Abstract
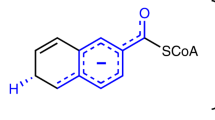
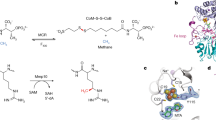
The pyridine nucleotides NADH and NADPH (NAD(P)H) are ubiquitous redox coenzymes that are present in all living cells. Although about 16% of all characterized enzymes use pyridine nucleotides as hydride donors or acceptors during catalysis, a detailed understanding of how the hydride is transferred between NAD(P)H and the corresponding substrate is lacking for many enzymes. Here we present evidence for a new mechanism that operates during enzymatic hydride transfers using crotonyl-CoA carboxylase/reductase (Ccr) as a case study. We observed a covalent ene intermediate between NADPH and the substrate, crotonyl-CoA, using NMR, high-resolution MS and stopped-flow spectroscopy. Preparation of the ene intermediate further allowed direct access to the catalytic cycle of other NADPH-dependent enzymes—including those from type II fatty acid biosynthesis—in an unprecedented way, suggesting that formation of NAD(P)H ene intermediates is a more general principle in catalysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Westheimer, F.H., Fisher, H.F., Conn, E.E. & Vennesland, B. The enzymatic transfer of hydrogen from alcohol to Dpn. J. Am. Chem. Soc. 73, 2403–2408 (1951).
Cheng, J.P., Lu, Y., Zhu, X.Q. & Mu, L.J. Energetics of multistep versus one-step hydride transfer reactions of reduced nicotinamide adenine dinucleotide (NADH) models with organic cations and p-quinones. J. Org. Chem. 63, 6108–6114 (1998).
Westheimer, F.H. Mechanism of action of the pyridine nucleotides. in Pyridine Nucleotide Coenzymes: Chemical, Biochemical, and Medical aspects Vol. 2 (eds. Dolphin, D., Poulson, R. & Avramovic, O.) 253–322 (Wiley, New York, 1986).
Abeles, R.H., Hutton, R.F. & Westheimer, F.H. The reduction of thioketones by a model for a coenzyme. J. Am. Chem. Soc. 79, 712–716 (1957).
Wu, Y.D. & Houk, K.N. Theoretical transition structures for hydride transfer to methyleniminium ion from methylamine and dihydropyridine. On the nonlinearity of hydride transfers. J. Am. Chem. Soc. 109, 2226–2227 (1987).
Bugg, T. Introduction to Enzyme and Coenzyme Chemistry (Wiley-Blackwell, Oxford, 2012).
Hamilton, G.A. in Progress in Bioorganic Chemistry Vol. 1 (eds. Kaiser, E.T. & Kezdy, F.J.) 83–157 (1971).
MacInnes, I., Nonhebel, D.C., Orszulik, S.T. & Suckling, C.J. On the mechanism of hydrogen transfer by nicotinamide coenzymes and alcohol-dehydrogenase. J. Chem. Soc., Perkin Trans. 1 1983, 2777–2779 (1983).
Hedlund, J., Jornvall, H. & Persson, B. Subdivision of the MDR superfamily of medium-chain dehydrogenases/reductases through iterative hidden Markov model refinement. BMC Bioinformatics 11, 534 (2010).
Agarwal, P.K., Webb, S.P. & Hammes-Schiffer, S. Computational studies of the mechanism for proton and hydride transfer in liver alcohol dehydrogenase. J. Am. Chem. Soc. 122, 4803–4812 (2000).
Bahnson, B.J., Park, D.H., Kim, K., Plapp, B.V. & Klinman, J.P. Unmasking of hydrogen tunneling in the horse liver alcohol dehydrogenase reaction by site-directed mutagenesis. Biochemistry 32, 5503–5507 (1993).
Roston, D. & Kohen, A. Elusive transition state of alcohol dehydrogenase unveiled. Proc. Natl. Acad. Sci. USA 107, 9572–9577 (2010).
Youn, B. et al. Mechanistic and structural studies of apoform, binary, and ternary complexes of the Arabidopsis alkenal double bond reductase At5g16970. J. Biol. Chem. 281, 40076–40088 (2006).
Porté, S. et al. Kinetic and structural evidence of the alkenal/one reductase specificity of human ζ-crystallin. Cell Mol. Life Sci. 68, 1065–1077 (2011).
Schlegel, B.P., Ratnam, K. & Penning, T.M. Retention of NADPH-linked quinone reductase activity in an aldo-keto reductase following mutation of the catalytic tyrosine. Biochemistry 37, 11003–11011 (1998).
Airenne, T.T. et al. Structure-function analysis of enoyl thioester reductase involved in mitochondrial maintenance. J. Mol. Biol. 327, 47–59 (2003).
Chen, Z.J. et al. Structural enzymological studies of 2-enoyl thioester reductase of the human mitochondrial FAS II pathway: new insights into its substrate recognition properties. J. Mol. Biol. 379, 830–844 (2008).
Wallace, K.K. et al. Purification of crotonyl-CoA reductase from Streptomyces collinus and cloning, sequencing and expression of the corresponding gene in Escherichia coli. Eur. J. Biochem. 233, 954–962 (1995).
Erb, T.J. et al. Synthesis of C5-dicarboxylic acids from C2-units involving crotonyl-CoA carboxylase/reductase: the ethylmalonyl-CoA pathway. Proc. Natl. Acad. Sci. USA 104, 10631–10636 (2007).
Erb, T.J., Brecht, V., Fuchs, G., Muller, M. & Alber, B.E. Carboxylation mechanism and stereochemistry of crotonyl-CoA carboxylase/reductase, a carboxylating enoyl-thioester reductase. Proc. Natl. Acad. Sci. USA 106, 8871–8876 (2009).
Wilson, M.C. & Moore, B.S. Beyond ethylmalonyl-CoA: the functional role of crotonyl-CoA carboxylase/reductase homologs in expanding polyketide diversity. Nat. Prod. Rep. 29, 72–86 (2012).
Erb, T.J. et al. Synthesis of C5-dicarboxylic acids from C2-units involving crotonyl-CoA carboxylase/reductase: The ethylmalonyl-CoA pathway. Proc. Natl. Acad. Sci. USA 104, 10631–10636 (2007).
Quade, N., Huo, L., Rachid, S., Heinz, D.W. & Muller, R. Unusual carbon fixation gives rise to diverse polyketide extender units. Nat. Chem. Biol. 8, 117–124 (2012).
Yoo, H.G. et al. Characterization of 2-octenoyl-CoA carboxylase/reductase utilizing pteB from Streptomyce avermitilis. Biosci. Biotechnol. Biochem. 75, 1191–1193 (2011).
Torkko, J.M. et al. Candida tropicalis Etr1p and Saccharomyces cerevisiae Ybr026p (Mrf1′p), 2-enoyl thioester reductases essential for mitochondrial respiratory competence. Mol. Cell Biol. 21, 6243–6253 (2001).
Todd, J.D., Curson, A.R., Sullivan, M.J., Kirkwood, M. & Johnston, A.W. The Ruegeria pomeroyi acuI gene has a role in DMSP catabolism and resembles yhdH of E. coli and other bacteria in conferring resistance to acrylate. PLoS ONE 7, e35947 (2012).
Schneider, K., Asao, M., Carter, M.S. & Alber, B.E. Rhodobacter sphaeroides uses a reductive route via propionyl coenzyme A to assimilate 3-hydroxypropionate. J. Bacteriol. 194, 225–232 (2012).
Asao, M. & Alber, B.E. Acrylyl-CoA reductase, an enzyme involved in the assimilation of 3-hydroxypropionate by Rhodobacter sphaeroides. J. Bacteriol. 195, 4716–4725 (2013).
Alder, K., Pascher, F. & Schmitz, A. Absorption of maleic acid-anhydride and azodicarbon acid-ester in simple unsaturated carbohydrogens. Information on the substitution processes in the allyl position. Chem. Ber. 76, 27–53 (1943).
Pindur, U. & Schneider, G.H. Pericyclic key reactions in biological-systems and biomimetic syntheses. Chem. Soc. Rev. 23, 409–415 (1994).
Lamb, A.L. Pericyclic reactions catalyzed by chorismate-utilizing enzymes. Biochemistry 50, 7476–7483 (2011).
Chook, Y.M., Gray, J.V., Ke, H. & Lipscomb, W.N. The monofunctional chorismate mutase from Bacillus subtilis. Structure determination of chorismate mutase and its complexes with a transition state analog and prephenate, and implications for the mechanism of the enzymatic reaction. J. Mol. Biol. 240, 476–500 (1994).
DeClue, M.S., Baldridge, K.K., Kunzler, D.E., Kast, P. & Hilvert, D. Isochorismate pyruvate lyase: a pericyclic reaction mechanism? J. Am. Chem. Soc. 127, 15002–15003 (2005).
Libby, R.D. & Mehl, R.A. Characterization of covalent Ene adduct intermediates in “hydride equivalent” transfers in a dihydropyridine model for NADH reduction reactions. Bioorg. Chem. 40, 57–66 (2012).
Jacobson, B.M. Aromatization of 1,4-cyclohexadiene with tetracyanoethylene-Ene reaction. J. Am. Chem. Soc. 102, 886–887 (1980).
Sulzbach, R.A. & Iqbal, F.M. Addition of acrylonitrile to derivatives of 1,4-dihydropyridine. Angew. Chem. Int. Ed. Engl. 10, 733 (1971).
Inagaki, S. & Hirabayashi, Y. Mechanism of NAD(P)H reduction reactions. Bull. Chem. Soc. Jpn. 50, 3360–3364 (1977).
Rozwarski, D.A., Grant, G.A., Barton, D.H., Jacobs, W.R. Jr. & Sacchettini, J.C. Modification of the NADH of the isoniazid target (InhA) from Mycobacterium tuberculosis. Science 279, 98–102 (1998).
Bull, H.G. et al. Mechanism-based inhibition of human steroid 5 α-reductase by finasteride: enzyme-catalyzed formation of NADP-dihydrofinasteride, a potent bisubstrate analog inhibitor. J. Am. Chem. Soc. 118, 2359–2365 (1996).
Wolfenden, R. Transition state analogues for enzyme catalysis. Nature 223, 704–705 (1969).
Kapmeyer, H., Pfleiderer, G. & Trommer, W.E. A transition state analogue for two pyruvate metabolizing enzymes, lactate dehydrogenase and alanine dehydrogenase. Biochemistry 15, 5024–5028 (1976).
Harden, A. & Young, W.J. The alcoholic ferment of yeast-juice. Proc. R. Soc. Lond., B 77, 405–420 (1906).
Simon, E.J. & Shemin, D. The preparation of S-succinyl coenzyme-A. J. Am. Chem. Soc. 75, 2520 (1953).
Peyraud, R. et al. Demonstration of the ethylmalonyl-CoA pathway by using 13C-metabolomics. Proc. Natl. Acad. Sci. USA 106, 4846–4851 (2009).
Dawson, R.M.C., Elliott, D.C., Elliott, W.H. & Jones, K.M. Data for Biochemical Research 3rd edn (Clarendon Press, Oxford, 1986).
Studier, F.W. Protein production by auto-induction in high-density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Kuntze, K. et al. Enzymes involved in the anaerobic degradation of meta-substituted halobenzoates. Mol. Microbiol. 82, 758–769 (2011).
Wu, L. et al. Quantitative analysis of the microbial metabolome by isotope dilution mass spectrometry using uniformly C-13–labeled cell extracts as internal standards. Anal. Biochem. 336, 164–171 (2005).
Acknowledgements
This work was supported by a Schweizerische Nationalfonds–Ambizione grant (PZ00P3_136828/1) and an ETH fellowship to T.J.E. We thank B.M. Jaun for valuable discussions and critical inspection of NMR data, A.J. Rosenthal for suggestions during the early phase of this work and S. Weidenweber and U. Ermler for discussions about a structural understanding of Ccr. We acknowledge K. Hiltunen, J. Torkko and A. Kastaniotis (University of Oulu, Finland) for providing the etr1p gene, G. Sturm (Albert-Ludwigs-University of Freiburg, Germany) for providing plasmid pGS5, A. Essig, M. Künzler and M. Aebi (ETH Zurich, Switzerland) for help with the Pichia pastoris expression system, R. Kissner and C. Molina (ETH Zurich, Switzerland) for stopped-flow experiments, S. Weidenweber and C. Vogel for the initial cloning of the Ccr and Etr1p constructs, respectively, and National BioResource Project (NBRP)-E. coli at the National Institute of Genetics (NIG) for providing E. coli strain JW3222.
Author information
Authors and Affiliations
Contributions
R.G.R. and T.J.E. conceived and designed all experiments, with the exception of the NMR experiments, which were designed together with M.-O.E., and the MS analyses, which were designed together with P.K. and J.A.V. NMR experiments were performed by R.G.R. and M.-O.E. MS experiments were performed by R.G.R. and P.K. Enzyme kinetic assays and stopped-flow measurements, as well as purification of the intermediate, were performed by R.G.R. and D.M.P. R.G.R. and T.J.E. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Figures 1–13 and Supplementary Tables 1–3. (PDF 6066 kb)
Rights and permissions
About this article
Cite this article
Rosenthal, R., Ebert, MO., Kiefer, P. et al. Direct evidence for a covalent ene adduct intermediate in NAD(P)H-dependent enzymes. Nat Chem Biol 10, 50–55 (2014). https://doi.org/10.1038/nchembio.1385
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1385
This article is cited by
-
Inhibition of homoserine dehydrogenase by formation of a cysteine-NAD covalent complex
Scientific Reports (2018)
-
A conserved threonine prevents self-intoxication of enoyl-thioester reductases
Nature Chemical Biology (2017)
-
A crotonyl-CoA reductase-carboxylase independent pathway for assembly of unusual alkylmalonyl-CoA polyketide synthase extender units
Nature Communications (2016)
-
Kovalente Zwischenprodukte in NAD(P)H-abhängigen Oxidoreduktasen
BIOspektrum (2016)
-
Do it your (path)way –synthetische Wege zur CO2-Fixierung
BIOspektrum (2016)