Abstract
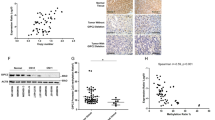
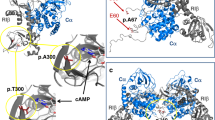
Pheochromocytomas, which are catecholamine-secreting tumors of neural crest origin, are frequently hereditary1. However, the molecular basis of the majority of these tumors is unknown2. We identified the transmembrane-encoding gene TMEM127 on chromosome 2q11 as a new pheochromocytoma susceptibility gene. In a cohort of 103 samples, we detected truncating germline TMEM127 mutations in approximately 30% of familial tumors and about 3% of sporadic-appearing pheochromocytomas without a known genetic cause. The wild-type allele was consistently deleted in tumor DNA, suggesting a classic mechanism of tumor suppressor gene inactivation. Pheochromocytomas with mutations in TMEM127 are transcriptionally related to tumors bearing NF1 mutations and, similarly, show hyperphosphorylation of mammalian target of rapamycin (mTOR) effector proteins. Accordingly, in vitro gain-of-function and loss-of-function analyses indicate that TMEM127 is a negative regulator of mTOR. TMEM127 dynamically associates with the endomembrane system and colocalizes with perinuclear (activated) mTOR, suggesting a subcompartmental-specific effect. Our studies identify TMEM127 as a tumor suppressor gene and validate the power of hereditary tumors to elucidate cancer pathogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Amar, L. et al. Genetic testing in pheochromocytoma or functional paraganglioma. J. Clin. Oncol. 23, 8812–8818 (2005).
Dahia, P.L. Evolving concepts in pheochromocytoma and paraganglioma. Curr. Opin. Oncol. 18, 1–8 (2006).
Dahia, P.L.M. et al. Novel pheochromocytoma susceptibility loci identified by integrative genomics. Cancer Res. 65, 9651–9658 (2005).
Sjöblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006).
Neumann, H.P. et al. Germ-line mutations in nonsyndromic pheochromocytoma. N. Engl. J. Med. 346, 1459–1466 (2002).
Jönsson, G. et al. High-resolution genomic profiles of breast cancer cell lines assessed by tiling BAC array comparative genomic hybridization. Genes Chromosom. Cancer 46, 543–558 (2007).
Snyder, C.M., Mardones, G.A., Ladinsky, M.S. & Howell, K.E. GMx33 associates with the trans-Golgi matrix in a dynamic manner and sorts within tubules exiting the Golgi. Mol. Biol. Cell 17, 511–524 (2006).
Dahia, P.L.M. et al. A HIF1alpha regulatory loop links hypoxia and mitochondrial signals in pheochromocytomas. PLoS Genet. 1, 72–80 (2005).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Martin, G.A. et al. The GAP-related domain of the neurofibromatosis type 1 gene product interacts with ras p21. Cell 63, 843–849 (1990).
Johannessen, C.M. et al. The NF1 tumor suppressor critically regulates TSC2 and mTOR. Proc. Natl. Acad. Sci. USA 102, 8573–8578 (2005).
Segouffin-Cariou, C. & Billaud, M. Transforming ability of MEN2A-RET requires activation of the phosphatidylinositol 3-Kinase/AKT signaling pathway. J. Biol. Chem. 275, 3568–3576 (2000).
Kwiatkowski, D.J. & Manning, B.D. Tuberous sclerosis: a GAP at the crossroads of multiple signaling pathways. Hum. Mol. Genet. 14 Spec No. 2, R251–R258 (2005).
Johannessen, C.M. et al. TORC1 is essential for NF1-associated malignancies. Curr. Biol. 18, 56–62 (2008).
Sarbassov, D.D., Guertin, D.A., Ali, S.M. & Sabatini, D.M. Phosphorylation and regulation of Akt/PKB by the rictor-mTOR complex. Science 307, 1098–1101 (2005).
Kim, D.H. et al. mTOR interacts with raptor to form a nutrient-sensitive complex that signals to the cell growth machinery. Cell 110, 163–175 (2002).
Sancak, Y. et al. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 320, 1496–1501 (2008).
Scott, K.L. et al. GOLPH3 modulates mTOR signalling and rapamycin sensitivity in cancer. Nature 459, 1085–1090 (2009).
Binda, M. et al. The Vam6 GEF controls TORC1 by activating the EGO complex. Mol. Cell 35, 563–573 (2009).
Dippold, H.C. et al. GOLPH3 bridges phosphatidylinositol-4- phosphate and actomyosin to stretch and shape the Golgi to promote budding. Cell 139, 337–351 (2009).
Ferner, R.E. et al. Guidelines for the diagnosis and management of individuals with neurofibromatosis 1. J. Med. Genet. 44, 81–88 (2007).
Schlisio, S. et al. The kinesin KIF1Bβ acts downstream from EglN3 to induce apoptosis and is a potential 1p36 tumor suppressor. Genes Dev. 22, 884–893 (2008).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−ΔΔC(T)) method. Methods 25, 402–408 (2001).
Aguiar, R.C. et al. PTPROt: an alternatively spliced and developmentally regulated B-lymphoid phosphatase that promotes G0/G1 arrest. Blood 94, 2403–2413 (1999).
Smith, P.G. et al. The phosphodiesterase PDE4B limits cAMP-associated PI3K/AKT-dependent apoptosis in diffuse large B-cell lymphoma. Blood 105, 308–316 (2005).
Sarbassov, D.D. et al. Rictor, a novel binding partner of mTOR, defines a rapamycin-insensitive and raptor-independent pathway that regulates the cytoskeleton. Curr. Biol. 14, 1296–1302 (2004).
Dahia, P.L.M. et al. PTEN is inversely correlated with the cell survival factor Akt/PKB and is inactivated via multiple mechanismsin haematological malignancies. Hum. Mol. Genet. 8, 185–193 (1999).
Inoki, K. et al. TSC2 integrates Wnt and energy signals via a coordinated phosphorylation by AMPK and GSK3 to regulate cell growth. Cell 126, 955–968 (2006).
Li, Q. et al. A syntaxin 1, Galpha(o), and N-type calcium channel complex at a presynaptic nerve terminal: analysis by quantitative immunocolocalization. J. Neurosci. 24, 4070–4081 (2004).
Acknowledgements
We thank S. Jiang for technical assistance, V. Frohlich and J. Wewer for help with optical imaging, J. Blenis for kindly providing the 293E cell line, W. Kaelin Jr. for his comments and suggestions, and M. Jech, C. Colin, N.V. Nguyen, M. Pujana, M. Vidal, D. Hill, J. Bruder and the Familial Pheochromocytoma Consortium members for their contribution to earlier phases of this project. We also thank the subjects and families that participated in the study for their cooperation. P.L.M.D. is a Kimmel Foundation Scholar and a recipient of a Voelcker Foundation Young Investigator Award and is supported by the Cancer Therapy and Research Center (CTRC) at the University of Texas Health Science Center at San Antonio (UTHSCSA) (NIH-P30 CA54174). R.C.T.A. is supported by the Voelcker Fund. Immunofluorescence images were generated in the Core Optical Imaging Facility which is supported by UTHSCSA, US National Institutes of Health (NIH)-National Cancer Institute P30 CA54174 (CTRC), NIH-National Institute on Aging (NIA) P30 AG013319 (Nathan Shock Center) and NIH-NIA P01AG19316.
Author information
Authors and Affiliations
Contributions
Y.Q., L.Y., E.E.K., K.B., R.E.L., E.S.C., J.D.L., F.S., R.A.T., R.C.T.A. and P.L.M.D. performed and analyzed experiments. M.S., N.A., F.S., F.B., G.O., R.A.T., S.P.A.T. and C.S. contributed reagents, clinical information and discussions. R.C.T.A., J.D.L. and P.L.M.D. designed experiments. R.C.T.A. and P.L.M.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–7. (PDF 2140 kb)
Rights and permissions
About this article
Cite this article
Qin, Y., Yao, L., King, E. et al. Germline mutations in TMEM127 confer susceptibility to pheochromocytoma. Nat Genet 42, 229–233 (2010). https://doi.org/10.1038/ng.533
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.533
This article is cited by
-
Single-nuclei and bulk-tissue gene-expression analysis of pheochromocytoma and paraganglioma links disease subtypes with tumor microenvironment
Nature Communications (2022)
-
N6-methyladenosine-dependent pri-miR-17-92 maturation suppresses PTEN/TMEM127 and promotes sensitivity to everolimus in gastric cancer
Cell Death & Disease (2020)
-
Master regulator analysis of paragangliomas carrying SDHx, VHL, or MAML3 genetic alterations
BMC Cancer (2019)
-
Human 5′ UTR design and variant effect prediction from a massively parallel translation assay
Nature Biotechnology (2019)
-
A nomogram for predicting the presence of germline mutations in pheochromocytomas and paragangliomas
Endocrine (2019)