Abstract
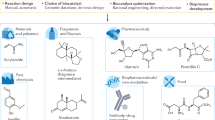
Enzymes are increasingly being used as biocatalysts in the generation of products that have until now been derived using traditional chemical processes. Such products range from pharmaceutical and agrochemical building blocks to fine and bulk chemicals and, more recently, components of biofuels. For a biocatalyst to be effective in an industrial process, it must be subjected to improvement and optimization, and in this respect the directed evolution of enzymes has emerged as a powerful enabling technology. Directed evolution involves repeated rounds of (i) random gene library generation, (ii) expression of genes in a suitable host and (iii) screening of libraries of variant enzymes for the property of interest. Both in vitro screening–based methods and in vivo selection–based methods have been applied to the evolution of enzyme function and properties. Significant developments have occurred recently, particularly with respect to library design, screening methodology, applications in synthetic transformations and strategies for the generation of new enzyme function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Schoemaker, H. et al. Dispelling the myths – biocatalysis in industrial synthesis. Science 299, 1694–1697 (2003).
Patel, R.N. Chemo-enzymatic synthesis of pharmaceutical intermediates. Expert Opin. Drug Discov. 3, 187–245 (2008).
Pollard, D.J. & Woodley, J.M. Biocatalysis for pharmaceutical intermediates: the future is now. Trends Biotechnol. 25, 66–73 (2007).
Panke, S., Held, M. & Wubbolts, M. Trends and innovations in industrial biocatalysis for the production of fine chemicals. Curr. Opin. Biotechnol. 15, 272–279 (2004).
Fox, R.J. & Clay, M.D. Catalytic effectiveness, a measure of enzyme proficiency for industrial applications. Trends Biotechnol. 27, 137–140 (2009).
Stemmer, W.P. Rapid evolution of a protein in vitro by DNA shuffling. Nature 370, 389–391 (1994).
Crameri, A. et al. DNA shuffling of a family of genes from diverse species accelerates directed evolution. Nature 391, 288–291 (1998).
Moore, J.C. & Arnold, F.H. Directed evolution of a para-nitrobenzyl esterase for aqueous-organic solvents. Nat. Biotechnol. 14, 458–467 (1996).
Reetz, M.T., Zonta, A., Schimossek, K., Liebeton, K. & Jaeger, K.-E. Creation of enantioselective biocatalysts for organic chemistry by in vitro evolution. Angew. Chem. Int. Ed. 36, 2830–2832 (1997).
Arnold, F.H. Directed evolution: creating biocatalysts for the future. Chem. Eng. Sci. 51, 5091–5102 (1996).
Tracewell, C.A. & Arnold, F.H. Directed enzyme evolution: climbing fitness peaks one amino acid at a time. Curr. Opin. Chem. Biol. 13, 3–9 (2009).
Alexeeva, M., Carr, R. & Turner, N.J. Directed evolution of enzymes: new biocatalysts for asymmetric synthesis. Org. Biomol. Chem. 1, 4133–4137 (2003).
Bershtein, S. & Tawfik, D.S. Advances in laboratory evolution of enzymes. Curr. Opin. Chem. Biol. 12, 151–158 (2008).
Turner, N.J. Directed evolution of enzymes new biocatalysts for organic synthesis. Chim. Oggi 26, 9–10 (2008).
Reetz, M.T. Directed evolution of enzymes for asymmetric syntheses. in Asymmetric Synthesis (eds. Christmann, M. & Bräse, S.) 207–211 (Wiley-VCH, Weinheim, Germany, 2007).
Johannes, T.W. & Zhao, H. Directed evolution of enzymes and biosynthetic pathways. Curr. Opin. Microbiol. 9, 261–267 (2006).
Sylvestre, J., Chautard, H., Cedrone, F. & Delcourt, M. Directed evolution of biocatalysts. Org. Process Res. Dev. 10, 562–571 (2006).
Hibbert, E.G. et al. Directed evolution of biocatalytic processes. Biomol. Eng. 22, 11–19 (2005).
Valetti, F. & Gilardi, G. Directed evolution of enzymes for product chemistry. Nat. Prod. Rep. 21, 490–511 (2004).
Lutz, S. & Patrick, W.M. Novel methods for directed evolution of enzymes: quality, not quantity. Curr. Opin. Biotechnol. 15, 291–297 (2004).
Montiel, C. & Bustos-Jaimes, I. Trends and challenges in directed evolution. Curr. Chem. Biol. 2, 50–59 (2008).
Fox, R.J. et al. Improving catalytic function by ProSAR-driven enzyme evolution. Nat. Biotechnol. 25, 338–344 (2007).
Fox, R.J. & Huisman, G.W. Enzyme optimization: moving from blind evolution to statistical exploration of sequence-function space. Trends Biotechnol. 26, 132–138 (2008).
Grate, J. Directed evolution of three biocatalysts to produce the key chiral building block for atorvastatin, the active ingredient in Lipitor. United States Environmental Protection Agency <http://www.epa.gov/greenchemistry/pubs/pgcc/winners/grca06.html> (2006).
Park, S. et al. Focusing mutations into the P. fluorescens esterase binding site increases enantioselectivity more effectively than distant mutations. Chem. Biol. 12, 45–54 (2005).
Morley, K.L. & Kazlauskas, R.J. Improving enzyme properties: when are closer mutations better? Trends Biotechnol. 23, 231–237 (2005).
Reetz, M.T., Kahakeaw, D. & Lohmer, R. Addressing the numbers problem in directed evolution. ChemBioChem 9, 1797–1804 (2008).
Reetz, M.T., Wang, L.-W. & Bocola, M. Directed evolution of enantioselective enzymes: iterative cycles of CASTing for probing protein-sequence space. Angew. Chem. Int. Ed. 45, 1236–1241 (2006).
Reetz, M.T. et al. Expanding the substrate scope of enzymes: combining mutations obtained by cASTing. Chem. Eur. J. 12, 6031–6038 (2006).
Muñoz, E. & Deem, M.W. Amino acid alphabet size in protein evolution experiments: better to search a small library thoroughly or a large library sparsely? Protein Eng. Des. Sel. 21, 311–317 (2008).
Reetz, M.T. & Wu, S. Greatly reduced amino acid alphabets in directed evolution: making the right choice for saturation mutagenesis at homologous enzyme positions. Chem. Commun. (Camb.) 5499–5501 (2008).
Hult, K. & Berglund, P. Enzyme promiscuity: mechanism and applications. Trends Biotechnol. 25, 231–238 (2007).
Taglieber, A. et al. Alternate-site enzyme promiscuity. Angew. Chem. Int. Ed. 46, 8597–8600 (2007).
Bornscheuer, U.T. & Kazlauskas, R.J. Reaction specificity of enzymes: catalytic promiscuity in biocatalysis: using old enzymes to form new bonds and follow new pathways. Angew. Chem. Int. Ed. 43, 6032–6040 (2004).
Peisajovich, S.G. & Tawfik, D.S. Protein engineers turned evolutionists. Nat. Methods 4, 991–994 (2007).
Bershtein, S., Goldin, K. & Tawfik, D.S. Intense neutral drifts yield robust and evolvable consensus proteins. J. Mol. Biol. 379, 1029–1044 (2008).
Aharoni, A. et al. The 'evolvability' of promiscuous protein functions. Nat. Genet. 37, 73–76 (2004).
Gupta, R.D. & Tawfik, D.S. Directed enzyme evolution via small and effective neutral drift libraries. Nat. Methods 5, 939–942 (2008).
Bloom, J.D., Romero, P.A., Lu, Z. & Arnold, F.H. Neutral genetic drift can alter promiscuous protein functions, potentially aiding functional evolution. Biol. Direct 2, 1–19 (2007).
Sakai, A. et al. Evolution of enzymatic activities in the enolase superfamily: stereochemically distinct mechanisms in two families of cis,cis-muconate lactonizing enzymes. Biochemistry 48, 1445–1453 (2009).
Grogan, G. Emergent mechanistic diversity of enzyme-catalysed β-diketone cleavage. Biochem. J. 388, 721–730 (2005).
Hamed, R.B., Batchelar, E.T., Clifton, I.J. & Schofield, C.J. Mechanisms and structures of crotonase superfamily enzymes-how nature controls enolate and oxyanion reactivity. Cell. Mol. Life Sci. 65, 2507–2527 (2008).
Hasnaoui-Dijoux, G., Majerić Elenkov, M., Lutje Spelberg, J.H., Hauer, B. & Janssen, D.B. Catalytic promiscuity of halohydrin dehalogenase and its application in enantioselective epoxide ring opening. ChemBioChem 9, 1048–1051 (2008).
Terao, Y., Miyamoto, K. & Ohta, H. Introduction of single mutation changes arylmalonate decarboxylase to racemase. Chem. Commun. (Camb.) 3600–3602 (2006).
Seebeck, F.P., Guainazzi, A., Amoreira, C., Baldridge, K.K. & Hilvert, D. Stereoselectivity and expanded substrate scope of an engineered PLP-dependent aldolase. Angew. Chem. Int. Ed. 45, 6824–6826 (2006).
Jochens, H. et al. Converting an esterase into an epoxide hydrolase. Angew. Chem. Int. Ed. 48, 3532–3535 (2009).
Keefe, A.D. & Szostak, J. Functional proteins from a random sequence library. Nature 410, 715–718 (2001).
Park, H.S. et al. Design and evolution of new catalytic activity with an existing protein scaffold. Science 311, 535–538 (2006).
Röthlisberger, D. et al. Kemp elimination catalysts by computational enzyme design. Nature 453, 190–195 (2008).
Jiang, L. et al. De novo computational design of retro-aldol enzymes. Science 319, 1387–1391 (2008).
Sterner, R., Merkl, R. & Raushel, F.M. Computational design of enzymes. Chem. Biol. 15, 421–423 (2008).
Damborsky, J. & Brezovsky, J. Computational tools for designing and engineering biocatalysts. Curr. Opin. Chem. Biol. 13, 26–34 (2009).
Smith, A.J.T. et al. Structural reorganization and preorganization in enzyme active sites: comparisons of experimental and theoretically ideal active site geometries in the multistep serine esterase reaction cycle. J. Am. Chem. Soc. 130, 15361–15373 (2008).
Seelig, B. & Szostak, J.W. Selection and evolution of enzymes from a partially randomized non-catalytic scaffold. Nature 448, 828–831 (2007).
Leemhuis, H., Kelly, R.M. & Dijkhuizen, L. Directed evolution of enzymes: library screening strategies. IUBMB Life 61, 222–228 (2009).
Belder, D., Ludwig, M., Wang, L.-W. & Reetz, M.T. Enantioselective catalysis and analysis on a chip. Angew. Chem. Int. Ed. 45, 2463–2466 (2006).
Mastrobattista, E. et al. High-throughput screening of enzyme libraries: in vitro evolution of a β-galactosidase by fluorescence-activated sorting of double emulsions. Chem. Biol. 12, 1291–1300 (2005).
Griffiths, A.D. & Tawfik, D.S. Directed evolution of an extremely fast phosphotriesterase by in vitro compartmentalization. EMBO J. 22, 24–35 (2003).
Fernandez-Gacio, A., Uguen, M. & Fastrez, J. Phage display as a tool for the directed evolution of enzymes. Trends Biotechnol. 21, 408–414 (2003).
Lipovsek, D. et al. Selection of horseradish peroxidase variants with enhanced enantioselectivity by yeast surface display. Chem. Biol. 14, 1176–1185 (2007).
Qian, Z., Fields, C.J. & Lutz, S. Investigating the structural and functional consequences of circular permutation on lipase B from Candida antarctica. ChemBioChem 8, 1989–1996 (2007).
Enright, A. et al. Stereoinversion of β- and γ-substituted-α-amino acids using a chemoenzymatic oxidation-reduction procedure. Chem. Commun. (Camb.) 2636–2637 (2003).
Roff, G.J., Lloyd, R.C. & Turner, N.J. A versatile chemo-enzymatic route to enantiomerically pure β-branched-α-amino acids. J. Am. Chem. Soc. 126, 4098–4099 (2004).
Alexeeva, M., Enright, A., Dawson, M.J., Mahmoudian, M. & Turner, N.J. Deracemisation of α-methylbenzylamine using an enzyme obtained by in vitro evolution. Angew. Chem. Int. Ed. 41, 3177–3180 (2002).
Carr, R. et al. Directed evolution of an amine oxidase possessing both broad substrate specificity and high enantioselectivity. Angew. Chem. Int. Ed. 42, 4807–4810 (2003).
Carr, R. et al. Directed evolution of an amine oxidase for the preparative deracemisation of cyclic secondary amines. ChemBioChem 6, 637–639 (2005).
Dunsmore, C.J., Carr, R., Fleming, T. & Turner, N.J. A chemo-enzymatic route to enantiomerically pure cyclic tertiary amines. J. Am. Chem. Soc. 128, 2224–2225 (2006).
Eve, T.S.C., Wells, A.S. & Turner, N.J. Enantioselective oxidation of O-methyl-N-hydroxylamines using MAO-N as catalyst. Chem. Commun. (Camb.) 1530–1531 (2007).
Bailey, K.R., Ellis, A.J., Reiss, R., Snape, T.J. & Turner, N.J. A template-based mnemonic for monoamine oxidase (MAO-N) catalyzed reactions and its application to the chemo-enzymatic deracemisation of the alkaloid (±)-crispine A. Chem. Commun. (Camb.) 3640–3642 (2007).
Atkin, K.E. et al. The structure of monoamine oxidase from Aspergillus niger provides a molecular context for improvements in activity obtained by directed evolution. J. Mol. Biol. 384, 1218–1231 (2008).
Jennewein, S. et al. Directed evolution of an industrial biocatalyst: 2-deoxy-D-ribose 5-phosphate aldolase. Biotechnol. J. 1, 537–548 (2006).
Ran, N. & Frost, J.W. Directed evolution of 2-keto-3-deoxy-6-phosphogalactonate aldolase to replace 3-deoxy-D-arabino-heptulosonic acid 7-phosphate synthase. J. Am. Chem. Soc. 129, 6130–6139 (2007).
Hsu, C.-C., Hong, Z., Wada, M., Franke, D. & Wong, C.-H. Directed evolution of D-sialic acid aldolase to L-3-deoxy-manno-2-octulosonic acid (L-KDO) aldolase. Proc. Natl. Acad. Sci. USA 102, 9122–9126 (2005).
Williams, G.J., Woodhall, T., Farnsworth, L.M., Nelson, A. & Berry, A. Creation of a pair of stereochemically complementary biocatalysts. J. Am. Chem. Soc. 128, 16238–16247 (2006).
Smith, M.E.B., Hibbert, E.G., Jones, A.B., Dalby, P.A. & Hailes, H.C. Enhancing and reversing the stereoselectivity of Escherichia coli transketolase via single-point mutations. Adv. Synth. Catal. 350, 2631–2638 (2008).
Tee, K.L. & Schwaneberg, U. A screening system for the directed evolution of epoxygenases: importance of position 184 in P450 BM3 for stereoselective styrene epoxidation. Angew. Chem. Int. Ed. 45, 5380–5383 (2006).
Reetz, M.T. et al. Directed evolution of an enantioselective epoxide hydrolase: uncovering the source of enantioselectivity at each evolutionary stage. J. Am. Chem. Soc. 131, 7334–7343 (2009).
Liu, Z. et al. Laboratory evolved biocatalysts for stereoselective syntheses of substituted benzaldehyde cyanohydrins. ChemBioChem 9, 58–61 (2008).
Koch, D.J., Chen, M.M., van Beilen, J.B. & Arnold, F.H. In vivo evolution of butane oxidation by terminal alkane hydroxylases AlkB and CYP153A6. Appl. Environ. Microbiol. 75, 337–344 (2009).
Landwehr, M. et al. Enantioselective alpha-hydroxylation of 2-arylacetic acid derivatives and buspirone catalyzed by engineered cytochrome P450 BM-3. J. Am. Chem. Soc. 128, 6058–6059 (2006).
Escalettes, F. & Turner, N.J. Directed evolution of galactose oxidase: generation of enantioselective secondary alcohol oxidases. ChemBioChem 9, 857–860 (2008).
Truppo, M.D., Escalettes, F. & Turner, N.J. Rapid determination of both the activity and enantioselectivity of ketoreductases. Angew. Chem. Int. Ed. 49, 2639–2641 (2008).
Klein, G. et al. Tailoring the active site of chemzymes by using a chemogenetic-optimization procedure: towards substrate-specific artificial hydrogenases based on the biotin-avidin technology. Angew. Chem. Int. Ed. 44, 7764–7767 (2005).
Ward, T.R. Artificial enzymes made to order: combination of computational design and directed evolution. Angew. Chem. Int. Ed. 47, 7802–7803 (2008).
Reetz, M.T., Peyralans, J.J.-P., Maichele, A., Fu, Y. & Maywald, M. Directed evolution of hybrid enzymes: evolving enantioselectivity of an achiral Rh-complex anchored to a protein. Chem. Commun. (Camb.) 4318–4320 (2006).
Jing, Q., Okrasa, K. & Kazlauskas, R.J. Stereoselective hydrogenation of olefins using rhodium-substituted carbonic anhydrase - a new reductase. Chem. Eur. J. 15, 1370–1376 (2009).
Pordea, A. et al. Artificial metalloenzyme for enantioselective sulfoxidation based on vanadyl-loaded streptavidin. J. Am. Chem. Soc. 130, 8085–8088 (2008).
Rousselot-Pailley, P. et al. The protein environment drives selectivity for sulfide oxidation by an artificial metalloenzyme. ChemBioChem 10, 545 (2009).
Pierron, J. et al. Artificial metalloenzymes for asymmetric allylic alkylation on the basis of the biotin-avidin technology. Angew. Chem. Int. Ed. 47, 701–705 (2008).
Umeno, D., Tobias, A.V. & Arnold, F.H. Diversifying carotenoid biosynthetic pathways by directed evolution. Microbiol. Mol. Biol. Rev. 69, 51–78 (2005).
Chatterjee, R. & Yuan, L. Directed evolution of metabolic pathways. Trends Biotechnol. 24, 28–38 (2006).
Fischbach, M.A., Lai, J.R., Roche, E.D., Walsh, C.T. & Liu, D.R. Directed evolution can rapidly improve the activity of chimeric assembly-line enzymes. Proc. Natl. Acad. Sci. USA 104, 11951–11956 (2007).
Chica, R.A., Doucet, N. & Pelletier, J.N. Semi-rational approaches to engineering enzyme activity: combining the benefits of directed evolution and rational design. Curr. Opin. Biotechnol. 16, 378–384 (2005).
Kazlauskas, R.J. & Lutz, S. Engineering enzymes by intelligent design. Curr. Opin. Chem. Biol. 13, 1–2 (2009).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Turner, N. Directed evolution drives the next generation of biocatalysts. Nat Chem Biol 5, 567–573 (2009). https://doi.org/10.1038/nchembio.203
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.203
This article is cited by
-
ACIDES: on-line monitoring of forward genetic screens for protein engineering
Nature Communications (2023)
-
Bioprospecting of microbial enzymes: current trends in industry and healthcare
Applied Microbiology and Biotechnology (2022)
-
Immobilization Horseradish Peroxidase onto UiO-66-NH2 for Biodegradation of Organic Dyes
Journal of Inorganic and Organometallic Polymers and Materials (2022)
-
Enhancement of protein thermostability by three consecutive mutations using loop-walking method and machine learning
Scientific Reports (2021)
-
Enhanced catalytic efficiency and coenzyme affinity of leucine dehydrogenase by comprehensive screening strategy for L-tert-leucine synthesis
Applied Microbiology and Biotechnology (2021)