Abstract
Loss of seed-coat impermeability was essential in the domestication of many leguminous crops to promote the production of their highly nutritious seeds. Here we show that seed-coat impermeability in wild soybean is controlled by a single gene, GmHs1-1, which encodes a calcineurin-like metallophosphoesterase transmembrane protein. GmHs1-1 is primarily expressed in the Malpighian layer of the seed coat and is associated with calcium content. The transition from impermeability to permeability in domesticated soybean was caused by artificial selection of a point mutation in GmHs1-1. Interestingly, a number of soybean landraces evaded selection for permeability because of an alternative selection for seed-coat cracking that also enables seed imbibition. Despite the single origin of the mutant allele Gmhs1-1, the distribution pattern of allelic variants in the context of soybean population structure and the detected signature of genomic introgression between wild and cultivated soybeans suggest that Gmhs1-1 may have experienced reselection for seed-coat permeability.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Fuller, D.Q. & Allaby, R. Seed dispersal and crop domestication: shattering, germination and seasonality in evolution under cultivation. Annual Plant Reviews 38, 238–295 (2009).
Abbo, S. et al. Plant domestication versus crop evolution: a conceptual framework for cereals and grain legumes. Trends Plant Sci. 19, 351–360 (2014).
Rolston, M.P. Water impermeable seed dormancy. Bot. Rev. 44, 365–396 (1978).
Foley, M.E. Seed dormancy: an update on terminology, physiological genetics, and quantitative trait loci regulating germinability. Weed Sci. 49, 305–317 (2001).
Ladizinsky, G. The origin of lentil and its wild gene pool. Euphytica 28, 179–187 (1979).
Gepts, P. & Debouck, D.G. In Common Beans: Research for Crop Improvement Origin, domestication, and evolution of the common bean, Phaseolus vulgaris. (eds. Voysest, O. & Van Schoonhoven, A.) pp. 7–53 (CAB International, 1991).
Abbo, S., Shtienberg, D., Lichtenzveig, J., Lev-Yadun, S. & Gopher, A. The chickpea, summer cropping, and a new model for pulse domestication in the ancient near east. Q. Rev. Biol. 78, 435–448 (2003).
Liu, B. et al. QTL Mapping of domestication-related traits in soybean (Glycine max). Ann. Bot. 100, 1027–1038 (2007).
Weeden, N.F. Genetic changes accompanying the domestication of Pisum sativum: is there a common genetic basis to the “domestication syndrome” for legumes? Ann. Bot. 100, 1017–1025 (2007).
Andargie, M., Pasquet, R.S., Gowda, B.S., Muluvi, G.M. & Timko, M.P. Molecular mapping of QTLs for domestication-related traits in cowpea (V. unguiculata (L.) Walp.). Euphytica 200, 401–412 (2014).
Keim, P., Diers, B.W. & Shoemaker, R.C. Genetic analysis of soybean hard seededness with molecular markers. Theor. Appl. Genet. 79, 465–469 (1990).
Carter, T., Nelson, R., Sneller, C. & Cui, Z. Genetic diversity in soybean. In Soybeans: Improvement Production and Uses 3rd edn. (eds. Boerma, H.R. & Specht, J.E.) pp. 303–416 (American Society of Agronomy–Crop Science Society of America–Soil Science Society of America, 2004).
Li, Y. et al. Genetic structure and diversity of cultivated soybean (Glycine max (L.) Merr.) landraces in China. Theor. Appl. Genet. 117, 857–871 (2008).
Potts, H.C., Duangpatra, J., Hairston, W.G. & Delouche, J.C. Some influences of hardseededness on soybean seed quality. Crop Sci. 18, 221–224 (1978).
Saio, K., Arai, K. & Watanabe, T. Fine structure of soybean seed coat and its changes on cooking. Cereal Science Today 18, 197–201 (1973).
Saio, K. Soybeans resistant to water absorption. Cereal Foods World 21, 168–173 (1976).
Zhang, B. et al. Quantitative trait loci mapping of seed hardness in soybean. Crop Sci. 48, 1341–1349 (2008).
Watanabe, S., Tajuddin, T., Yamanaka, N., Hayashi, M. & Harada, K. Analysis of QTLs for reproductive development and seed quality traits in soybean using recombinant inbred lines. Breed. Sci. 54, 399–407 (2004).
Schmutz, J. et al. Genome sequence of the paleopolyploid soybean (Glycine max (L.) Merr.). Nature 463, 178–183 (2010).
Kim, M.Y. et al. Whole-genome sequencing and intensive analysis of the undomesticated soybean (Glycine soja Sieb. and Zucc.) genome. Proc. Natl. Acad. Sci. USA 107, 22032–22037 (2010).
Li, Y.H. et al. De novo assembly of soybean wild relatives for pan-genome analysis of diversity and agronomic traits. Nat. Biotechnol. 32, 1045–1052 (2014).
Lush, W.M. & Evan, L.T. The seed coats of cowpeas and other grain legumes: structure in relation to function. Field Crops Res. 3, 267–286 (1980).
Miller, S.S., Bowman, L.A., Gijzen, M. & Miki, B.L.A. Early development of the seed coat of soybean (Glycine max). Ann. Bot. 84, 297–304 (1999).
Harris, W.H. Comparative ultrastructure of developing seed coats of 'hard-seeded' and 'soft-seeded' varieties of soybean, Glycine max (L.) Merril. Bot. Gaz. 148, 324–331 (1987).
Russi, L., Cocks, P.S. & Roberts, E.H. Coat thickness and hard-seededness in some Medicago and Trifolium species. Seed Sci. Res. 2, 243–249 (1992).
Valenti, G.S., Melone, L., Ferro, M. & Bozzini, A. Comparative studies on testa structure of 'hard-seeded' and 'soft-seeded' varieties of Lupinus angustifolius L. (Leguminosae) and on mechanisms of water entry. Seed Science and Technology 17, 563–581 (1989).
Kingsbury, T.J. & Cunningham, K.W. A conserved family of calcineurin regulators. Genes Dev. 14, 1595–1604 (2000).
Rusnak, F. & Mertz, P. Calcineurin: form and function. Physiol. Rev. 80, 1483–1521 (2000).
Aramburu, J., Heitman, J. & Crabtree, G.R. Calcineurin: a central controller of signaling in eukaryotes. EMBO Rep. 5, 343–348 (2004).
Zhou, Z. et al. Resequencing 302 wild and cultivated accessions identifies genes related to domestication and improvement in soybean. Nat. Biotechnol. 33, 408–414 (2015).
Hyten, D.L. et al. Impacts of genetic bottlenecks on soybean genome diversity. Proc. Natl. Acad. Sci. USA 103, 16666–16671 (2006).
Tian, Z. et al. Artificial selection for determinate growth habit in soybean. Proc. Natl. Acad. Sci. USA 107, 8563–8568 (2010).
Nakamura, T., Yang, D., Kalaiselvi, S., Uematsu, Y. & Takahashi, R. Genetic analysis of net-like cracking in soybean seed coats. Euphytica 133, 179–184 (2003).
Ma, F., Cholewa, E., Mohamed, T., Peterson, C.A. & Gijzen, M. Cracks in the palisade cuticle of soybean seed coats correlate with their permeability to water. Ann. Bot. 94, 213–228 (2004).
Ping, J. et al. Dt2 is a gain-of-function MADS-domain factor gene that specifies semideterminacy in soybean. Plant Cell 26, 2831–2842 (2014).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Flores, T. et al. Silencing of GmFAD3 gene by siRNA leads to low α-linolenic acids (18:3) of fad3-mutant phenotype in soybean [Glycine max (Merr.)]. Transgenic Res. 17, 839–850 (2008).
Shi, H. et al. BR-SIGNALING KINASE1 physically associates with FLAGELLIN SENSING2 and regulates plant innate immunity in Arabidopsis. Plant Cell 25, 1143–1157 (2013).
Yang, J. et al. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E.L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Ziegler, G. et al. Ionomic screening of field-grown soybean identifies mutants with altered seed elemental composition. Plant Genome 10.3835/plantgenome2012.07.0012 (2013).
Bradbury, P.J. et al. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23, 2633–2635 (2007).
Olsen, K.M. et al. Selection under domestication: evidence for a sweep in the rice waxy genomic region. Genetics 173, 975–983 (2006).
Bosse, M. et al. Genomic analysis reveals selection for Asian genes in European pigs following human-mediated introgression. Nat. Commun. 5, 4392 (2014).
Chia, J.M. et al. Maize HapMap2 identifies extant variation from a genome in flux. Nat. Genet. 44, 803–807 (2012).
Acknowledgements
We thank Z. Tian, J. Du, F. Lin, J. Ping, L. Zhu and A. Johnson for their involvement in developing the mapping populations at Purdue University, and we thank D. Szymanski for valuable comments on the manuscript. This work was mainly supported by soybean checkoff funds from the Indiana Soybean Alliance (00049970 and 14055567) and was partially supported by the Taishan Scholarship and High-Level Talents Foundation of Qingdao Agricultural University (631304), the North Central Soybean Research Program (58-3611-4-017), Ag Alumni Seed and the Purdue Agricultural Research Program. The US Department of Agriculture–Agricultural Research Service is an equal-opportunity provider and employer.
Author information
Authors and Affiliations
Contributions
L.S., R.L.N. and J.M. designed the research; L.S., Z.M., C.C., D.Z., M.Z., Y.W., X.Z., S.A.S. and L.Z. performed the research; L.S., Z.M., C.C., Z.J.Z. and J.M. analyzed the data; J.M. wrote the manuscript with input from L.S., S.A.S., Z.J.Z. and R.L.N.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
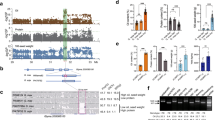
Supplementary Figure 1 Expression of Glyma02g43700.1 and Glyma02g43710.1 in the mapped GmHs1-1 region in seed coats 4 weeks post-anthesis (WPA).
Expression levels of genes relative to that of the actin-expressing gene in PI 468916 and Williams 82 and in the F1:2 seed coats were detected by qRT-PCR. Expression levels are shown as mean ± s.e.m. from three replicates.
Supplementary Figure 2 Structural prediction of the GmHs1-1 protein.
(a) Posterior probabilities for transmembrane helix, inside, and outside displayed for the GmHs1-1 protein as determined by TMHMM 2.0. Two transmembrane segments were predicted. The asterisk indicates the location of the amino acid change resulting from the C>T mutation in GmHs1-1 that resulted in the Gmhs1-1 allele. (b,c) Predicted three-dimensional structures of the GmHs1-1 and Gmhs1-1 proteins, which differ at a single amino acid from threonine to methionine resulting from the C>T mutation. Arrows indicate the predicted structural differences around the mutation sites between the GmHs1-1 and Gmhs1-1 proteins.
Supplementary Figure 3 Comparison of the promoter sequences of the GmHs1-1 locus between G. max and G. soja.
(a) Phylogenetic relationships of the G. soja and G. max accessions whose promoter and CDS regions at the GmHs1-1 locus were completely sequenced in this study and which were also previously re-sequenced30. The phylogenetic tree of the 302 re-sequenced accessions was built with the genome-wide SNPs detected by resequencing30. (b) Sequence variation in the complete 2.5-kb genomic region upstream of the GmHs1-1 CDS. The positions of variation sites upstream of the start codon are labeled with negative numbers. The portions of identical sequences within the promoter region shared by all these accessions are indicated by dots. Accessions sharing an identical promoter are highlighted. Stars indicate the landraces showing seed-coat cracking. "H", hard seededness; "P", permeable seed coat; "P/C", permeable seed coat due to seed-coat cracking; "In", insertion; "Del", deletion.
Supplementary Figure 4 Distribution of six landraces showing seed-coat cracking in the context of the genetic structure of a soybean population.
Each vertical bar represents a single soybean accession and is partitioned into K (3) colored segments. Each color represents one cluster, and the length of the colored segment shows the accession’s estimated proportion of membership in that cluster as calculated by STRUCTURE in a typical run at the value of K (3). The genotypes of individual cultivars are marked by thin vertical lines above the plot of population structure, and their phenotypes are indicated by solid (hard-seededness) and hollow (seed-coat permeability) diamonds and stars (seed-coat cracking), respectively. The detailed information of this population is shown in Supplementary Table 3. The population structure was constructed on the basis of genetic diversity revealed by sequences from 102 genes as described previously31,32.
Supplementary Figure 5 Distribution of the wild-type GmHs1-1 allele in a mini-core collection of Chinese soybean landraces.
(a) Genetic distribution of the GmHs1-1 allele in the context of the genetic structure of the landrace population. Each vertical bar represents a single soybean landrace and is partitioned into K (3) colored segments. Each color represents one cluster, and the length of the colored segment shows the landrace’s estimated proportion of membership in that cluster as calculated by STRUCTURE in a typical run at the value of K (3). Abbreviations above brackets indicate soybean-cultivated eco-regions or subregions in China. CSp, Changjiang (Yangtze river) spring-type subregion; HR, Huang-Huai region; HSp, Huang-Huai spring-type subregion; HSu, Huang-Hai summer-type subregion; NESp, Northeast spring-type subregion; NR, North region; NSp, North spring-type subregion; Sau, South autumn planting ecotype subregion; SR, South region; SSp, South spring-type subregion; Ssu, South summer-type subregion. (b) Geographic distribution of the GmHs1-1 allele carried by landraces in the mini-core collection in China. The detailed information of this population is shown in Supplementary Table 4. The population structure was described previously13,32.
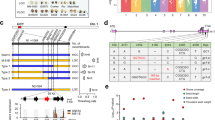
Supplementary Figure 6 Selective sweep in the GmHs1-1 region and putative genomic introgression.
(a) An illustration of a selective sweep that spans an ~160-kb region that harbors 19 genes including the GmHs1-1 locus. The red arrowhead points to the GmHs1-1 locus. Each blue bar represents one gene. The smallest bars are equivalent to a ratio of 0. The order of and distances among the bars indicate the order of the gene in the region but do not reflect the sizes of the genes or intergenic distances. (b) Ratios of the nucleotide diversity among the Gmhs1-1 accessions (πGmhs1-1) to the nucleotide diversity among the GmHs1-1 accessions (πGmHs1-1) estimated for all genes on chromosome 2. Among the 109 accessions (Supplementary Table 3) genotyped for the GmHs1-1 locus by CAPS markers and sequencing of DNA fragments in this study, a total of 87 (18 G. soja accessions and 69 G. max accessions) were resequenced30 and used for genome-wide genetic diversity analysis. All SNPs in genetic sequences along chromosome 2 were included in the analysis of genetic diversity. The blue vertical lines represent individual genes and their orders along chromosome 2. Sizes of individual genes and intergenic distances are not reflected in the plots. The red box marks a genomic region comprising 25 genes upstream and 25 genes downstream of the GmHs1-1 locus, and the two vertical pink lines beneath the red box indicate the ~160-kb selective sweep. (c–f) Putative introgression of genomic segments from G. soja into each of the four resequenced landraces carrying GmHs1-1 along chromosome 2. Two vertical pink lines indicate the ~160-kb genomic region showing the selective sweep. No clear genomic introgression was detected in the ~160-kb selective sweep region, but signatures of genomic introgression were detected in its flanking regions in three of the four landraces. (g–j) Exemplification of putative introgression of genomic segments from G. soja into each of the four resequenced landraces on other chromosomes. In each of the panels from c to j, the x axis shows the full length of a specific chromosome, and the y axis represents relative identicals by descent (rIBD) that reflect the relative frequency of the shared haplotypes between each of the four landraces and the 18 G. soja accessions versus the shared haplotypes between each of the four landraces and the 69 G. soja accessions along individual chromosomes (Online Methods). All SNPs detected by genome resequencing30 were used to identify putative genomic introgression.
Supplementary Figure 7 Comparison of linkage disequilibrium (LD) in the genomic region surrounding the selective sweep that harbors the GmHs1-1 locus.
(a) LD in the G. max subpopulation. (b) A genomic region surrounding the selective sweep as shown in Supplementary Fig. 6a. (c) LD in the G. max subpopulation. Dashed gray lines connecting panels a and b and panels b and c mark the same regions. The red dashed lines indicate the position of the GmHs1-1 locus. All SNPs from 87 resequenced accessions30 were used for LD analysis. Strong LD was detected in the selective sweep region in the G. max subpopulation, whereas no clear LD was detected in the corresponding region in the G. soja subpopulation. The causative (C>T) mutation for the transition from GmHs1-1 to Gmhs1-1 was not detected from the resequencing data30; thus the potential LD between the C>T mutation and other SNPs in the region is not reflected.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–6 (PDF 1618 kb)
Rights and permissions
About this article
Cite this article
Sun, L., Miao, Z., Cai, C. et al. GmHs1-1, encoding a calcineurin-like protein, controls hard-seededness in soybean. Nat Genet 47, 939–943 (2015). https://doi.org/10.1038/ng.3339
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3339
This article is cited by
-
An omics strategy increasingly improves the discovery of genetic loci and genes for seed-coat color formation in soybean
Molecular Breeding (2023)
-
Accelerating crop domestication through genome editing for sustainable agriculture
Journal of Plant Biochemistry and Biotechnology (2023)
-
Identification of KASP markers and candidate genes for drought tolerance in wheat using 90K SNP array genotyping of near-isogenic lines targeting a 4BS quantitative trait locus
Theoretical and Applied Genetics (2023)
-
Development of a versatile resource for post-genomic research through consolidating and characterizing 1500 diverse wild and cultivated soybean genomes
BMC Genomics (2022)
-
Improved pea reference genome and pan-genome highlight genomic features and evolutionary characteristics
Nature Genetics (2022)