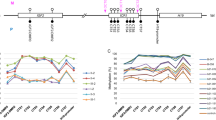
Abstract
Constitutional abnormalities at the imprinted 11p15 growth regulatory region cause syndromes characterized by disordered growth, some of which include a risk of Wilms tumor1,2,3. We explored their possible contribution to nonsyndromic Wilms tumor and identified constitutional 11p15 abnormalities in genomic lymphocyte DNA from 13 of 437 individuals (3%) with sporadic Wilms tumor without features of growth disorders, including 12% of bilateral cases (P = 0.001) and in one familial Wilms tumor pedigree. No abnormality was detected in 220 controls (P = 0.006). Abnormalities identified included H19 DMR epimutations, uniparental disomy 11p15 and H19 DMR imprinting center mutations (one microinsertion and one microdeletion), thus identifying microinsertion as a new class of imprinting center mutation. Our data identify constitutional 11p15 defects as one of the most common known causes of Wilms tumor, provide mechanistic insights into imprinting disruption and reveal clinically important epigenotype-phenotype associations. The impact on clinical management dictates that constitutional 11p15 analysis should be considered in all individuals with Wilms tumor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Gicquel, C. et al. Epimutation of the telomeric imprinting center region on chromosome 11p15 in Silver-Russell syndrome. Nat. Genet. 37, 1003–1007 (2005).
Weksberg, R., Shuman, C. & Smith, A.C. Beckwith-Wiedemann syndrome. Am. J. Med. Genet. C. Semin. Med. Genet. 137, 12–23 (2005).
Scott, R.H., Stiller, C.A., Walker, L.L. & Rahman, N. Syndromes and constitutional chromosomal abnormalities associated with Wilms tumour. J. Med. Genet. 43, 705–715 (2006).
Little, S.E. et al. Frequency and heritability of WT1 mutations in nonsyndromic Wilms' tumor patients: a UK Children's Cancer Study Group Study. J. Clin. Oncol. 22, 4140–4146 (2004).
Reik, W. et al. Imprinting mutations in the Beckwith-Wiedemann syndrome suggested by altered imprinting pattern in the IGF2–H19 domain. Hum. Mol. Genet. 4, 2379–2385 (1995).
Niemitz, E. et al. Microdeletion of LIT1 in familial Beckwith-Wiedemann syndrome. Am. J. Hum. Genet. 75, 844–849 (2004).
Sparago, A. et al. Microdeletions in the human H19 DMR result in loss of IGF2 imprinting and Beckwith-Wiedemann syndrome. Nat. Genet. 36, 958–960 (2004).
Prawitt, D. et al. Microdeletion of target sites for insulator protein CTCF in a chromosome 11p15 imprinting center in Beckwith-Wiedemann syndrome and Wilms' tumor. Proc. Natl. Acad. Sci. USA 102, 4085–4090 (2005).
Prawitt, D. et al. Microdeletions in the human H19 DMR result in loss of IGF2 imprinting and Beckwith-Wiedemann syndrome. Nat. Genet. 37, 785–787 (2005).
Sparago, A. et al. Mechanisms causing imprinting defects in familial Beckwith-Wiedemann syndrome with Wilms' tumour. Hum. Mol. Genet. 16, 254–264 (2007).
Rainier, S. et al. Relaxation of imprinted genes in human cancer. Nature 362, 747–749 (1993).
Ogawa, O. et al. Relaxation of insulin-like growth factor II gene imprinting implicated in Wilms' tumour. Nature 362, 749–751 (1993).
Moulton, T. et al. Epigenetic lesions at the H19 locus in Wilms' tumour patients. Nat. Genet. 7, 440–447 (1994).
Steenman, M.J. et al. Loss of imprinting of IGF2 is linked to reduced expression and abnormal methylation of H19 in Wilms' tumour. Nat. Genet. 7, 433–439 (1994).
Okamoto, K. et al. Epigenetic changes at the insulin-like growth factor II/H19 locus in developing kidney is an early event in Wilms tumorigenesis. Proc. Natl. Acad. Sci. USA 94, 5367–5371 (1997).
Scott, R.H. et al. Methylation-specific multiplex ligation-dependent probe amplification (MS-MLPA) robustly detects and distinguishes 11p15 abnormalities associated with overgrowth and growth retardation. J. Med. Genet. 45, 106–113 (2008).
Buiting, K. et al. Inherited microdeletions in the Angelman and Prader-Willi syndromes define an imprinting centre on human chromosome 15. Nat. Genet. 9, 395–400 (1995).
Bastepe, M. et al. Deletion of the NESP55 differentially methylated region causes loss of maternal GNAS imprints and pseudohypoparathyroidism type Ib. Nat. Genet. 37, 25–27 (2005).
Kagami, M. et al. Deletions and epimutations affecting the human 14q32.2 imprinted region in individuals with paternal and maternal upd(14)-like phenotypes. Nat. Genet. 40, 237–242 (2008).
Porteus, M.H. et al. Characteristics and outcome of children with Beckwith-Wiedemann syndrome and Wilms' tumor: a report from the National Wilms Tumor Study Group. J. Clin. Oncol. 18, 2026–2031 (2000).
Beckwith, J.B. Kiviat, N.B. & Bonadio, J.F. Nephrogenic rests, nephroblastomatosis, and the pathogenesis of Wilms' tumor. Pediatr. Pathol. 10, 1–36 (1990).
Ferner, R.E. et al. Guidelines for the diagnosis and management of individuals with neurofibromatosis 1. J. Med. Genet. 44, 81–88 (2007).
Tatton-Brown, K. et al. Genotype-phenotype associations in Sotos syndrome: an analysis of 266 individuals with NSD1 aberrations. Am. J. Hum. Genet. 77, 193–204 (2005).
Engel, J.R. et al. Epigenotype-phenotype correlations in Beckwith-Wiedemann syndrome. J. Med. Genet. 37, 921–926 (2000).
Gaston, V. et al. Analysis of the methylation status of the KCNQ1OT and H19 genes in leukocyte DNA for the diagnosis and prognosis of Beckwith-Wiedemann syndrome. Eur. J. Hum. Genet. 9, 409–418 (2001).
Weksberg, R. et al. Tumor development in the Beckwith-Wiedemann syndrome is associated with a variety of constitutional molecular 11p15 alterations including imprinting defects of KCNQ1OT1. Hum. Mol. Genet. 10, 2989–3000 (2001).
DeBaun, M.R. et al. Epigenetic alterations of H19 and LIT1 distinguish patients with Beckwith-Wiedemann syndrome with cancer and birth defects. Am. J. Hum. Genet. 70, 604 (2002).
Bliek, J. et al. Epigenotyping as a tool for the prediction of tumor risk and tumor type in patients with Beckwith-Wiedemann syndrome (BWS). J. Pediatr. 145, 796–799 (2004).
Cooper, W.N. et al. Molecular subtypes and phenotypic expression of Beckwith-Wiedemann syndrome. Eur. J. Hum. Genet. 13, 1025–1032 (2005).
Scott, R.H. et al. Surveillance for Wilms tumour in at-risk children: pragmatic recommendations for best practice. Arch. Dis. Child. 91, 995–999 (2006).
Acknowledgements
We thank the affected individuals and families involved in the research. We thank the many physicians and nurses who referred families and provided samples and clinical information. The following members of the FACT Collaboration provided familial Wilms tumor pedigrees analyzed in this study: L. Arbour, C. Bonaïti-Pellié, L. Cannon-Albright, A. Chompret, T. Cole, C. Dhooge, W. Dupuis, A. Foot, W. Foulkes, H. Galvin, A. Gnekow, N. Graf, D. King, J. Kingston, I. Lewis, A. O'Meara, F. Millot, H. Price, B. Royer-Pokora, V. Schumacher, C. Schwartz, R. Shannon, E. Sheridan, J. Skeen, P. Tonin and A. Weirich. We thank B. Messahel, R. Al-Saadi, M. Cohen, N. Coleman and M. Malone for providing Wilms tumor specimens. We thank M. Stratton and C. Turnbull for their comments on the manuscript. The research was carried out as part of the Investigation of Familial Wilms Tumour Genes and the Factors Associated with Childhood Tumours (FACT) study, which are UK Children's Cancer and Leukaemia Group (CCLG) studies. The Childhood Cancer Research Group receives funding from the Department of Health and the Scottish Ministers. The views expressed in this publication are those of the authors and not necessarily those of the Department of Health and the Scottish Ministers. R.H.S. is supported by the Michael and Betty Kadoorie Cancer Genetics Research Programme. This work was supported by grants from Medical Research Council (UK), Cancer Research UK and the Institute of Cancer Research (UK).
Author information
Authors and Affiliations
Consortia
Contributions
The study was designed by R.H.S. and N.R. The molecular analyses were performed by R.H.S., J.D., L.B., K.B. and S.H. Samples and clinical information were provided by members of the FACT Collaboration, R.H.S., A.C., M.G., J.A.K., G.A.L., S.P., B.P., M.D.R., D.W., J.A.C., P.P., E.R.M., J.M.B., C.A.S., K.P.-J. and N.R. and coordinated by N.H. The manuscript was written by R.H.S. and N.R., with input from the other authors.
Corresponding author
Additional information
A full list of members is provided in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–3, Supplementary Figures 1–3 (PDF 897 kb)
Rights and permissions
About this article
Cite this article
Scott, R., Douglas, J., Baskcomb, L. et al. Constitutional 11p15 abnormalities, including heritable imprinting center mutations, cause nonsyndromic Wilms tumor. Nat Genet 40, 1329–1334 (2008). https://doi.org/10.1038/ng.243
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.243
This article is cited by
-
Cancer predisposition signaling in Beckwith-Wiedemann Syndrome drives Wilms tumor development
British Journal of Cancer (2024)
-
Hallmark discoveries in the biology of Wilms tumour
Nature Reviews Urology (2024)
-
Imprinting disorders
Nature Reviews Disease Primers (2023)
-
Genetic and epigenetic features of bilateral Wilms tumor predisposition in patients from the Children’s Oncology Group AREN18B5-Q
Nature Communications (2023)
-
Methylation Statuses of H19DMR and KvDMR at WT2 in Wilms Tumors in Taiwan
Pathology & Oncology Research (2020)