Abstract
Protein aggregation is mostly viewed as deleterious and irreversible causing several pathologies. However, reversible protein aggregation has recently emerged as a novel concept for cellular regulation. Here, we characterize stress-induced, reversible aggregation of yeast pyruvate kinase, Cdc19. Aggregation of Cdc19 is regulated by oligomerization and binding to allosteric regulators. We identify a region of low compositional complexity (LCR) within Cdc19 as necessary and sufficient for reversible aggregation. During exponential growth, shielding the LCR within tetrameric Cdc19 or phosphorylation of the LCR prevents unscheduled aggregation, while its dephosphorylation is necessary for reversible aggregation during stress. Cdc19 aggregation triggers its localization to stress granules and modulates their formation and dissolution. Reversible aggregation protects Cdc19 from stress-induced degradation, thereby allowing cell cycle restart after stress. Several other enzymes necessary for G1 progression also contain LCRs and aggregate reversibly during stress, implying that reversible aggregation represents a conserved mechanism regulating cell growth and survival.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Broach, J. R. Nutritional control of growth and development in yeast. Genetics 192, 73–105 (2012).
De Virgilio, C. & Loewith, R. Cell growth control: little eukaryotes make big contributions. Oncogene 25, 6392–6415 (2006).
Kroemer, G., Marino, G. & Levine, B. Autophagy and the integrated stress response. Mol. Cell 40, 280–293 (2010).
Protter, D. S. & Parker, R. Principles and properties of stress granules. Trends Cell Biol. 26, 668–679 (2016).
Buchan, J. R. mRNP granules. Assembly, function, and connections with disease. RNA Biol. 11, 1019–1030 (2014).
Wippich, F. et al. Dual specificity kinase DYRK3 couples stress granule condensation/dissolution to mTORC1 signaling. Cell 152, 791–805 (2013).
Mnaimneh, S. et al. Exploration of essential gene functions via titratable promoter alleles. Cell 118, 31–44 (2004).
Narayanaswamy, R. et al. Widespread reorganization of metabolic enzymes into reversible assemblies upon nutrient starvation. Proc. Natl Acad. Sci. USA 106, 10147–10152 (2009).
Saad, S., Peter, M. & Dechant, R. In scarcity and abundance: metabolic signals regulating cell growth. Physiology (Bethesda) 28, 298–309 (2013).
Kryndushkin, D., Pripuzova, N., Burnett, B. G. & Shewmaker, F. Non-targeted identification of prions and amyloid-forming proteins from yeast and mammalian cells. J. Biol. Chem. 288, 27100–27111 (2013).
Wallace, E. W. et al. Reversible, specific, active aggregates of endogenous proteins assemble upon heat stress. Cell 162, 1286–1298 (2015).
Alberti, S., Mateju, D., Mediani, L. & Carra, S. Granulostasis: protein quality control of RNP granules. Front. Mol. Neurosci. 10, 84 (2017).
Aguzzi, A. & Altmeyer, M. Phase separation: linking cellular compartmentalization to disease. Trends Cell Biol. 26, 547–558 (2016).
Molliex, A. et al. Phase separation by low complexity domains promotes stress granule assembly and drives pathological fibrillization. Cell 163, 123–133 (2015).
Patel, A. et al. A liquid-to-solid phase transition of the ALS protein FUS accelerated by disease mutation. Cell 162, 1066–1077 (2015).
Kato, M. et al. Cell-free formation of RNA granules: low complexity sequence domains form dynamic fibers within hydrogels. Cell 149, 753–767 (2012).
Xiang, S. et al. The LC domain of hnRNPA2 adopts similar conformations in hydrogel polymers, liquid-like droplets, and nuclei. Cell 163, 829–839 (2015).
Letunic, I., Doerks, T. & Bork, P. SMART: recent updates, new developments and status in 2015. Nucleic Acids Res. 43, D257–D260 (2015).
Jurica, M. S. et al. The allosteric regulation of pyruvate kinase by fructose-1,6-bisphosphate. Structure 6, 195–210 (1998).
Anastasiou, D. et al. Pyruvate kinase M2 activators promote tetramer formation and suppress tumorigenesis. Nat. Chem. Biol. 8, 839–847 (2012).
Merrins, M. J., Van Dyke, A. R., Mapp, A. K., Rizzo, M. A. & Satin, L. S. Direct measurements of oscillatory glycolysis in pancreatic islet beta-cells using novel fluorescence resonance energy transfer (FRET) biosensors for pyruvate kinase M2 activity. J. Biol. Chem. 288, 33312–33322 (2013).
Yang, W. et al. PKM2 phosphorylates histone H3 and promotes gene transcription and tumorigenesis. Cell 150, 685–696 (2012).
Yang, W. et al. Nuclear PKM2 regulates β-catenin transactivation upon EGFR activation. Nature 480, 118–122 (2011).
Gao, X., Wang, H., Yang, J. J., Liu, X. & Liu, Z. R. Pyruvate kinase M2 regulates gene transcription by acting as a protein kinase. Mol. Cell 45, 598–609 (2012).
Fenton, A. W. & Blair, J. B. Kinetic and allosteric consequences of mutations in the subunit and domain interfaces and the allosteric site of yeast pyruvate kinase. Arch. Biochem. Biophys. 397, 28–39 (2002).
Kresnowati, M. T. A. P. et al. When transcriptome meets metabolome: fast cellular responses of yeast to sudden relief of glucose limitation. Mol. Syst. Biol. 2, 49 (2006).
Boles, E. et al. Characterization of a glucose-repressed pyruvate kinase (Pyk2p) in Saccharomyces cerevisiae that is catalytically insensitive to fructose-1,6-bisphosphate. J. Bacteriol. 179, 2987–2993 (1997).
Grüning, N.-M. et al. Pyruvate kinase triggers a metabolic feedback loop that controls redox metabolism in respiring cells. Cell Metab. 14, 415–427 (2011).
Protter, D. S. & Parker, R. Principles and properties of stress granules. Trends Cell Biol. 26, 668–679 (2016).
Buchan, J. R., Muhlrad, D. & Parker, R. P bodies promote stress granule assembly in Saccharomyces cerevisiae. J. Cell Biol. 183, 441–455 (2008).
Brengues, M. & Parker, R. Accumulation of polyadenylated mRNA, Pab1p, eIF4E, and eIF4G with P-bodies in Saccharomyces cerevisiae. Mol. Biol. Cell 18, 2592–2602 (2007).
Beckmann, B. M. et al. The RNA-binding proteomes from yeast to man harbour conserved enigmRBPs. Nat. Commun. 6, 10127 (2015).
Kramer, K. et al. Photo-cross-linking and high-resolution mass spectrometry for assignment of RNA-binding sites in RNA-binding proteins. Nat. Methods 11, 1064–1070 (2014).
Berchowitz, L. E. et al. Regulated formation of an amyloid-like translational repressor governs gametogenesis. Cell 163, 406–418 (2015).
Han, T. W. et al. Cell-free formation of RNA granules: bound RNAs identify features and components of cellular assemblies. Cell 149, 768–779 (2012).
Kwon, I. et al. Phosphorylation-regulated binding of RNA polymerase II to fibrous polymers of low-complexity domains. Cell 155, 1049–1060 (2013).
Wu, R. et al. A large-scale method to measure absolute protein phosphorylation stoichiometries. Nat. Methods 8, 677–683 (2011).
Jorgensen, P. et al. A dynamic transcriptional network communicates growth potential to ribosome synthesis and critical cell size. Genes Dev. 18, 2491–2505 (2004).
Dechant, R., Saad, S., Ibanez, A. J. & Peter, M. Cytosolic pH regulates cell growth through distinct GTPases, Arf1 and Gtr1, to promote Ras/PKA and TORC1 activity. Mol. Cell 55, 409–421 (2014).
Urban, J. et al. Sch9 is a major target of TORC1 in Saccharomyces cerevisiae. Mol. Cell 26, 663–674 (2007).
Quintas, A., Vaz, D. C., Cardoso, I., Saraiva, M. J. & Brito, R. M. Tetramer dissociation and monomer partial unfolding precedes protofibril formation in amyloidogenic transthyretin variants. J. Biol. Chem. 276, 27207–27213 (2001).
Westermark, G. T. & Westermark, P. Prion-like aggregates: infectious agents in human disease. Trends Mol. Med. 16, 501–507 (2010).
Castello, A., Hentze, M. W. & Preiss, T. Metabolic enzymes enjoying new partnerships as RNA-binding proteins. Trends Endocrinol. Metab. 26, 746–757 (2015).
Dechant, R. et al. Cytosolic pH is a second messenger for glucose and regulates the PKA pathway through V-ATPase. EMBO J. 29, 2515–2526 (2010).
Pelet, S., Dechant, R., Lee, S. S., van Drogen, F. & Peter, M. An integrated image analysis platform to quantify signal transduction in single cells. Integr. Biol. 4, 1274–1282 (2012).
Feng, Y. et al. Global analysis of protein structural changes in complex proteomes. Nat. Biotechnol. 32, 1036–1044 (2014).
Acknowledgements
We thank S. Alberti (Max Planck Institute of Molecular Cell Biology and Genetics, Dresden, Germany), M. Ralser (Department of Biochemistry and Cambridge Systems Biology Centre, University of Cambridge, Cambridge, UK) and K. Weis (Institute of Biochemistry, Department of Biology, ETH Zürich, Zürich, Switzerland) for providing reagents; S. McKnight, D. Frantz and L. Berchowitz for help with b-isox experiments; A. Timofiiva for help with microscopy; and T. Mayor, A. Smith, P. Kimmig and members of the Peter laboratory for helpful discussions and comments on the manuscript. This work was funded by the Swiss National Science Foundation (SNF, project grants 31003A_166513 to R.D. and 310030B_160312 to M.P.), the European Research Council (ERC, Rubinet) and ETH Zürich.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.S., M.P. and R.D.; investigation: S.S., G.C. and Y.F.; formal analysis: R.D.; writing—original draft preparation: S.S. and R.D.; writing—review and editing: S.S., G.C., Y.F., P.P., M.P. and R.D.; visualization: S.S. and R.D.; supervision: P.P., M.P. and R.D.; funding acquisition: M.P. and R.D.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cdc19 forms reversible aggregates upon stress and tetrameric Cdc19ΔPEP is unable to support cell growth in yeast.
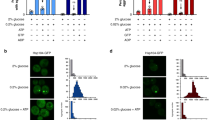
(a,b) Major protein degradation pathways do not significantly contribute to reversibility of Cdc19 aggregates. Cells expressing Cdc19-GFP and the indicated mutations were grown to stationary phase, loaded into a microfluidic chip and reversibility of aggregates was scored upon glucose readdition in the presence of (a) PMSF (prevents vacuolar degradation) or (b) MG132 to inhibit proteasomal degradation. Note that expression of the Pdr1-Cyc8 fusion necessary for efficient inhibition of proteasomal activity by MG132. The reason for faster solubilisation of aggregates in this strain is unclear. Arrowheads indicate frames with near-complete solubilisation of detectable aggregates. Time relative to glucose readdition is indicated. Data shown are representative of 3 independent experiments. (c) Cdc19-aggregates in stationary phase are solid-like. Cells were grown to stationary phase and Cdc19-GFP aggregates were subjected to FRAP analysis as described in Fig. 1. Quantification of fluorescence recovery for all analyzed cells is shown as the mean of the relative fluorescence intensity as a function of time (red line) together with the individual cell traces (light and dark gray) (n = 3 independent experiments). (d) Cells harboring the tetO7-CDC19 allele expressing wild-type or mutant Cdc19 or an empty vector were grown in the presence or absence of doxycycline (10 μg/ml, added at t = 0h) and growth was analyzed using a BioLector (m2p-labs). Accumulation of biomass is shown as a function of time. Data shown are representative of 2 independent experiments. (e) Cytoplasmic acidification is not required for the formation of Cdc19 aggregates. Cells were grown as in Fig. 1a and representative images of Cdc19-GFP upon glucose starvation in C-starvation media (pH 4.6 or pH 7.4) or PBS (pH 7.4) from 3 independent experiments are shown. In all images, the scale bar represents 3 μm. Statistical source data for c can be found in Supplementary Table 4.
Supplementary Figure 2 Cdc19 colocalizes with RNA containing stress granules and failure of recruitment renders Cdc19 unstable.
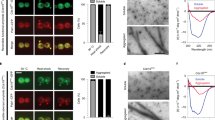
(a) Cdc19 aggregates colocalize with the stress granule marker Pub1. Cells expressing Cdc19-GFP and Pub1-RFP were starved and localization of Cdc19 aggregates relative to Pub1 was analyzed. Data shown are representative of 2 independent experiments. (b) Cdc19 aggregates colocalize with the stress granule marker Pab1, but not with the P-body marker Edc3. Cells expressing Cdc19-GFP, Pab1-CFP and Edc3-RFP were grown as in Fig. 1 and timing of the appearance of P-bodies and stress granules and their localization was analyzed relative to Cdc19 aggregation. Data shown are representative of 3 independent experiments. Time relative to glucose readdition is indicated. Note that Cdc19-GFP was imaged using YFP filter sets to avoid any spectral mixing in cells expressing Pab1-CFP. (c) Cells expressing wild-type Cdc19-TAP or Cdc19ΔPEP-TAP and Hxk2-TAP as control were grown in SD media and UV treated. Cross-linked protein-mRNA complexes were isolated using poly-dT and analyzed by western-blot. Data shown are representative of 3 independent experiments. (d) Cells expressing wild-type or mutant Cdc19-GFP expressing Pab1-CFP were starved for glucose and localization of Cdc19 and Pab1 was analyzed at the indicated time-points. Note that Cdc19-GFP was imaged using YFP filter sets to avoid any spectral mixing in cells expressing Pab1-CFP. Data shown are representative of 3 independent experiments. (e) Cells expressing wild-type or mutant Cdc19 were grown as in Fig. 3g and relative abundance of Cdc19 was determined by mass-spectrometry and plotted as the mean ± S.E.M. of three technical replicates. In all images, the scale bar represents 3 μm. Uncropped blot is shown in Supplementary Fig. 5.
Supplementary Figure 3 Cdc19 forms Thioflavin-T positive aggregates in vitro and LCRs might generally contribute to aggregation.
(a) Wild-type and mutant Cdc19 incubated at the indicated temperatures were mixed with ThT as in Fig. 4e and emission spectra were recorded. Data shown are representative of 3 independent experiments. (b) Sbp1 and Ssa2 colocalize with stress granules upon glucose starvation. Cells expressing Sbp1-GFP or Ssa2-GFP and Pab1-CFP were grown as in Fig. 1 and scored for localization of the indicated proteins. Data shown are representative of 3 independent experiments. Note that Sbp1-GFP and Ssa2-GFP were imaged using YFP filter sets to avoid any spectral mixing in cells expressing Pab1-CFP. The scale bar represents 3 μm.
Supplementary Figure 4 Dephosphorylation of the Cdc19 LCR is required for reversible aggregation.
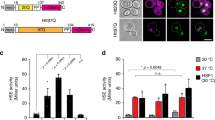
(a) Schematic representation of the MS-workflow to determine the fraction of phosphorylated peptides. Unphosphorylated peptides were measured with or without phosphatase treatment (PPase) and abundance relative to a control peptide was determined. (b) The LCR of Cdc19 is part of the subunit interface in Cdc19 tetramers. Crystal structure of Cdc19 showing a part of the subunit interface of Cdc19 tetramers. Blue: ribbon representation of Cdc19 with the region corresponding to the LCR shown in yellow. Thr372, Thr376, Ser377 and Ser385 are shown in space filling representation. Other subunits of the Cdc19 tetramer (light gray, dark-gray, red) are shown as surface representations. Note that Ser385 forms intersubunit contacts (hydrogen bond via side chain-OH), strongly suggesting that phosphorylation at this site would be incompatible with phosphorylation. (c) Cells expressing wild-type or mutant Cdc19-GFP expressing Pab1-CFP were starved for glucose and localization of Cdc19 and Pab1 was analyzed at the indicated time-points. Note that Cdc19-GFP was imaged using YFP filter sets to avoid any spectral mixing in cells expressing Pab1-CFP. Data shown are representative of 3 independent experiments. (d) Cells as in Fig. 7a were grown and analyzed for aggregate formation upon heat shock (30 min, 42 °C) and after recovery at 30 °C for 60 min. Representative images of 3 independent experiments for the indicated conditions are shown. In all images, the scale bar represents 3 μm.
Supplementary information
Supplementary Information
Supplementary Information (PDF 4512 kb)
Supplementary Table 1
Supplementary Information (XLSX 30 kb)
Supplementary Table 2
Supplementary Information (XLSX 28 kb)
Supplementary Table 3
Supplementary Information (XLSX 44 kb)
Supplementary Table 4
Supplementary Information (XLSX 145 kb)
Rights and permissions
About this article
Cite this article
Saad, S., Cereghetti, G., Feng, Y. et al. Reversible protein aggregation is a protective mechanism to ensure cell cycle restart after stress. Nat Cell Biol 19, 1202–1213 (2017). https://doi.org/10.1038/ncb3600
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3600
This article is cited by
-
Differentiation granules, a dynamic regulator of T. brucei development
Nature Communications (2024)
-
Identification of CDK gene family and functional analysis of CqCDK15 under drought and salt stress in quinoa
BMC Genomics (2023)
-
The Pga59 cell wall protein is an amyloid forming protein involved in adhesion and biofilm establishment in the pathogenic yeast Candida albicans
npj Biofilms and Microbiomes (2023)
-
Polymorphic amyloid nanostructures of hormone peptides involved in glucose homeostasis display reversible amyloid formation
Nature Communications (2023)
-
Activation of L-lactate oxidase by the formation of enzyme assemblies through liquid–liquid phase separation
Scientific Reports (2023)