Abstract
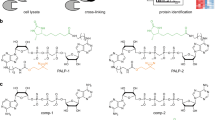
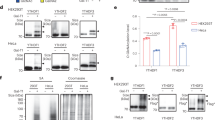
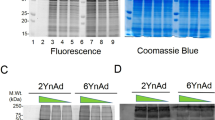
ADP-ribosylation is a post-translational modification that is known to be involved in cellular homeostasis and stress but has been challenging to analyze biochemically. To facilitate the detection of ADP-ribosylated proteins, we show that an alkyne–adenosine analog, N6-propargyl adenosine (N6pA), is metabolically incorporated in mammalian cells and enables fluorescence detection and proteomic analysis of ADP-ribosylated proteins. Notably, our analysis of N6pA-labeled proteins that are upregulated by oxidative stress revealed differential ADP-ribosylation of small GTPases. We discovered that oxidative stress induced ADP-ribosylation of Hras on Cys181 and Cys184 in the C-terminal hypervariable region, which are normally S-fatty-acylated. Downstream Hras signaling is impaired by ADP-ribosylation during oxidative stress, but is rescued by ADP-ribosyltransferase inhibitors. Our study demonstrates that ADP-ribosylation of small GTPases not only is mediated by bacterial toxins but is endogenously regulated in mammalian cells. N6pA provides a useful tool to characterize ADP-ribosylated proteins and their regulatory mechanisms in cells.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Chambon, P., Weill, J.D. & Mandel, P. Nicotinamide mononucleotide activation of new DNA-dependent polyadenylic acid synthesizing nuclear enzyme. Biochem. Biophys. Res. Commun. 11, 39–43 (1963).
Leung, A.K.L. Poly(ADP-ribose): an organizer of cellular architecture. J. Cell Biol. 205, 613–619 (2014).
Kraus, W.L. PARPs and ADP-ribosylation: 50 years ... and counting. Mol. Cell 58, 902–910 (2015).
Barkauskaite, E., Jankevicius, G. & Ahel, I. Structures and mechanisms of enzymes employed in the synthesis and degradation of PARP-dependent protein ADP-ribosylation. Mol. Cell 58, 935–946 (2015).
Lord, C.J. & Ashworth, A. The DNA damage response and cancer therapy. Nature 481, 287–294 (2012).
Meder, V.S., Boeglin, M., de Murcia, G. & Schreiber, V. PARP-1 and PARP-2 interact with nucleophosmin/B23 and accumulate in transcriptionally active nucleoli. J. Cell Sci. 118, 211–222 (2005).
Smith, S. & de Lange, T. Tankyrase promotes telomere elongation in human cells. Curr. Biol. 10, 1299–1302 (2000).
Gagné, J.-P. et al. Proteome-wide identification of poly(ADP-ribose) binding proteins and poly(ADP-ribose)-associated protein complexes. Nucleic Acids Res. 36, 6959–6976 (2008).
Cho, S.H. et al. PARP-14, a member of the B aggressive lymphoma family, transduces survival signals in primary B cells. Blood 113, 2416–2425 (2009).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Westcott, N.P. & Hang, H.C. Chemical reporters for exploring ADP-ribosylation and AMPylation at the host-pathogen interface. Curr. Opin. Chem. Biol. 23, 56–62 (2014).
Honjo, T., Nishizuka, Y. & Hayaishi, O. Diphtheria toxin-dependent adenosine diphosphate ribosylation of aminoacyl transferase II and inhibition of protein synthesis. J. Biol. Chem. 243, 3553–3555 (1968).
Ganesan, A.K., Vincent, T.S., Olson, J.C. & Barbieri, J.T. Pseudomonas aeruginosa exoenzyme S disrupts Ras-mediated signal transduction by inhibiting guanine nucleotide exchange factor-catalyzed nucleotide exchange. J. Biol. Chem. 274, 21823–21829 (1999).
Aktories, K. Bacterial protein toxins that modify host regulatory GTPases. Nat. Rev. Microbiol. 9, 487–498 (2011).
Williams, G.T., Shall, S. & Ford, C.C. Direct radioactive labelling of poly(ADP-ribose) in developing Xenopus laevis embryos. Biosci. Rep. 3, 461–467 (1983).
Jungmichel, S. et al. Proteome-wide identification of poly(ADP-Ribosyl)ation targets in different genotoxic stress responses. Mol. Cell 52, 272–285 (2013).
Zhang, Y., Wang, J., Ding, M. & Yu, Y. Site-specific characterization of the Asp- and Glu-ADP-ribosylated proteome. Nat. Methods 10, 981–984 (2013).
Morgan, R.K. & Cohen, M.S. A clickable aminooxy probe for monitoring cellular ADP-ribosylation. ACS Chem. Biol. 10, 1778–1784 (2015).
Jiang, H., Kim, J.H., Frizzell, K.M., Kraus, W.L. & Lin, H. Clickable NAD analogues for labeling substrate proteins of poly(ADP-ribose) polymerases. J. Am. Chem. Soc. 132, 9363–9372 (2010).
Carter-O'Connell, I. et al. Identifying family-member-specific targets of mono-ARTDs by using a chemical genetics approach. Cell Rep. 14, 621–631 (2016).
Wallrodt, S., Buntz, A., Wang, Y., Zumbusch, A. & Marx, A. Bioorthogonally functionalized NAD(+) analogues for in-cell visualization of poly(ADP-ribose) formation. Angew. Chem. Int. Ed. Engl. 55, 7660–7664 (2016).
Gibson, B.A. et al. Chemical genetic discovery of PARP targets reveals a role for PARP-1 in transcription elongation. Science 353, 45–50 (2016).
Daniels, C.M., Ong, S.E. & Leung, A.K. The promise of proteomics for the study of ADP-ribosylation. Mol. Cell 58, 911–924 (2015).
Grammel, M. & Hang, H.C. Chemical reporters for biological discovery. Nat. Chem. Biol. 9, 475–484 (2013).
Grammel, M., Hang, H. & Conrad, N.K. Chemical reporters for monitoring RNA synthesis and poly(A) tail dynamics. ChemBioChem 13, 1112–1115 (2012).
Chapman, J.D., Gagné, J.-P., Poirier, G.G. & Goodlett, D.R. Mapping PARP-1 auto-ADP-ribosylation sites by liquid chromatography-tandem mass spectrometry. J. Proteome Res. 12, 1868–1880 (2013).
Daniels, C.M., Ong, S.-E. & Leung, A.K.L. Phosphoproteomic approach to characterize protein mono- and poly(ADP-ribosyl)ation sites from cells. J. Proteome Res. 13, 3510–3522 (2014).
Ogata, N., Ueda, K. & Hayaishi, O. ADP-ribosylation of histone H2B. Identification of glutamic acid residue 2 as the modification site. J. Biol. Chem. 255, 7610–7615 (1980).
Kanda, T., Sullivan, K.F. & Wahl, G.M. Histone-GFP fusion protein enables sensitive analysis of chromosome dynamics in living mammalian cells. Curr. Biol. 8, 377–385 (1998).
Wahlberg, E. et al. Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors. Nat. Biotechnol. 30, 283–288 (2012).
Leung, A.K.L. et al. Poly(ADP-ribose) regulates stress responses and microRNA activity in the cytoplasm. Mol. Cell 42, 489–499 (2011).
Choudhury, A. et al. Rab proteins mediate Golgi transport of caveola-internalized glycosphingolipids and correct lipid trafficking in Niemann-Pick C cells. J. Clin. Invest. 109, 1541–1550 (2002).
Roth, A.F. et al. Global analysis of protein palmitoylation in yeast. Cell 125, 1003–1013 (2006).
Ahearn, I.M., Haigis, K., Bar-Sagi, D. & Philips, M.R. Regulating the regulator: post-translational modification of RAS. Nat. Rev. Mol. Cell Biol. 13, 39–51 (2011).
Percher, A. et al. Mass-tag labeling reveals site-specific and endogenous levels of protein S-fatty acylation. Proc. Natl. Acad. Sci. USA 113, 4302–4307 (2016).
Wan, J., Roth, A.F., Bailey, A.O. & Davis, N.G. Palmitoylated proteins: purification and identification. Nat. Protoc. 2, 1573–1584 (2007).
Yount, J.S. et al. Palmitoylome profiling reveals S-palmitoylation-dependent antiviral activity of IFITM3. Nat. Chem. Biol. 6, 610–614 (2010).
Avraham, R. & Yarden, Y. Feedback regulation of EGFR signalling: decision making by early and delayed loops. Nat. Rev. Mol. Cell Biol. 12, 104–117 (2011).
Wu, R.-F., Ma, Z., Liu, Z. & Terada, L.S. Nox4-derived H2O2 mediates endoplasmic reticulum signaling through local Ras activation. Mol. Cell. Biol. 30, 3553–3568 (2010).
Choudhary, S., Wang, K.-K.A. & Wang, H.-C.R. Oncogenic H-Ras, FK228, and exogenous H2O2 cooperatively activated the ERK pathway in selective induction of human urinary bladder cancer J82 cell death. Mol. Carcinog. 50, 215–219 (2011).
Kolch, W. Coordinating ERK/MAPK signalling through scaffolds and inhibitors. Nat. Rev. Mol. Cell Biol. 6, 827–837 (2005).
Kauppinen, T.M., Suh, S.W., Berman, A.E., Hamby, A.M. & Swanson, R.A. Inhibition of poly(ADP-ribose) polymerase suppresses inflammation and promotes recovery after ischemic injury. J. Cereb. Blood Flow Metab. 29, 820–829 (2009).
West, R.E. Jr., Moss, J., Vaughan, M., Liu, T. & Liu, T.Y. Pertussis toxin-catalyzed ADP-ribosylation of transducin. Cysteine 347 is the ADP-ribose acceptor site. J. Biol. Chem. 260, 14428–14430 (1985).
Bivona, T.G. et al. PKC regulates a farnesyl-electrostatic switch on K-Ras that promotes its association with Bcl-XL on mitochondria and induces apoptosis. Mol. Cell 21, 481–493 (2006).
Ahearn, I.M. et al. FKBP12 binds to acylated H-ras and promotes depalmitoylation. Mol. Cell 41, 173–185 (2011).
Goodwin, J.S. et al. Depalmitoylated Ras traffics to and from the Golgi complex via a nonvesicular pathway. J. Cell Biol. 170, 261–272 (2005).
Burgoyne, J.R. et al. Oxidation of HRas cysteine thiols by metabolic stress prevents palmitoylation in vivo and contributes to endothelial cell apoptosis. FASEB J. 26, 832–841 (2012).
Evangelista, A.M., Kohr, M.J. & Murphy, E. S-nitrosylation: specificity, occupancy, and interaction with other post-translational modifications. Antioxid. Redox Signal. 19, 1209–1219 (2013).
Ho, G.P.H. et al. S-nitrosylation and S-palmitoylation reciprocally regulate synaptic targeting of PSD-95. Neuron 71, 131–141 (2011).
Leonard, S.E., Reddie, K.G. & Carroll, K.S. Mining the thiol proteome for sulfenic acid modifications reveals new targets for oxidation in cells. ACS Chem. Biol. 4, 783–799 (2009).
Nguyen, T. & Francis, M.B. Practical synthetic route to functionalized rhodamine dyes. Org. Lett. 5, 3245–3248 (2003).
Grammel, M., Luong, P., Orth, K. & Hang, H.C. A chemical reporter for protein AMPylation. J. Am. Chem. Soc. 133, 17103–17105 (2011).
Acknowledgements
N.P.W. is a Leukemia and Lymphoma Society postdoctoral fellow. The authors thank R. Pagano (Mayo Clinic) for GFP–HA–Rab7, G. Wahl (Salk Institute) for the histone H2B–GFP, M. Phillips (New York University) for the HA–Kras4A and Flag–Kras4B constructs and V. Schreiber (University of Strasbourg) for the ARTD1−/− (PARP1−/−) MEFs. H.C.H. acknowledges support from NIH-NIGMS R01 GM087544 grant.
Author information
Authors and Affiliations
Contributions
H.C.H. and N.P.W. designed the experiments, analyzed data and wrote the manuscript. H.M. helped with mass spectrometry analysis. J.P.F. performed and helped analyze the NAD metabolite mass spectrometry. N.P.W. performed all other experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Figures 1–22. (PDF 2533 kb)
Supplementary Dataset 1
List of proteins with a fudge factor greater than 1 and a false discovery rate of 0.05 for the unstressed samples. (XLSX 115 kb)
Supplementary Dataset 2
List of proteins with a fudge factor greater than 1 and a false discovery rate of 0.05 for the stressed samples. (XLSX 111 kb)
Supplementary Dataset 3
List of proteins with a difference of means greater than 2 with the probability of that difference P <0.05 for the stressed vs. the unstressed samples. (XLSX 54 kb)
Rights and permissions
About this article
Cite this article
Westcott, N., Fernandez, J., Molina, H. et al. Chemical proteomics reveals ADP-ribosylation of small GTPases during oxidative stress. Nat Chem Biol 13, 302–308 (2017). https://doi.org/10.1038/nchembio.2280
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2280
This article is cited by
-
Full-length single-molecule protein fingerprinting
Nature Nanotechnology (2024)
-
FICD activity and AMPylation remodelling modulate human neurogenesis
Nature Communications (2020)
-
An Integrated Chemical Proteomics Approach for Quantitative Profiling of Intracellular ADP-Ribosylation
Scientific Reports (2019)
-
Mining the cellular inventory of pyridoxal phosphate-dependent enzymes with functionalized cofactor mimics
Nature Chemistry (2018)