Abstract
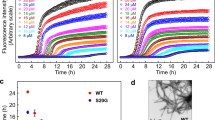
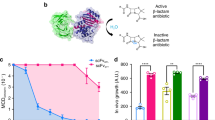
Protein aggregation underlies an array of human diseases, yet only one small-molecule therapeutic targeting this process has been successfully developed to date. Here, we introduce an in vivo system, based on a β-lactamase tripartite fusion construct, that is capable of identifying aggregation-prone sequences in the periplasm of Escherichia coli and inhibitors that prevent their aberrant self-assembly. We demonstrate the power of the system using a range of proteins, from small unstructured peptides (islet amyloid polypeptide and amyloid β) to larger, folded immunoglobulin domains. Configured in a 48-well format, the split β-lactamase sensor readily differentiates between aggregation-prone and soluble sequences. Performing the assay in the presence of 109 compounds enabled a rank ordering of inhibition and revealed a new inhibitor of islet amyloid polypeptide aggregation. This platform can be applied to both amyloidogenic and other aggregation-prone systems, independent of sequence or size, and can identify small molecules or other factors able to ameliorate or inhibit protein aggregation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Sipe, J.D. et al. Nomenclature 2014: amyloid fibril proteins and clinical classification of the amyloidosis. Amyloid 21, 221–224 (2014).
Knowles, T.P.J., Vendruscolo, M. & Dobson, C.M. The amyloid state and its association with protein misfolding diseases. Nat. Rev. Mol. Cell Biol. 15, 384–396 (2014).
Tipping, K.W., van Oosten-Hawle, P., Hewitt, E.W. & Radford, S.E. Amyloid fibres: inert end-stage aggregates or key players in disease? Trends Biochem. Sci. 40, 719–727 (2015).
Pieri, L., Madiona, K., Bousset, L. & Melki, R. Fibrillar α-synuclein and huntingtin exon 1 assemblies are toxic to the cells. Biophys. J. 102, 2894–2905 (2012).
Bulawa, C.E. et al. Tafamidis, a potent and selective transthyretin kinetic stabilizer that inhibits the amyloid cascade. Proc. Natl. Acad. Sci. USA 109, 9629–9634 (2012).
Meng, F., Marek, P., Potter, K.J., Verchere, C.B. & Raleigh, D.P. Rifampicin does not prevent amyloid fibril formation by human islet amyloid polypeptide but does inhibit fibril thioflavin-T interactions: implications for mechanistic studies of β-cell death. Biochemistry 47, 6016–6024 (2008).
Butterfield, S., Hejjaoui, M., Fauvet, B., Awad, L. & Lashuel, H.A. Chemical strategies for controlling protein folding and elucidating the molecular mechanisms of amyloid formation and toxicity. J. Mol. Biol. 421, 204–236 (2012).
Foit, L. et al. Optimizing protein stability in vivo. Mol. Cell 36, 861–871 (2009).
Kim, W. et al. A high-throughput screen for compounds that inhibit aggregation of the Alzheimer's peptide. ACS Chem. Biol. 1, 461–469 (2006).
Lee, L.L., Ha, H., Chang, Y.T. & DeLisa, M.P. Discovery of amyloid-beta aggregation inhibitors using an engineered assay for intracellular protein folding and solubility. Protein Sci. 18, 277–286 (2009).
Espargaró, A., Sabate, R. & Ventura, S. Thioflavin-S staining coupled to flow cytometry. A screening tool to detect in vivo protein aggregation. Mol. Biosyst. 8, 2839–2844 (2012).
Romero, D., Sanabria-Valentín, E., Vlamakis, H. & Kolter, R. Biofilm inhibitors that target amyloid proteins. Chem. Biol. 20, 102–110 (2013).
McKoy, A.F., Chen, J., Schupbach, T. & Hecht, M.H. A novel inhibitor of amyloid β (Aβ) peptide aggregation: from high throughput screening to efficacy in an animal model of Alzheimer disease. J. Biol. Chem. 287, 38992–39000 (2012).
Morell, M., de Groot, N.S., Vendrell, J., Avilés, F.X. & Ventura, S. Linking amyloid protein aggregation and yeast survival. Mol. Biosyst. 7, 1121–1128 (2011).
Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 67, 593–656 (2003).
Hailu, T.T., Foit, L. & Bardwell, J.C.A. In vivo detection and quantification of chemicals that enhance protein stability. Anal. Biochem. 434, 181–186 (2013).
Quan, S. et al. Genetic selection designed to stabilize proteins uncovers a chaperone called Spy. Nat. Struct. Mol. Biol. 18, 262–269 (2011).
Chen, Y.-R. & Glabe, C.G. Distinct early folding and aggregation properties of Alzheimer amyloid-β peptides Abeta40 and Abeta42: stable trimer or tetramer formation by Abeta42. J. Biol. Chem. 281, 24414–24422 (2006).
Bitan, G., Lomakin, A. & Teplow, D.B. Amyloid β-protein oligomerization: prenucleation interactions revealed by photo-induced cross-linking of unmodified proteins. J. Biol. Chem. 276, 35176–35184 (2001).
Young, L.M., Cao, P., Raleigh, D.P., Ashcroft, A.E. & Radford, S.E. Ion mobility spectrometry-mass spectrometry defines the oligomeric intermediates in amylin amyloid formation and the mode of action of inhibitors. J. Am. Chem. Soc. 136, 660–670 (2014).
Koo, B.W. & Miranker, A.D. Contribution of the intrinsic disulfide to the assembly mechanism of islet amyloid. Protein Sci. 14, 231–239 (2005).
Qiao, Q., Bowman, G.R. & Huang, X. Dynamics of an intrinsically disordered protein reveal metastable conformations that potentially seed aggregation. J. Am. Chem. Soc. 135, 16092–16101 (2013).
Sgourakis, N.G., Yan, Y., McCallum, S.A., Wang, C. & Garcia, A.E. The Alzheimer's peptides Abeta40 and 42 adopt distinct conformations in water: a combined MD/NMR study. J. Mol. Biol. 368, 1448–1457 (2007).
Young, L.M. et al. Screening and classifying small-molecule inhibitors of amyloid formation using ion mobility spectrometry-mass spectrometry. Nat. Chem. 7, 73–81 (2015).
Nesterov, E.E. et al. In vivo optical imaging of amyloid aggregates in brain: design of fluorescent markers. Angew. Chem. Int. Ed. Engl. 44, 5452–5456 (2005).
O'Nuallain, B. & Wetzel, R. Conformational Abs recognizing a generic amyloid fibril epitope. Proc. Natl. Acad. Sci. USA 99, 1485–1490 (2002).
Eichner, T., Kalverda, A.P., Thompson, G.S., Homans, S.W. & Radford, S.E. Conformational conversion during amyloid formation at atomic resolution. Mol. Cell 41, 161–172 (2011).
Valleix, S. et al. Hereditary systemic amyloidosis due to Asp76Asn variant β2-microglobulin. N. Engl. J. Med. 366, 2276–2283 (2012).
Mangione, P.P. et al. Structure, folding dynamics, and amyloidogenesis of D76N β2-microglobulin: roles of shear flow, hydrophobic surfaces, and α-crystallin. J. Biol. Chem. 288, 30917–30930 (2013).
Jespers, L., Schon, O., Famm, K. & Winter, G. Aggregation-resistant domain antibodies selected on phage by heat denaturation. Nat. Biotechnol. 22, 1161–1165 (2004).
Jespers, L., Schon, O., James, L.C., Veprintsev, D. & Winter, G. Crystal structure of HEL4, a soluble, refoldable human V(H) single domain with a germ-line scaffold. J. Mol. Biol. 337, 893–903 (2004).
Daval, M. et al. The effect of curcumin on human islet amyloid polypeptide misfolding and toxicity. Amyloid 17, 118–128 (2010).
Sparks, S., Liu, G., Robbins, K.J. & Lazo, N.D. Curcumin modulates the self-assembly of the islet amyloid polypeptide by disassembling α-helix. Biochem. Biophys. Res. Commun. 422, 551–555 (2012).
Meng, F. et al. The sulfated triphenyl methane derivative acid fuchsin is a potent inhibitor of amyloid formation by human islet amyloid polypeptide and protects against the toxic effects of amyloid formation. J. Mol. Biol. 400, 555–566 (2010).
Meng, F. & Raleigh, D.P. Inhibition of glycosaminoglycan-mediated amyloid formation by islet amyloid polypeptide and proIAPP processing intermediates. J. Mol. Biol. 406, 491–502 (2011).
Palhano, F.L., Lee, J., Grimster, N.P. & Kelly, J.W. Toward the molecular mechanism(s) by which EGCG treatment remodels mature amyloid fibrils. J. Am. Chem. Soc. 135, 7503–7510 (2013).
Cheng, B. et al. Coffee components inhibit amyloid formation of human islet amyloid polypeptide in vitro: possible link between coffee consumption and diabetes mellitus. J. Agric. Food Chem. 59, 13147–13155 (2011).
Cheng, B. et al. Silibinin inhibits the toxic aggregation of human islet amyloid polypeptide. Biochem. Biophys. Res. Commun. 419, 495–499 (2012).
Tu, L.-H. et al. Mutational analysis of the ability of resveratrol to inhibit amyloid formation by islet amyloid polypeptide: critical evaluation of the importance of aromatic-inhibitor and histidine-inhibitor interactions. Biochemistry 54, 666–676 (2015).
Aitken, J.F., Loomes, K.M., Konarkowska, B. & Cooper, G.J.S. Suppression by polycyclic compounds of the conversion of human amylin into insoluble amyloid. Biochem. J. 374, 779–784 (2003).
Aarabi, M.H. & Mirhashemi, S.M. The role of two natural flavonoids on human amylin aggregation. Afr. J. Pharm. Pharmacol. 6, 2374–2379 (2012).
Zelus, C. et al. Myricetin inhibits islet amyloid polypeptide (IAPP) aggregation and rescues living mammalian cells from IAPP toxicity. Open Biochem. J. 6, 66–70 (2012).
Porat, Y., Mazor, Y., Efrat, S. & Gazit, E. Inhibition of islet amyloid polypeptide fibril formation: a potential role for heteroaromatic interactions. Biochemistry 43, 14454–14462 (2004).
Wu, C., Lei, H., Wang, Z., Zhang, W. & Duan, Y. Phenol red interacts with the protofibril-like oligomers of an amyloidogenic hexapeptide NFGAIL through both hydrophobic and aromatic contacts. Biophys. J. 91, 3664–3672 (2006).
Noor, H., Cao, P. & Raleigh, D.P. Morin hydrate inhibits amyloid formation by islet amyloid polypeptide and disaggregates amyloid fibers. Protein Sci. 21, 373–382 (2012).
Li, J., Zhu, M., Manning-Bog, A.B., Di Monte, D.A. & Fink, A.L. Dopamine and L-dopa disaggregate amyloid fibrils: implications for Parkinson's and Alzheimer's disease. FASEB J. 18, 962–964 (2004).
Lee, H.-J. et al. Dopamine promotes formation and secretion of non-fibrillar alpha-synuclein oligomers. Exp. Mol. Med. 43, 216–222 (2011).
Baell, J. & Walters, M.A. Chemistry: chemical con artists foil drug discovery. Nature 513, 481–483 (2014).
Baell, J.B. & Holloway, G.A. New substructure filters for removal of pan assay interference compounds (PAINS) from screening libraries and for their exclusion in bioassays. J. Med. Chem. 53, 2719–2740 (2010).
Jelsch, C., Mourey, L., Masson, J.-M. & Samama, J.-P. Crystal structure of Escherichia coli TEM1 β-lactamase at 1.8 A resolution. Proteins 16, 364–383 (1993).
Williamson, J.A. & Miranker, A.D. Direct detection of transient α-helical states in islet amyloid polypeptide. Protein Sci. 16, 110–117 (2007).
Walsh, D.M. et al. A facile method for expression and purification of the Alzheimer's disease-associated amyloid β-peptide. FEBS J. 276, 1266–1281 (2009).
Kad, N.M., Thomson, N.H., Smith, D.P., Smith, D.A. & Radford, S.E. β(2)-microglobulin and its deamidated variant, N17D form amyloid fibrils with a range of morphologies in vitro. J. Mol. Biol. 313, 559–571 (2001).
Marek, P., Woys, A.M., Sutton, K., Zanni, M.T. & Raleigh, D.P. Efficient microwave-assisted synthesis of human islet amyloid polypeptide designed to facilitate the specific incorporation of labeled amino acids. Org. Lett. 12, 4848–4851 (2010).
Giles, K. et al. Applications of a travelling wave-based radio-frequency-only stacked ring ion guide. Rapid Commun. Mass Spectrom. 18, 2401–2414 (2004).
Schrödinger Release 9.3 (Schrödinger LLC, New York, 2014-2).
Acknowledgements
J.C.S. is co-funded by Innovate UK (131841) and the Biotechnology and Biological Sciences Research Council (BBSRC) (BB/M01259X/1) and was previously funded by a BBSRC CASE studentship (grant number BB/H014713/1) sponsored by Avacta Analytical plc, Wetherby, UK. L.M.Y. is funded by a BBSRC CASE studentship (grant number BB/I015361/1) sponsored by Micromass UK Ltd./Waters Corporation, Manchester, UK. R.A.M. is funded by a BBSRC studentship (grant number BB/F01614X/1). The Synapt HDMS mass spectrometer was purchased with funds from the BBSRC (BB/E012558/1). M.P.J., D.J.B. and S.E.R. also acknowledge funding from the European Research Council (ERC) under the European Union's Seventh Framework Programme (FP7/2007-2013)/ERC grant agreement number 322408. R.J.F. and C.H.R. acknowledge the Biomedical Health Research Centre (University of Leeds) for funding. We thank J. Bardwell (University of Michigan) for his longstanding collaboration and advice at the beginning of the study. We thank S. Webster (formerly at Avacta Analytical plc) for his advice and helpful discussions. We are very grateful to D. Raleigh and L.-H. Tu (Stony Brook University) for kindly providing the synthetic hIAPP and rIAPP peptides. We thank R. Wetzel (University of Pittsburgh) for providing the WO1 antibody. We also acknowledge our collaborators and all members of our groups for helpful discussions.
Author information
Authors and Affiliations
Contributions
J.C.S. and L.M.Y. contributed equally to this work. J.C.S. designed the study, purified the β-lactamase constructs, performed the in vivo experiments, performed the β-lactamase activity assays in vitro, performed TEM, analyzed results and wrote the manuscript. L.M.Y. conceived, designed and performed mass spectrometry experiments, performed TEM and thioflavin T fluorometry and analyzed results. R.A.M. purified Aβ40 and performed TEM and thioflavin T fluorometry. M.P.J. performed western blots, dot blots, and thioflavin T and NIAD-4 fluorometry. C.H.R. and R.J.F. designed and prepared the small-molecule screening library. R.J.F. also performed all PAINS analyses. D.A.S. contributed to experiment design. A.E.A. conceived and designed mass spectrometry experiments. D.J.B. and S.E.R. conceived and designed the experiments and wrote the manuscript. All authors contributed to manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–4 and Supplementary Figures 1–15. (PDF 6345 kb)
Supplementary Data Set 1
Full list of all small molecules used in this study and their structures and chemical properties. (PDF 4471 kb)
Rights and permissions
About this article
Cite this article
Saunders, J., Young, L., Mahood, R. et al. An in vivo platform for identifying inhibitors of protein aggregation. Nat Chem Biol 12, 94–101 (2016). https://doi.org/10.1038/nchembio.1988
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1988
This article is cited by
-
Protein assembly systems in natural and synthetic biology
BMC Biology (2020)
-
An in vivo platform to select and evolve aggregation-resistant proteins
Nature Communications (2020)
-
Molecular Targets and Therapeutic Strategies in Spinocerebellar Ataxia Type 7
Neurotherapeutics (2019)
-
An integrated bacterial system for the discovery of chemical rescuers of disease-associated protein misfolding
Nature Biomedical Engineering (2017)
-
Enabling stop codon read-through translation in bacteria as a probe for amyloid aggregation
Scientific Reports (2017)