Lys) variant. Ann. Hum. Genet. 73, 335–345 (2009).
Takeuchi, F. et al. Confirmation of ALDH2 as a major locus of drinking behavior and of its variants regulating multiple metabolic phenotypes in Japanese. Circ. J. 75, 911–918 (2011).
Voight, B.F., Kudaravalli, S., Wen, X. & Pritchard, J.K. A map of recent positive selection in the human genome. PLoS Biol. 4, e72 (2006).
Sabeti, P.C. et al. Genome-wide detection and characterization of positive selection in human populations. Nature 449, 913–918 (2007).
Soranzo, N. et al. A genome-wide meta-analysis identifies 22 loci associated with eight hematological parameters in the HaemGen consortium. Nat. Genet. 41, 1182–1190 (2009).
Zollner, S. & Pritchard, J.K. Overcoming the winner's curse: estimating penetrance parameters from case-control data. Am. J. Hum. Genet. 80, 605–615 (2007).
Ong, R.T. & Teo, Y.Y. varLD: a program for quantifying variation in linkage disequilibrium patterns between populations. Bioinformatics 26, 1269–1270 (2010).
Stranger, B.E. et al. Population genomics of human gene expression. Nat. Genet. 39, 1217–1224 (2007).
Dixon, A.L. et al. A genome-wide association study of global gene expression. Nat. Genet. 39, 1202–1207 (2007).
Mizutani, S. et al. New insights into the importance of aminopeptidase A in hypertension. Heart Fail. Rev. 13, 273–284 (2008).
Cox, G.A., Mahaffey, C.L., Nystuen, A., Letts, V.A. & Frankel, W.N. The mouse fidgetin gene defines a new role for AAA family proteins in mammalian development. Nat. Genet. 26, 198–202 (2000).
Goenaga, D. et al. Molecular determinants of Grb14-mediated inhibition of insulin signaling. Mol. Endocrinol. 23, 1043–1051 (2009).
Heid, I.M. et al. Meta-analysis identifies 13 new loci associated with waist-hip ratio and reveals sexual dimorphism in the genetic basis of fat distribution. Nat. Genet. 42, 949–960 (2010).
Katoh, M. Molecular cloning and characterization of ST7R (ST7-like, ST7L) on human chromosome 1p13, a novel gene homologous to tumor suppressor gene ST7 on human chromosome 7q31. Int. J. Oncol. 20, 1247–1253 (2002).
Brooks, P.J., Enoch, M.A., Goldman, D., Li, T.K. & Yokoyama, A. The alcohol flushing response: an unrecognized risk factor for esophageal cancer from alcohol consumption. PLoS Med. 6, e50 (2009).
Harada, S., Misawa, S., Agarwal, D.P. & Goedde, H.W. Liver alcohol dehydrogenase and aldehyde dehydrogenase in the Japanese: isozyme variation and its possible role in alcohol intoxication. Am. J. Hum. Genet. 32, 8–15 (1980).
Barter, P.J. et al. Effects of torcetrapib in patients at high risk for coronary events. N. Engl. J. Med. 357, 2109–2122 (2007).
Tabara, Y. et al. Common variants in the ATP2B1 gene are associated with susceptibility to hypertension: the Japanese Millennium Genome Project. Hypertension 56, 973–980 (2010).
Hong, K.W. et al. Recapitulation of two genomewide association studies on blood pressure and essential hypertension in the Korean population. J. Hum. Genet. 55, 336–341 (2010).
Johnson, A.D. et al. SNAP: a web-based tool for identification and annotation of proxy SNPs using HapMap. Bioinformatics 24, 2938–2939 (2008).
Pickrell, J.K. et al. Signals of recent positive selection in a worldwide sample of human populations. Genome Res. 19, 826–837 (2009).
Li, Y., Willer, C.J., Ding, J., Scheet, P. & Abecasis, G.R. MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 34, 816–834 (2010).
Marchini, J. et al. A new multipoint method for genome-wide association studies by imputation of genotypes. Nat. Genet. 39, 906–913 (2007).
Browning, B.L. & Browning, S.R. A unified approach to genotype imputation and haplotype-phase inference for large data sets of trios and unrelated individuals. Am. J. Hum. Genet. 84, 210–223 (2009).
Cui, J.S., Hopper, J.L. & Harrap, S.B. Antihypertensive treatments obscure familial contributions to blood pressure variation. Hypertension 41, 207–210 (2003).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Higgins, J.P., Thompson, S.G., Deeks, J.J. & Altman, D.G. Measuring inconsistency in meta-analyses. Br. Med. J. 327, 557–560 (2003).
Ge, D. et al. WGAViewer: software for genomic annotation of whole genome association studies. Genome Res. 18, 640–643 (2008).
Acknowledgements
The authors acknowledge the essential role of the Asian Genetic Epidemiology Network (AGEN) in developing and supporting this manuscript. AGEN members include the Cardio-metabolic Genome Epidemiology (CAGE) Network, Genetic Epidemiology Network of Salt-Sensitivity (GenSalt), Korean Association Resource (KARE) Project, Shanghai Hypertension Study, Singapore Malay Eye Survey (SiMES), Singapore Prospective Study (SP2) Program, Suita Study and Taiwan Super Control Study.
CAGE: The CAGE Network Studies were supported by grants for the Core Research for Evolutional Science and Technology (CREST) from the Japan Science Technology Agency; the Program for Promotion of Fundamental Studies in Health Sciences, National Institute of Biomedical Innovation Organization (NIBIO); KAKENHI (Grant-in-Aid for Scientific Research) on Priority Areas 'Applied Genomics' from the Ministry of Education, Culture, Sports, Science and Technology of Japan; and the Grant of National Center for Global Health and Medicine (NCGM).
GenSalt: The Genetic Epidemiology Network of Salt Sensitivity is supported by research grants (U01HL072507, R01HL087263 and R01HL090682) from the National Heart, Lung, and Blood Institute, National Institutes of Health (Bethesda, Maryland, USA). T.N.K. is supported partially by Award Number K12HD043451 from the Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health (Bethesda, Maryland, USA).
KARE: KARE and HEXA-shared control studies were supported by grants from Korea Centers for Disease Control & Prevention, Republic of Korea (4845-301, 4851-302, 4851-307).
Shanghai: This work was supported by the Chinese National Key Program for Basic Research (grants 973:2004CB518603, 2006CB503804 and 2009CB521905) and Chinese National High Tech Program (grants 863:2009AA022703 and 2006AA02Z179) and the Ministry of Science and Technology, National Natural Science Foundation (30871361).
SiMES: The Singapore Malay Eye Study (SiMES) was funded by the National Medical Research Council (NMRC 0796/2003 and NMRC/STaR/0003/2008) and the Biomedical Research Council (BMRC, 09/1/35/19/616).
SP2: The Singapore Prospective Study Program (SP2) was funded through grants from the Biomedical Research Council of Singapore (BMRC 05/1/36/19/413 and 03/1/27/18/216) and the National Medical Research Council of Singapore (NMRC/1174/2008). Y.Y.T. acknowledges support from the Singapore National Research Foundation (NRF-RF-2010-05). E.S.T. also receives additional support from the National Medical Research Council through a clinician scientist award (NMRC/CSA/008/2009).
Suita/Ehime Study: The Ehime Study was supported by Grants for Scientific Research (Priority Areas 'Medical Genome Science (Millennium Genome Project)' and 'Applied Genomics') from the Ministry of Education, Culture, Sports, Science and Technology, Japan; a Grants-in-Aid (H15-longevity-005, H17-longevity-003, H16-kenko-001, H18-longevity (kokusai)) from the Ministry of Health, Labor and Welfare, Health and Labor Sciences Research Grants, Japan; a Science and Technology Incubation Program in Advanced Regions, Japan Science and Technology Agency; and the Japan Atherosclerosis Prevention Fund.
Taiwan Super Control Study: This study was supported by Academia Sinica Genomic Medicine Multicenter Study; National Research Program for Genomic Medicine, National Science Council, Taiwan (National Clinical Core, NSC97-3112-B-001-014; and National Genotyping Center, NSC97-3112-B-001-015).
Author information
Authors and Affiliations
Contributions
Principal investigators: N.K., J.H.
Project coordination leaders: N.K., J.H., Y.T.
Manuscript writing group: N.K., F.T., T.N.K., J.H., Y.Y.T., Y.S.C., E.S.T.
Project data management: T.N.K.
Genotyping and quality control: F.T., M.I., K.Y., Y.T., N.I., Y.K., X.S., W.T.T., Y.Y.T.
Phenotype collection, data management: CAGE: N.K., K.Y., T.K., T.N., M.Y., K.O., Y.Y., E.N., T.S., R.T., S.K., T.O.; GenSalt: T.N.K., D.G., J.H.; KARE: J.-P.J., S.S.K., Y.S.C.; Shanghai: Y.Z., X. Zhang, X. Zhou, D.Z.; SiMES/SP2: T.A., T.Y.W., E.S.T.; Suita: N.I., Y.K., Y.T., T.M.; Taiwan: C.-H.C., L.-c.C., Y.-T.C., J.-Y.W.
Genome-wide genotyping: CAGE: N.K., M.I.; GenSalt: J.E.H., Y.J.S.; KARE: J.-Y.L., B.-G.H., Y.S.C.; Shanghai: W.H.; SiMES/SP2: M.S., J.J.L.; Suita: N.I.; Taiwan: Y.-T.C., J.-Y.W.
Data analysis and data interpretation: N.K., F.T., T.N.K., Y.T., Y.Y.T., E.S.T., M.J.G., Y.J.K., X.S., W.T.T., R.T.H.O., C.-H.C., L.-c.C., C.E.J., J.H.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Figures 1–10 and Supplementary Tables 1–8. (PDF 2422 kb)
Rights and permissions
About this article
Cite this article
Kato, N., Takeuchi, F., Tabara, Y. et al. Meta-analysis of genome-wide association studies identifies common variants associated with blood pressure variation in east Asians. Nat Genet 43, 531–538 (2011). https://doi.org/10.1038/ng.834
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.834
This article is cited by
-
Cold-induced vasodilation response in a Japanese cohort: insights from cold-water immersion and genome-wide association studies
Journal of Physiological Anthropology (2023)
-
Comparison of cardiovascular disease risk factors among FiLWHEL (2014–2016), NNS (2013) and KNHANES (2013–2015) women
BMC Women's Health (2023)
-
CASZ1: a promising factor modulating aldosterone biosynthesis and mineralocorticoid receptor activity
Hypertension Research (2023)
-
The role of aldehyde dehydrogenase 2 in cardiovascular disease
Nature Reviews Cardiology (2023)
-
ALDH2 polymorphism rs671 and alcohol consumption: possible explanatory factors for race/ethnic differences in bone density
Osteoporosis International (2023)