Abstract
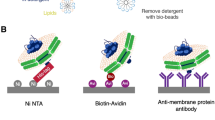
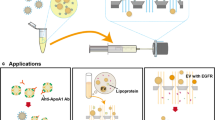
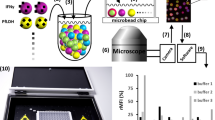
Membrane proteomics, the large-scale global analysis of membrane proteins, is often constrained by the efficiency of separating and extracting membrane proteins. Recent approaches involve conjugating membrane proteins with the small molecule biotin and using the receptor streptavidin to extract the labelled proteins. Despite the many advantages of this method, several shortcomings remain, including potential contamination by endogenously biotinylated molecules and interference by streptavidin during analytical stages. Here, we report a supramolecular fishing method for membrane proteins using the synthetic receptor–ligand pair cucurbit[7]uril–1-trimethylammoniomethylferrocene (CB[7]–AFc). CB[7]-conjugated beads selectively capture AFc-labelled proteins from heterogeneous protein mixtures, and AFc-labelling of cells results in the efficient capture of membrane proteins by these beads. The captured proteins can be recovered easily at room temperature by treatment with a strong competitor such as 1,1′-bis(trimethylammoniomethyl)ferrocene. This synthetic but biocompatible host–guest system may be a useful alternative to streptavidin–biotin for membrane proteomics as well as other biological and biotechnological applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Alberts, B. et al. Molecular Biology of the Cell (Garland Science, 2002).
Rabilloud, T. Membrane proteins ride shotgun. Nature Biotechnol. 21, 508–510 (2003).
Josic, D. J. & Clifton, G. Mammalian plasma membrane proteomics. Proteomics 7, 3010–3029 (2007).
Leonard, R. T. & Vanderwoude, W. J. Isolation of plasma membranes from corn roots by sucrose density gradient centrifugation. Plant Physiol. 57, 105–114 (1976).
Cao, R. et al. Integration of a two-phase partition method into proteomics research on rat liver plasma membrane proteins. J. Proteome Res. 5, 634–642 (2006).
Josic, D. et al. Use of selective extraction and fast chromatographic separation combined with electrophoretic methods for mapping of membrane proteins. Electrophoresis 26, 2809–2822 (2005).
Zhang L. et al. Immunoaffinity purification of plasma membrane with secondary antibody superparamagnetic beads for proteomic analysis. J. Proteome Res. 6, 34–43 (2007).
Lawson, E. L. et al. Use of magnetic beads with immobilized monoclonal antibodies for isolation of highly pure plasma membranes. Electrophoresis 27, 2747–2758 (2006).
Takechi, R., Taniguchi, A., Ebara, S., Fukui, T. & Watanabe, T. Biotin deficiency affects the proliferation of human embryonic palatal mesenchymal cells in culture. J. Nutr. 138, 680–684 (2008).
Zhang, W., Zhou, G., Zhao, Y., White, M. A. & Zhao, Y. Affinity enrichment of plasma membrane for proteomics analysis. Electrophoresis 24, 2855–2863 (2003).
Jaffrey, S. R., Erdjument-Bromage, H., Ferris, C. D., Tempst, P. & Snyder, S. H. Protein S-nitrosylation: a physiological signal for neuronal nitric oxide. Nat. Cell Biol. 3, 193–197 (2001).
Fonovic, M., Verhelst, S. H. L., Sorum, M. T. & Bogyo, M. Proteomics evaluation of chemically cleavable activity-based probes. Mol. Cell Proteomics 6, 1761–1770 (2007).
González, M. et al. Interaction of biotin with streptavidin thermostability and conformational changes upon binding. J. Biol. Chem. 272, 11288–11294 (1997).
Chivers, C. E. et al. A streptavidin variant with slower biotin dissociation and increased mechanostability. Nature Methods 7, 391–393 (2010).
Nguyen, T., Joshi, N. S. & Francis, M. B. An affinity-based method for the purification of fluorescently-labeled biomolecules. Bioconjug. Chem. 17, 869–872 (2006).
Chung, J. A. et al. Purification of prenylated proteins by affinity chromatography on cyclodextrin-modified agarose. Anal. Biochem. 386, 1–8 (2009).
Lee, J. W., Samal, S., Selvapalam, N., Kim, H.-J. & Kim, K. Cucurbituril homologues and derivatives: new opportunities in supramolecular chemistry. Acc. Chem. Res. 36, 621–630 (2003).
Lagona, J., Mukhopadhyay, P., Chakrabarti, S. & Isaacs, L. The cucurbit[n]uril family. Angew. Chem. Int. Ed. 44, 4844–4870 (2005).
Kim, K. et al. Functionalized cucurbiturils and their applications. Chem. Soc. Rev. 36, 267–279 (2007).
Jeon, W. S. et al. Complexation of ferrocene derivatives by the cucurbit[7]uril host: a comparative study of the cucurbituril and cyclodextrin host families. J. Am. Chem. Soc. 127, 12984–12989 (2005).
Liu, S. et al. The cucurbit[n]uril family: prime component for self-sorting systems. J. Am. Chem. Soc. 127, 15959–15967 (2005).
Rekharsky, M. V. et al. A synthetic host–guest system achieves avidin–biotin affinity by overcoming enthalpy–entropy compensation. Proc. Natl Acad. Sci USA 104, 20737–20742 (2007).
Hennig, A., Bakirci, H. & Nau, W. M. Label-free continuous enzyme assays with macrocycle–fluorescent dye complexes. Nature Methods 4, 629–632 (2007)
Shaikh, M. et al. Salt-induced guest relocation from a macrocyclic cavity into a biomolecular pocket: interplay between cucurbit[7]uril and albumin. Chem. Commun. 3681–3683 (2008).
Ghosh, S. & Isaacs, L. Biological catalysis regulated by cucurbit[7]uril molecular containers. J. Am. Chem. Soc. 132, 4445–4454 (2010).
Hwang, I. et al. Noncovalent immobilization of proteins on a solid surface by cucurbit[7]uril–ferrocenemethylammonium pair, a potential replacement of biotin–avidin pair. J. Am. Chem. Soc. 129, 4170–4171 (2007).
Young, J. F. et al. Strong and reversible monovalent supramolecular protein immobilization. ChemBioChem 11, 180–183 (2010).
Prulière-Escabasse, V. et al. Modulation of epithelial sodium channel trafficking and function by sodium 4-phenylbutyrate in human nasal epithelial cells. J. Biol. Chem. 282, 34048–34057 (2007).
Wong, K. A. & Lodish, H. F. A revised model for AMP-activated protein kinase structure: the alpha-subunit binds to both the beta- and gamma-subunits although there is no direct binding between the beta- and gamma-subunits. J. Biol. Chem. 281, 36434–36442 (2006).
Tan, S., Tan, H. T. & Chung, M. C. M. Membrane proteins and membrane proteomics. Proteomics 8, 3924–3932 (2008).
Acknowledgements
This work was supported by the Creative Research Initiative and Brain Korea 21 Program of the Korean Ministry of Education, Science, and Technology (MOEST), the World Class University (WCU) program through the Korea Science and Engineering Foundation funded by MOEST (project no. R31-2008-000-10059-0). The authors also thank J. Mark Kim for helpful discussions.
Author information
Authors and Affiliations
Contributions
K.K. and S.H.R conceived and designed the project. D.W.L., K.M.P., B.M., K.S., P.S. and N.S. performed the synthesis. D.W.L. and H.J. performed the protein and cell experiments. D.W.L., K.M.P. and S.H.H. analysed the data. T.L. performed mass spectrometry. J.K. provided invaluable advice. K.K., D.W.L. and K.M.P. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 879 kb)
Rights and permissions
About this article
Cite this article
Lee, DW., Park, K., Banerjee, M. et al. Supramolecular fishing for plasma membrane proteins using an ultrastable synthetic host–guest binding pair. Nature Chem 3, 154–159 (2011). https://doi.org/10.1038/nchem.928
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.928
This article is cited by
-
Adaptive insertion of a hydrophobic anchor into a poly(ethylene glycol) host for programmable surface functionalization
Nature Chemistry (2023)
-
Cucurbituril curiosities
Nature Chemistry (2023)
-
Evaluation of the stability of cucurbit[8]uril-based ternary host−guest complexation in physiological environment and the fabrication of a supramolecular theranostic nanomedicine
Journal of Nanobiotechnology (2021)
-
Coordination cages as permanently porous ionic liquids
Nature Chemistry (2020)
-
Purification of protein therapeutics via high-affinity supramolecular host–guest interactions
Nature Biomedical Engineering (2020)