Abstract
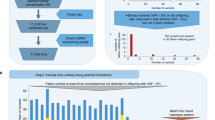
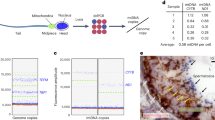
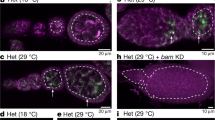
A genetic bottleneck explains the marked changes in mitochondrial DNA (mtDNA) heteroplasmy that are observed during the transmission of pathogenic mutations, but the precise timing of these changes remains controversial, and it is not clear whether selection has a role. These issues are important for the genetic counseling of prospective mothers and for the development of treatments aimed at disease prevention. By studying mice transmitting a heteroplasmic single-base-pair deletion in the mitochondrial tRNAMet gene, we show that the extent of mammalian mtDNA heteroplasmy is principally determined prenatally within the developing female germline. Although we saw no evidence of mtDNA selection prenatally, skewed heteroplasmy levels were observed in the offspring of the next generation, consistent with purifying selection. High percentages of mtDNA genomes with the tRNAMet mutation were linked to a compensatory increase in overall mitochondrial RNA levels, ameliorating the biochemical phenotype and explaining why fecundity is not compromised.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Upholt, W.B. & Dawid, I.B. Mapping of mitochondrial DNA of individual sheep and goats: rapid evolution in the D loop region. Cell 11, 571–583 (1977).
Olivo, P.D., Van de Walle, M.J., Laipis, P.J. & Hauswirth, W.W. Nucleotide sequence evidence for rapid genotypic shifts in the bovine mitochondrial DNA D-loop. Nature 306, 400–402 (1983).
Cao, L. et al. The mitochondrial bottleneck occurs without reduction of mtDNA content in female mouse germ cells. Nat. Genet. 39, 386–390 (2007).
Cree, L.M. et al. A reduction of mitochondrial DNA molecules during embryogenesis explains the rapid segregation of genotypes. Nat. Genet. 40, 249–254 (2008).
Wai, T., Teoli, D. & Shoubridge, E.A. The mitochondrial DNA genetic bottleneck results from replication of a subpopulation of genomes. Nat. Genet. 40, 1484–1488 (2008).
Wonnapinij, P., Chinnery, P.F. & Samuels, D.C. Previous estimates of mitochondrial DNA mutation level variance did not account for sampling error: comparing the mtDNA genetic bottleneck in mice and humans. Am. J. Hum. Genet. 86, 540–550 (2010).
Samuels, D.C., Wonnapinij, P., Cree, L.M. & Chinnery, P.F. Reassessing evidence for a postnatal mitochondrial genetic bottleneck. Nat. Genet. 42, 471–472 (2010).
Chinnery, P.F. et al. The inheritance of mitochondrial DNA heteroplasmy: random drift, selection or both? Trends Genet. 16, 500–505 (2000).
Uusimaa, J. et al. Prevalence, segregation, and phenotype of the mitochondrial DNA 3243A>G mutation in children. Ann. Neurol. 62, 278–287 (2007).
Trifunovic, A. et al. Premature ageing in mice expressing defective mitochondrial DNA polymerase. Nature 429, 417–423 (2004).
Stewart, J.B. et al. Strong purifying selection in transmission of mammalian mitochondrial DNA. PLoS Biol. 6, e10 (2008).
Stewart, J.B., Freyer, C., Elson, J.L. & Larsson, N.G. Purifying selection of mtDNA and its implications for understanding evolution and mitochondrial disease. Nat. Rev. Genet. 9, 657–662 (2008).
Manfredi, G. et al. Identification of a mutation in the mitochondrial tRNACys gene associated with mitochondrial encephalopathy. Hum. Mutat. 7, 158–163 (1996).
Santorelli, F.M. et al. Mitochondrial tRNACys gene mutation (A5814G): a second family with mitochondrial encephalopathy. Neuromuscul. Disord. 7, 156–159 (1997).
Wonnapinij, P., Chinnery, P.F. & Samuels, D.C. The distribution of mitochondrial DNA heteroplasmy due to random genetic drift. Am. J. Hum. Genet. 83, 582–593 (2008).
Payer, B. et al. Generation of stella-GFP transgenic mice: a novel tool to study germ cell development. Genesis 44, 75–83 (2006).
Fan, W. et al. A mouse model of mitochondrial disease reveals germline selection against severe mtDNA mutations. Science 319, 958–962 (2008).
Metodiev, M.D. et al. Methylation of 12S rRNA is necessary for in vivo stability of the small subunit of the mammalian mitochondrial ribosome. Cell Metab. 9, 386–397 (2009).
Cámara, Y. et al. MTERF4 regulates translation by targeting the methyltransferase NSUN4 to the mammalian mitochondrial ribosome. Cell Metab. 13, 527–539 (2011).
Boulet, L., Karpati, G. & Shoubridge, E.A. Distribution and threshold expression of the tRNALys mutation in skeletal muscle of patients with myoclonic epilepsy and ragged-red fibers (MERRF). Am. J. Hum. Genet. 51, 1187–1200 (1992).
Brown, D.T., Samuels, D.C., Michael, E.M., Turnbull, D.M. & Chinnery, P.F. Random genetic drift determines the level of mutant mitochondrial DNA in human primary oocytes. Am. J. Hum. Genet. 68, 533–536 (2001).
Freyer, C. et al. Maintenance of respiratory chain function in mouse hearts with severely impaired mtDNA transcription. Nucleic Acids Res. 38, 6577–6588 (2010).
Schägger, H. & von Jagow, G. Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal. Biochem. 199, 223–231 (1991).
Acknowledgements
We would like to thank C. Göttlinger and G. Rappl for help with the fluorescence-activated cell sorting (FACS) analysis. T.W. is supported by a long-term Human Frontier Science Program (HFSP) fellowship. P.F.C. is a Wellcome Trust Senior Fellow in Clinical Science and a National Institute for Health Research (NIHR) Senior Investigator, who also receives funding from the Wellcome Centre for Mitochondrial Research, the Medical Research Council (UK) Translational Muscle Centre, the UK NIHR Biomedical Research Centre for Ageing and Age-Related Disease and NIHR Dementia Biomedical Research Unit awards to the Newcastle upon Tyne Foundation Hospitals National Health Service (NHS) Trust. N.-G.L. is supported by the Swedish Research Council, the Leducq foundation and a European Research Council Advanced Investigator grant.
Author information
Authors and Affiliations
Contributions
P.F.C., C.F., L.M.C. and N.-G.L. conceived the study and supervised the laboratory work, which was carried out by C.F., L.M.C., A.M., J.B.S., C.K., V.I.F., D.M., T.W., E.H., R.J.W. and E.E.C. D.C.S. performed the statistical analysis. P.F.C. wrote the manuscript with C.F., which was modified after critical appraisal from the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Supplementary Tables 1–3 and 5 (PDF 25942 kb)
Supplementary Table 4
Raw heteroplasmy data from the mothers, oogonia, oocytes and offspring. See excel file. (XLSX 26 kb)
Rights and permissions
About this article
Cite this article
Freyer, C., Cree, L., Mourier, A. et al. Variation in germline mtDNA heteroplasmy is determined prenatally but modified during subsequent transmission. Nat Genet 44, 1282–1285 (2012). https://doi.org/10.1038/ng.2427
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2427
This article is cited by
-
Modulating Mitochondrial DNA Heteroplasmy with Mitochondrially Targeted Endonucleases
Annals of Biomedical Engineering (2022)
-
Oxygen tension modulates the mitochondrial genetic bottleneck and influences the segregation of a heteroplasmic mtDNA variant in vitro
Communications Biology (2021)
-
Extreme heterogeneity of human mitochondrial DNA from organelles to populations
Nature Reviews Genetics (2021)
-
Cell competition acts as a purifying selection to eliminate cells with mitochondrial defects during early mouse development
Nature Metabolism (2021)
-
Step-wise elimination of α-mitochondrial nucleoids and mitochondrial structure as a basis for the strict uniparental inheritance in Cryptococcus neoformans
Scientific Reports (2020)