Abstract
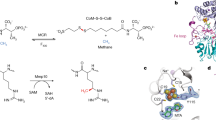
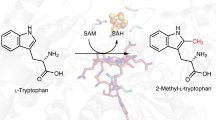
How living organisms create carbon-sulfur bonds during the biosynthesis of critical sulfur-containing compounds is still poorly understood. The methylthiotransferases MiaB and RimO catalyze sulfur insertion into tRNAs and ribosomal protein S12, respectively. Both belong to a subgroup of radical–S-adenosylmethionine (radical-SAM) enzymes that bear two [4Fe-4S] clusters. One cluster binds S-adenosylmethionine and generates an Ado• radical via a well-established mechanism. However, the precise role of the second cluster is unclear. For some sulfur-inserting radical-SAM enzymes, this cluster has been proposed to act as a sacrificial source of sulfur for the reaction. In this paper, we report parallel enzymological, spectroscopic and crystallographic investigations of RimO and MiaB, which provide what is to our knowledge the first evidence that these enzymes are true catalysts and support a new sulfation mechanism involving activation of an exogenous sulfur cosubstrate at an exchangeable coordination site on the second cluster, which remains intact during the reaction.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Fontecave, M., Ollagnier-de-Choudens, S. & Mulliez, E. Biological radical sulfur insertion reactions. Chem. Rev. 103, 2149–2166 (2003).
Jenner, L., Demeshkina, N., Yusupova, G. & Yusupov, M. Structural rearrangements of the ribosome at the tRNA proofreading step. Nat. Struct. Mol. Biol. 17, 1072–1078 (2010).
Wei, F.Y. et al. Deficit of tRNA(Lys) modification by Cdkal1 causes the development of type 2 diabetes in mice. J. Clin. Invest. 121, 3598–3608 (2011).
Atta, M. et al. S-Adenosylmethionine–dependent radical-based modification of biological macromolecules. Curr. Opin. Struct. Biol. 20, 684–692 (2010).
Kowalak, J.A. & Walsh, K.A. β-methylthio-aspartic acid: identification of a novel posttranslational modification in ribosomal protein S12 from Escherichia coli. Protein Sci. 5, 1625–1632 (1996).
Sofia, H.J., Chen, G., Hetzler, B.G., Reyes-Spindola, J.F. & Miller, N.E. Radical SAM, a novel protein superfamily linking unresolved steps in familiar biosynthetic pathways with radical mechanisms: functional characterization using new analysis and information visualization methods. Nucleic Acids Res. 29, 1097–1106 (2001).
Lee, K.H. et al. Characterization of RimO, a new member of the methylthiotransferase subclass of the radical SAM superfamily. Biochemistry 48, 10162–10174 (2009).
Hernández, H.L. et al. MiaB, a bifunctional radical-S-adenosylmethionine enzyme involved in the thiolation and methylation of tRNA, contains two essential [4Fe-4S] clusters. Biochemistry 46, 5140–5147 (2007).
Arragain, S. et al. Identification of eukaryotic and prokaryotic methylthiotransferase for biosynthesis of 2-methylthio-N-6-threonylcarbamoyladenosine in tRNA. J. Biol. Chem. 285, 28425–28433 (2010).
Arragain, S. et al. Post-translational modification of ribosomal proteins. structural and functional characterization of RimO from Thermotoga maritima, a radical S-adenosylmethionine methylthiotransferase. J. Biol. Chem. 285, 5792–5801 (2010).
Anantharaman, V., Koonin, E.V. & Aravind, L. TRAM, a predicted RNA-binding domain, common to tRNA uracil methylation and adenine thiolation enzymes. FEMS Microbiol. Lett. 197, 215–221 (2001).
Booker, S.J., Cicchillo, R.M. & Grove, T.L. Self-sacrifice in radical S-adenosylmethionine proteins. Curr. Opin. Chem. Biol. 11, 543–552 (2007).
Ugulava, N.B., Sacanell, C.J. & Jarrett, J.T. Spectroscopic changes during a single turnover of biotin synthase: destruction of a [2Fe-2S] cluster accompanies sulfur insertion. Biochemistry 40, 8352–8358 (2001).
Frey, P.A., Hegeman, A.D. & Ruzicka, F.J. The radical SAM superfamily. Crit. Rev. Biochem. Mol. Biol. 43, 63–88 (2008).
Pierrel, F., Douki, T., Fontecave, M. & Atta, M. MiaB protein is a bifunctional radical-S-adenosylmethionine enzyme involved in thiolation and methylation of tRNA. J. Biol. Chem. 279, 47555–47563 (2004).
Vey, J.L. & Drennan, C.L. Structural insights into radical generation by the radical SAM superfamily. Chem. Rev. 111, 2487–2506 (2011).
Duschene, K.S., Veneziano, S.E., Silver, S.C. & Broderick, J.B. Control of radical chemistry in the AdoMet radical enzymes. Curr. Opin. Chem. Biol. 13, 74–83 (2009).
Zhang, Q. & Liu, W. Complex biotransformations catalyzed by radical S-adenosylmethionine enzymes. J. Biol. Chem. 286, 30245–30252 (2011).
Branden, C. & Tooze, J. Introduction to Protein Structure Vol. 2, 59–60 (Garland Publishing, New York, 1999).
Holm, L. & Rosenstrom, P. Dali server: conservation mapping in 3D. Nucleic Acids Res. 38, W545–W549 (2010).
Andreeva, A. et al. Data growth and its impact on the SCOP database: new developments. Nucleic Acids Res. 36, D419–D425 (2008).
Baikalov, I. et al. Structure of the Escherichia coli response regulator NarL. Biochemistry 35, 11053–11061 (1996).
Schnell, R., Agren, D. & Schneider, G. 1.9-Å structure of the signal receiver domain of the putative response regulator NarL from Mycobacterium tuberculosis. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 64, 1096–1100 (2008).
Maris, A.E. et al. Dimerization allows DNA target site recognition by the NarL response regulator. Nat. Struct. Biol. 9, 771–778 (2002).
Porter, S.L., Wadhams, G.H. & Armitage, J.P. Signal processing in complex chemotaxis pathways. Nat. Rev. Microbiol. 9, 153–165 (2011).
Berkovitch, F., Nicolet, Y., Wan, J.T., Jarrett, J.T. & Drennan, C.L. Crystal structure of biotin synthase, an S-adenosylmethionine–dependent radical enzyme. Science 303, 76–79 (2004).
Hänzelmann, P. & Schindelin, H. Crystal structure of the S-adenosylmethionine-dependent enzyme MoaA and its implications for molybdenum cofactor deficiency in humans. Proc. Natl. Acad. Sci. USA 101, 12870–12875 (2004).
Lees, N.S. et al. ENDOR spectroscopy shows that guanine N1 binds to [4Fe-4S] cluster II of the S-adenosylmethionine-dependent enzyme MoaA: mechanistic implications. J. Am. Chem. Soc. 131, 9184–9185 (2009).
Zheng, B., Chen, X.D., Zheng, S.L. & Holm, R.H. Selenium as a structural surrogate of sulfur: template-assisted assembly of five types of tungsten-iron-sulfur/selenium clusters and the structural fate of chalcogenide reactants. J. Am. Chem. Soc. 134, 6479–6490 (2012).
Wilker, J.J. & Lippard, S.J. Methylation of iron-sulfur complexes by trimethyl phosphate. Inorg. Chem. 38, 3569–3574 (1999).
Petrey, D. & Honig, B. GRASP2: visualization, surface properties, and electrostatics of macromolecular structures and sequences. Methods Enzymol. 374, 492–509 (2003).
Iwig, D.F. & Booker, S.J. Insight into the polar reactivity of the onium chalcogen analogues of S-adenosyl-L-methionine. Biochemistry 43, 13496–13509 (2004).
Tse Sum Bui, B., Mattioli, T.A., Florentin, D., Bolbach, G. & Marquet, A. Escherichia coli biotin synthase produces selenobiotin. Further evidence of the involvement of the [2Fe-2S]2+ cluster in the sulfur insertion step. Biochemistry 45, 3824–3834 (2006).
Syper, L. & Mlochowski, J. The convenient syntheses of organoselenium reagents. Synthesis, 439–442 (1984).
Fish, W.W. Rapid colorimetric micromethod for the quantitation of complexed iron in biological samples. Methods Enzymol. 158, 357–364 (1988).
Beinert, H. Semi-micro methods for analysis of labile sulfide and of labile sulfide plus sulfane sulfur in unusually stable iron sulfur proteins. Anal. Biochem. 131, 373–378 (1983).
Then, J. & Truper, H.G. Sulfide oxidation in Ectothiorhodospira abdelmalekii—evidence for the catalytic role of cytochrome c-551. Arch. Microbiol. 135, 254–258 (1983).
Pierrel, F., Hernandez, H.L., Johnson, M.K., Fontecave, M. & Atta, M. MiaB protein from Thermotoga maritima—characterization of an extremely thermophilic tRNA-methylthiotransferase. J. Biol. Chem. 278, 29515–29524 (2003).
Gehrke, C.W. & Kuo, K.C. Ribonucleoside analysis by reversed-phase high-performance liquid-chromatography. J. Chromatogr. 471, 3–36 (1989).
te Velde, G. & Baerends, E.J. Numerical integration for polyatomic systems. J. Comput. Phys. 99, 84–98 (1992).
Vosko, S.H., Wilk, L. & Nusair, M. Accurate spin-dependent electron liquid correlation energies for local spin-density calculations—a critical analysis. Can. J. Phys. 58, 1200–1211 (1980).
Becke, A.D. Density-functional exchange-energy approximation with correct asymptotic-behavior. Phys. Rev. A 38, 3098–3100 (1988).
Perdew, J.P. Density-functional approximation for the correlation energy of the inhomogeneous electron gas. Phys. Rev. B Condens. Matter 33, 8822–8824 (1986).
Szilagyi, R.K. & Winslow, M.A. On the accuracy of density functional theory for iron-sulfur clusters. J. Comput. Chem. 27, 1385–1397 (2006).
Noodleman, L., Peng, C.Y., Case, D.A. & Mouesca, J.M. Orbital interactions, electron delocalization and spin coupling in iron-sulfur clusters. Coord. Chem. Rev. 144, 199–244 (1995).
Acton, T.B. et al. Robotic cloning and protein production platform of the Northeast Structural Genomics Consortium. Methods Enzymol. 394, 210–243 (2005).
Jansson, M. et al. High-level production of uniformly 15N- and 13C-enriched fusion proteins in Escherichia coli. J. Biomol. NMR 7, 131–141 (1996).
Doublié, S. et al. Crystallization and preliminary X-ray analysis of the 9 kDa protein of the mouse signal recognition particle and the selenomethionyl-SRP9. FEBS Lett. 384, 219–221 (1996).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Macromol. Crystallogr. A. 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
McRee, D.E. XtalView Xfit—a versatile program for manipulating atomic coordinates and electron density. J. Struct. Biol. 125, 156–165 (1999).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Acknowledgements
We thank R. Abramowitz and J. Schwanof for assistance with synchrotron data collection, B. Gibney for advice on Fe-S reconstitution for crystallization and R. Breslow for discussion of the reaction mechanism. We thank O. Hamelin for GC/MS analysis and J.-M. Moulis for providing 77Se (both from Chemistry and Biology of Metals Laboratory, Grenoble). This work was supported by the US National Institutes of Health Protein Structure Initiative grants U54-GM074958 and U54-GM094597 to the NeSG (http://www.nesg.org/), a Groupement d′Intérêt Scientifique–CNRS fellowship to S.A., Agence Nationale de la Recherche–Blanc 2010 grant INSERAD and Région Rhône-Alpes grant CIBLE 2008-2011.
Author information
Authors and Affiliations
Contributions
E.M., S.A., M.A. and M.F. designed the biochemical and enzymological experiments, which were conducted by E.M. and S.A. J.-M.M. and S.G. designed and conducted the EPR, HYSCORE and DFT experiments. S.K.-J. conducted the HPLC/MS experiments. T.B.A., R.X., J.F.H., J.S. and G.T.M. designed the target selection and protein purification–crystallization pipeline of the NeSG, which purified a wide variety of MTTases for this project. F.F., with advice from J.F.H., developed reconstitution methods for crystallization, which was performed by M.H. F.F. solved and refined related crystal structures. M.F., E.M., J.F.H., F.F., M.A. and G.T.M. interpreted the results, and M.F., E.M., J.F.H. and F.F. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Note 1 (PDF 3738 kb)
Rights and permissions
About this article
Cite this article
Forouhar, F., Arragain, S., Atta, M. et al. Two Fe-S clusters catalyze sulfur insertion by radical-SAM methylthiotransferases. Nat Chem Biol 9, 333–338 (2013). https://doi.org/10.1038/nchembio.1229
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1229
This article is cited by
-
Monitoring Fe–S cluster occupancy across the E. coli proteome using chemoproteomics
Nature Chemical Biology (2023)
-
Radical thioesterification via nickel-catalysed sensitized electron transfer
Nature Synthesis (2023)
-
Structural basis for tRNA methylthiolation by the radical SAM enzyme MiaB
Nature (2021)
-
Identification of the key functional genes in salt-stress tolerance of Cyanobacterium Phormidium tenue using in silico analysis
3 Biotech (2021)
-
Bio-additive-based screening: toward evaluation of the biocompatibility of chemical reactions
Nature Protocols (2019)