Abstract
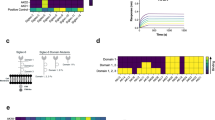
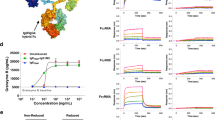
Development of specific inhibitors of allergy has had limited success, in part, owing to a lack of experimental models that reflect the complexity of allergen-IgE interactions. We designed a heterotetravalent allergen (HtTA) system, which reflects epitope heterogeneity, polyclonal response and number of immunodominant epitopes observed in natural allergens, thereby providing a physiologically relevant experimental model to study mast cell degranulation. The HtTA design revealed the importance of weak-affinity epitopes in allergy, particularly when presented with high-affinity epitopes. The effect of selective inhibition of weak-affinity epitope-IgE interactions was investigated with heterobivalent inhibitors (HBIs) designed to simultaneously target the antigen- and nucleotide-binding sites on the IgE Fab. HBI demonstrated enhanced avidity for the target IgE and was a potent inhibitor of degranulation in vitro and in vivo. These results demonstrate that partial inhibition of allergen-IgE interactions was sufficient to prevent mast cell degranulation, thus establishing the therapeutic potential of the HBI design.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Metzger, H. Transmembrane signaling—the joy of aggregation. J. Immunol. 149, 1477–1487 (1992).
Blank, U. & Rivera, J. Assays for regulated exocytosis of mast cell granules. Curr. Protoc. Cell. Biol. 15.11, 32 (2006).
Wang, J. et al. Correlation of IgE/IgG4 milk epitopes and affinity of milk-specific IgE antibodies with different phenotypes of clinical milk allergy. J. Allergy Clin. Immunol. 125, 695–702 (2010).
López-Torrejón, G. et al. An experimental and modeling-based approach to locate IgE epitopes of plant profilin allergens. J. Allergy Clin. Immunol. 119, 1481–1488 (2007).
Christensen, L.H., Holm, J., Lund, G., Riise, E. & Lund, K. Several distinct properties of the IgE repertoire determine effector cell degranulation in response to allergen challenge. J. Allergy Clin. Immunol. 122, 298–304 (2008).
Rougé, P. et al. Mapping and conformational analysis of IgE-binding epitopic regions on the molecular surface of the major Ara h 3 legumin allergen of peanut (Arachis hypogaea). Mol. Immunol. 46, 1067–1075 (2009).
Pacios, L.F. et al. Mimotope mapping as a complementary strategy to define allergen IgE-epitopes: peach Pru p 3 allergen as a model. Mol. Immunol. 45, 2269–2276 (2008).
Stanley, J.S. et al. Identification and mutational analysis of the immunodominant IgE binding epitopes of the major peanut allergen Ara h 2. Arch. Biochem. Biophys. 342, 244–253 (1997).
Tanabe, S. Epitope peptides and immunotherapy. Curr. Protein Pept. Sci. 8, 109–118 (2007).
Cerecedo, I. et al. Mapping of the IgE and IgG4 sequential epitopes of milk allergens with a peptide microarray-based immunoassay. J. Allergy Clin. Immunol. 122, 589–594 (2008).
Cong, Y., Lou, F., Xue, W., Li, L. & Chen, M. Characterisation of the IgE-binding immunodominant epitopes on Ara h1. Food Agric. Immunol. 19, 175–185 (2008).
Denery-Papini, S. et al. Immunoglobulin-E–binding epitopes of wheat allergens in patients with food allergy to wheat and in mice experimentally sensitized to wheat proteins. Clin. Exp. Allergy 41, 1478–1492 (2011).
Tordesillas, L. et al. Characterization of IgE epitopes of Cuc m 2, the major melon allergen, and their role in cross-reactivity with pollen profilins. Clin. Exp. Allergy 40, 174–181 (2010).
Guex, N. & Peitsch, M. SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18, 2714–2723 (1997).
Passante, E. & Frankish, N. The RBL-2H3 cell line: its provenance and suitability as a model for the mast cell. Inflamm. Res. 58, 737–745 (2009).
Andrews, N.L. et al. Small, mobile FcɛR1 receptor aggregates are signaling competent. Immunity 31, 469–479 (2009).
Handlogten, M.W., Kiziltepe, T., Moustakas, D.T. & Bilgicer, B. Design of a heterobivalent ligand to inhibit IgE clustering on mast cells. Chem. Biol. 18, 1179–1188 (2011).
Collins, A.M., Basil, M., Nguyen, K. & Thelian, D. Rat basophil leukaemia (RBL) cells sensitized with low affinity IgE respond to high valency antigen. Clin. Exp. Allergy 26, 964–970 (1996).
Hsu, C.L., Neilsen, C.V. & Bryce, P.J. IL-33 is produced by mast cells and regulates IgE-dependent inflammation. PLoS ONE 5, e11944 (2010).
James, L.C. & Tawfik, D.S. The specificity of cross-reactivity: promiscuous antibody binding involves specific hydrogen bonds rather than nonspecific hydrophobic stickiness. Protein Sci. 12, 2183–2193 (2003).
Hlavacek, W.S., Posner, R.G. & Perelson, A.S. Steric effects on multivalent ligand-receptor binding: exclusion of ligand sites by bound cell surface receptors. Biophys. J. 76, 3031–3043 (1999).
Xu, K., Goldstein, B., Holowka, D. & Baird, B. Kinetics of multivalent antigen DNP-BSA binding to IgE-FcɛRI in relationship to the stimulated tyrosine phosphorylation of FcɛRI. J. Immunol. 160, 3225–3235 (1998).
Rajagopalan, K. et al. Novel unconventional binding site in the variable region of immunoglobulins. Proc. Natl. Acad. Sci. USA 93, 6019–6024 (1996).
Alves, N.J. et al. Selective photocrosslinking of functional ligands to antibodies via the conserved nucleotide binding site. Biomaterials 34, 5700–5710 (2013).
Alves, N.J. et al. A small molecule based affinity chromatography method for antibody purification via nucleotide binding site targeting. Anal. Chem. 84, 7721–7728 (2012).
Schweitzer-Stenner, R., Licht, A., Luscher, I. & Pecht, I. Oligomerization and ring-closure of immunoglobulin-E class antibodies by divalent haptens. Biochemistry 26, 3602–3612 (1987).
Hunt, J. et al. A fluorescent biosensor reveals conformational changes in human immunoglobulin E Fc implications for mechanisms of receptor binding, inhibition, and allergen recognition. J. Biol. Chem. 287, 17459–17470 (2012).
Baird, E.J., Holowka, D., Coates, G.W. & Baird, B. Highly effective poly(ethylene glycol) architectures for specific inhibition of immune receptor activation. Biochemistry 42, 12739–12748 (2003).
Handlogten, M.W., Kiziltepe, T., Alves, N.J. & Bilgicer, B. Synthetic allergen design reveals the significance of moderate affinity epitopes in mast cell degranulation. ACS Chem. Biol. 7, 1796–1801 (2012).
Stefanick, J.F., Kiziltepe, T., Handlogten, M.W., Alves, N.J. & Bilgicer, B. Enhancement of antibody selectivity via bicyclic complex formation. J. Phys. Chem. Lett. 3, 598–602 (2012).
Bilgiçer, B., Moustakas, D.T. & Whitesides, G.M. A synthetic trivalent hapten that aggregates anti-2,4-DNP IgG into bicyclic trimers. J. Am. Chem. Soc. 129, 3722–3728 (2007).
Bilgiçer, B. et al. A non-chromatographic method for the purification of a bivalently active monoclonal IgG antibody from biological fluids. J. Am. Chem. Soc. 131, 9361–9367 (2009).
Handlogten, M.W., Kiziltepe, T. & Bilgicer, B. Design of a heterotetravalent synthetic allergen that reflects epitope heterogeneity and IgE antibody variability to study mast cell degranulation. Biochem. J. 449, 91–99 (2013).
Sil, D., Lee, J.B., Luo, D., Holowka, D. & Baird, B. Trivalent ligands with rigid DNA spacers reveal structural requirements for IgE receptor signaling in RBL mast cells. ACS Chem. Biol. 2, 674–684 (2007).
Posner, R.G. et al. Trivalent antigens for degranulation of mast cells. Org. Lett. 9, 3551–3554 (2007).
Baird, B., Zheng, Y. & Holowka, D. Structural mapping of IgE-Fc-ɛ-RI, an immunoreceptor complex. Acc. Chem. Res. 26, 428–434 (1993).
Dinneswara Reddy, G., Park, S., Cho, H.M., Kim, T. & Lee, M.E. Antiallergic activity profile in vitro RBL-2H3 and in vivo passive cutaneous anaphylaxis mouse model of new sila-substituted 1,3,4-oxadiazoles. J. Med. Chem. 55, 6438–6444 (2012).
Lund, G. et al. Antibody repertoire complexity and effector cell biology determined by assays for IgE-mediated basophil and T-cell activation. J. Immunol. Methods 383, 4–20 (2012).
Acknowledgements
We thank B. Wilson (University of New Mexico) for generously providing us with IgEDNP and the RBL cells, and P. Bryce (Northwestern University) for providing protocols and advice on the PCA model. We thank B. Boggess at the Mass Spectrometry and Proteomics Facility in the University of Notre Dame for the use of MS instrumentation. This work was supported by the National Institutes of Health–National Institute of Allergy and Infectious Diseases (grant number R03 AI085485 (to B.B.) and R01 AI095282 (to M.H.K.)).
Author information
Authors and Affiliations
Contributions
B.B. supervised and coordinated all of the research activities, designed the experiments and funded the study. M.W.H. designed the experiments, synthesized and characterized all of the molecules, performed the in vitro experiments and analyzed all of the data. T.K. designed and supervised the in vitro assays. M.H.K. designed and supervised the in vivo studies. A.P.S. performed the in vivo experiments. M.W.H., T.K. and B.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Figures 1–9. (PDF 1211 kb)
Rights and permissions
About this article
Cite this article
Handlogten, M., Kiziltepe, T., Serezani, A. et al. Inhibition of weak-affinity epitope-IgE interactions prevents mast cell degranulation. Nat Chem Biol 9, 789–795 (2013). https://doi.org/10.1038/nchembio.1358
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1358
This article is cited by
-
Exploring the potential and safety of quantum dots in allergy diagnostics
Microsystems & Nanoengineering (2023)
-
Determination of Crucial Immunogenic Epitopes in Major Peanut Allergy Protein, Ara h2, via Novel Nanoallergen Platform
Scientific Reports (2017)
-
The production and regulation of IgE by the immune system
Nature Reviews Immunology (2014)
-
Allergy's Achilles' heel?
Nature Chemical Biology (2013)