Abstract
Cells test the rigidity of the extracellular matrix by applying forces to it through integrin adhesions. Recent measurements show that these forces are applied by local micrometre-scale contractions, but how contraction force is regulated by rigidity is unknown. Here we performed high temporal- and spatial-resolution tracking of contractile forces by plating cells on sub-micrometre elastomeric pillars. We found that actomyosin-based sarcomere-like contractile units (CUs) simultaneously moved opposing pillars in net steps of ∼2.5 nm, independent of rigidity. What correlated with rigidity was the number of steps taken to reach a force level that activated recruitment of α-actinin to the CUs. When we removed actomyosin restriction by depleting tropomyosin 2.1, we observed larger steps and higher forces that resulted in aberrant rigidity sensing and growth of non-transformed cells on soft matrices. Thus, we conclude that tropomyosin 2.1 acts as a suppressor of growth on soft matrices by supporting proper rigidity sensing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Wang, H. B., Dembo, M. & Wang, Y. L. Substrate flexibility regulates growth and apoptosis of normal but not transformed cells. Am. J. Physiol. Cell Physiol. 279, C1345–C1350 (2000).
Engler, A. J., Sen, S., Sweeney, H. L. & Discher, D. E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Ingber, D. E. Mechanobiology and diseases of mechanotransduction. Ann. Med. 35, 564–577 (2003).
Jaalouk, D. E. & Lammerding, J. Mechanotransduction gone awry. Nat. Rev. Mol. Cell Biol. 10, 63–73 (2009).
Freedman, V. H. & Shin, S. I. Cellular tumorigenicity in nude mice: correlation with cell growth in semi-solid medium. Cell 3, 355–359 (1974).
Stoker, M., O’Neill, C., Berryman, S. & Waxman, V. Anchorage and growth regulation in normal and virus-transformed cells. Int. J. Cancer 3, 683–693 (1968).
Vogel, V. & Sheetz, M. P. Cell fate regulation by coupling mechanical cycles to biochemical signaling pathways. Curr. Opin. Cell Biol. 21, 38–46 (2009).
Plotnikov, S. V., Pasapera, A. M., Sabass, B. & Waterman, C. M. Force fluctuations within focal adhesions mediate ECM-rigidity sensing to guide directed cell migration. Cell 151, 1513–1527 (2012).
Ghassemi, S. et al. Cells test substrate rigidity by local contractions on submicrometer pillars. Proc. Natl Acad. Sci. USA 109, 5328–5333 (2012).
Chan, C. E. & Odde, D. J. Traction dynamics of filopodia on compliant substrates. Science 322, 1687–1691 (2008).
Elosegui-Artola, A. et al. Rigidity sensing and adaptation through regulation of integrin types. Nat. Mater. 13, 631–637 (2014).
Choquet, D., Felsenfeld, D. P. & Sheetz, M. P. Extracellular matrix rigidity causes strengthening of integrin-cytoskeleton linkages. Cell 88, 39–48 (1997).
Jiang, G., Huang, A. H., Cai, Y., Tanase, M. & Sheetz, M. P. Rigidity sensing at the leading edge through αvβ3 integrins and RPTPα. Biophys. J. 90, 1804–1809 (2006).
Geeves, M. A., Hitchcock-DeGregori, S. E. & Gunning, P. W. A systematic nomenclature for mammalian tropomyosin isoforms. J. Muscle Res. Cell Motil. 36, 147–153 (2015).
Roca-Cusachs, P. et al. Integrin-dependent force transmission to the extracellular matrix by α-actinin triggers adhesion maturation. Proc. Natl Acad. Sci. USA 110, E1361–E1370 (2013).
Burkholder, T. J. & Lieber, R. L. Sarcomere length operating range of vertebrate muscles during movement. J. Exp. Biol. 204, 1529–1536 (2001).
Spudich, J. A., Huxley, H. E. & Finch, J. T. Regulation of skeletal muscle contraction. II. Structural studies of the interaction of the tropomyosin-troponin complex with actin. J. Mol. Biol. 72, 619–632 (1972).
Fischer, R. S., Fritz-Six, K. L. & Fowler, V. M. Pointed-end capping by tropomodulin3 negatively regulates endothelial cell motility. J. Cell Biol. 161, 371–380 (2003).
Giannone, G. et al. Periodic lamellipodial contractions correlate with rearward actin waves. Cell 116, 431–443 (2004).
Dubin-Thaler, B. J., Giannone, G., Dobereiner, H. G. & Sheetz, M. P. Nanometer analysis of cell spreading on matrix-coated surfaces reveals two distinct cell states and STEPs. Biophys. J. 86, 1794–1806 (2004).
Cai, Y. et al. Nonmuscle myosin IIA-dependent force inhibits cell spreading and drives F-actin flow. Biophys. J. 91, 3907–3920 (2006).
Niederman, R. & Pollard, T. D. Human platelet myosin. II. In vitro assembly and structure of myosin filaments. J. Cell Biol. 67, 72–92 (1975).
Schevzov, G., Whittaker, S. P., Fath, T., Lin, J. J. & Gunning, P. W. Tropomyosin isoforms and reagents. Bioarchitecture 1, 135–164 (2011).
Cox, S. et al. Bayesian localization microscopy reveals nanoscale podosome dynamics. Nat. Methods 9, 195–200 (2012).
Verkhovsky, A. B., Svitkina, T. M. & Borisy, G. G. Myosin II filament assemblies in the active lamella of fibroblasts: their morphogenesis and role in the formation of actin filament bundles. J. Cell Biol. 131, 989–1002 (1995).
Sweeney, H. L. & Houdusse, A. Structural and functional insights into the Myosin motor mechanism. Ann. Rev. Biophys. 39, 539–557 (2010).
Kaya, M. & Higuchi, H. Stiffness, working stroke, and force of single-myosin molecules in skeletal muscle: elucidation of these mechanical properties via nonlinear elasticity evaluation. Cell. Mol. Life Sci. 70, 4275–4292 (2013).
Gelles, J., Schnapp, B. J. & Sheetz, M. P. Tracking kinesin-driven movements with nanometre-scale precision. Nature 331, 450–453 (1988).
Little, M. A. et al. Steps and bumps: precision extraction of discrete states of molecular machines. Biophys. J. 101, 477–485 (2011).
Kuo, S. C., Gelles, J., Steuer, E. & Sheetz, M. P. A model for kinesin movement from nanometer-level movements of kinesin and cytoplasmic dynein and force measurements. J. Cell Sci. Suppl. 14, 135–138 (1991).
Svoboda, K., Schmidt, C. F., Schnapp, B. J. & Block, S. M. Direct observation of kinesin stepping by optical trapping interferometry. Nature 365, 721–727 (1993).
Kaya, M. & Higuchi, H. Nonlinear elasticity and an 8-nm working stroke of single myosin molecules in myofilaments. Science 329, 686–689 (2010).
Jiang, G., Giannone, G., Critchley, D. R., Fukumoto, E. & Sheetz, M. P. Two-piconewton slip bond between fibronectin and the cytoskeleton depends on talin. Nature 424, 334–337 (2003).
Perz-Edwards, R. J. et al. X-ray diffraction evidence for myosin-troponin connections and tropomyosin movement during stretch activation of insect flight muscle. Proc. Natl Acad. Sci. USA 108, 120–125 (2011).
Wu, S. et al. Electron tomography of cryofixed, isometrically contracting insect flight muscle reveals novel actin-myosin interactions. PLoS ONE 5, e12643 (2010).
Prager-Khoutorsky, M. et al. Fibroblast polarization is a matrix-rigidity-dependent process controlled by focal adhesion mechanosensing. Nat. Cell Biol. 13, 1457–1465 (2011).
Prasad, G. L., Fuldner, R. A. & Cooper, H. L. Expression of transduced tropomyosin 1 cDNA suppresses neoplastic growth of cells transformed by the ras oncogene. Proc. Natl Acad. Sci. USA 90, 7039–7043 (1993).
Raval, G. N. et al. Loss of expression of tropomyosin-1, a novel class II tumor suppressor that induces anoikis, in primary breast tumors. Oncogene 22, 6194–6203 (2003).
Bhattacharya, B., Prasad, G. L., Valverius, E. M., Salomon, D. S. & Cooper, H. L. Tropomyosins of human mammary epithelial cells: consistent defects of expression in mammary carcinoma cell lines. Cancer Res. 50, 2105–2112 (1990).
Piazzesi, G. et al. Skeletal muscle performance determined by modulation of number of myosin motors rather than motor force or stroke size. Cell 131, 784–795 (2007).
Kad, N. M., Kim, S., Warshaw, D. M., VanBuren, P. & Baker, J. E. Single-myosin crossbridge interactions with actin filaments regulated by troponin-tropomyosin. Proc. Natl Acad. Sci. USA 102, 16990–16995 (2005).
Liu, X. & Pollack, G. H. Stepwise sliding of single actin and myosin filaments. Biophys. J. 86, 353–358 (2004).
Debold, E. P., Patlak, J. B. & Warshaw, D. M. Slip sliding away: load-dependence of velocity generated by skeletal muscle myosin molecules in the laser trap. Biophys. J. 89, L34–L36 (2005).
Walcott, S., Fagnant, P. M., Trybus, K. M. & Warshaw, D. M. Smooth muscle heavy meromyosin phosphorylated on one of its two heads supports force and motion. J. Biol. Chem. 284, 18244–18251 (2009).
Fusi, L., Brunello, E., Reconditi, M., Piazzesi, G. & Lombardi, V. The non-linear elasticity of the muscle sarcomere and the compliance of myosin motors. J. Physiology 592, 1109–1118 (2014).
Luo, W. et al. Analysis of the local organization and dynamics of cellular actin networks. J. Cell Biol. 202, 1057–1073 (2013).
Erdmann, T. & Schwarz, U. S. Stochastic force generation by small ensembles of myosin II motors. Physical Rev. Lett. 108, 188101 (2012).
Kovacs, M., Thirumurugan, K., Knight, P. J. & Sellers, J. R. Load-dependent mechanism of nonmuscle myosin 2. Proc. Natl Acad. Sci. USA 104, 9994–9999 (2007).
Hai, C. M. & Murphy, R. A. Cross-bridge phosphorylation and regulation of latch state in smooth muscle. Am. J. Physiol. 254, C99–C106 (1988).
Hai, C. M. & Murphy, R. A. Regulation of shortening velocity by cross-bridge phosphorylation in smooth muscle. Am. J. Physiol. 255, C86–C94 (1988).
Galbraith, C. G., Yamada, K. M. & Sheetz, M. P. The relationship between force and focal complex development. J. Cell Biol. 159, 695–705 (2002).
Mahadev, K. et al. Suppression of the transformed phenotype of breast cancer by tropomyosin-1. Exp. Cell Res. 279, 40–51 (2002).
Wolfenson, H., Bershadsky, A., Henis, Y. I. & Geiger, B. Actomyosin-generated tension controls the molecular kinetics of focal adhesions. J. Cell Sci. 124, 1425–1432 (2011).
Lavelin, I. et al. Differential effect of actomyosin relaxation on the dynamic properties of focal adhesion proteins. PLoS ONE 8, e73549 (2013).
Moore, S. W., Roca-Cusachs, P. & Sheetz, M. P. Stretchy proteins on stretchy substrates: the important elements of integrin-mediated rigidity sensing. Dev. Cell 19, 194–206 (2010).
Su, J., Muranjan, M. & Sap, J. Receptor protein tyrosine phosphatase alpha activates Src-family kinases and controls integrin-mediated responses in fibroblasts. Curr. Biol. 9, 505–511 (1999).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using microManager. Curr. Protoc. Mol. Biol. 92, 14.20.11–14.20.17 (2010).
Schnapp, B. J., Gelles, J. & Sheetz, M. P. Nanometer-scale measurements using video light microscopy. Cell Motil. Cytoskeleton 10, 47–53 (1988).
Savitzky, A. & Golay, M. J. E. Smoothing + differentiation of data by simplified least squares procedures. Analytical Chem. 36, 1627 (1964).
Cox, S. et al. Bayesian localization microscopy reveals nanoscale podosome dynamics. Nat. Methods 9, 195–200 (2012).
Acknowledgements
We thank all of the members of the Sheetz laboratory for their help, in particular to S. Moore and N. Biais for experimental support and helpful comments on the manuscript. We thank E. Rosten (University of Cambridge, Cambridge, UK) for the technical support during the 3B imaging analysis. We thank J. Fernandez (Columbia University) for helpful discussions about step detection. This work was financially supported by a National Institutes of Health (NIH) grant ‘Analysis of 120 nm local contractions linked to rigidity sensing’ (1 R01 GM100282-01), and by the NIH Common Fund Nanomedicine programme (PN2 EY016586). H.W. was supported by a Marie Curie International Outgoing Fellowship within the Seventh European Commission Framework Programme (PIOF-GA-2012-332045). T.I. was supported by a Postdoctoral Fellowship from the American Heart Association. M.P.S. was partially supported by the Mechanobiology Institute, National University of Singapore.
Author information
Authors and Affiliations
Contributions
H.W., G.M., S.L., P.R.-C., J.H. and M.P.S. designed the study; H.W., G.M., S.L. and T.I. performed the experiments; H.W., G.M., S.L., M.R.S., T.I. and B.O’S. were involved in data analysis; S.L. and S.G. fabricated pillar moulds; H.W. and S.L. wrote Matlab codes for analysis of pillar movements; H.W., G.M., J.H. and M.P.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Myosin is involved in CU formation.
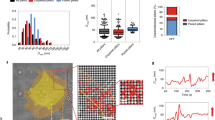
(a) CUs observed in a cell spreading on pillars coated with 10 μg ml−1 collagen (red arrows, CUs; yellow arrows, unpaired displacements). Experiment was repeated twice (5 videos altogether). (b) Additional knockdown of myosin-IIA to that shown in Fig. 1e shows ∼75% decrease in myosin-IIA expression (n = 2 separate repeats of KD) and a major decrease in CU formation. [AU: please provide and define the n number and define error bars. If the n number represents technical replicates (rather than biological replicates performed independently of each other) or n < 3 the error bars should be removed HW: n = 2, therefore the error bars were removed] Experiment was repeated twice. (c) Bleaching analysis to measure the number of active myosin heads between pillars. Brightfield (left, top) and fluorescence (left, middle) images of a cell that was fixed after 20 minutes of spreading on 0.5 μm pillars, and stained for p-MLC. Cells were imaged until all fluorescence was bleached, which enabled the detection of bleaching events of single fluorophores (left, bottom) in isolated clusters (zoom-in in middle image). In this way, the number of fluorophores (before bleaching) in single clusters was measured; the resulting histogram of the number of fluorophores per cluster is presented on the right (mean ± s.d. = 112 ± 62; n = 78 in 5 cells). Next, the number of p-MLC molecules in each cluster was calculated (see Methods). Experiment was repeated twice. (d) Tpm and p-MLC co-localize between pillars. Experiment was repeated twice.
Supplementary Figure 2 Tmod is distributed between nascent adhesions in CUs.
(a) Tmod localizes between the pillars at the cell edge, with some overlap with α-actinin. The plot on the right shows normalized average fluorescence intensities (±s.d.) of Tmod on 0.5 μm pillars measured between two adjacent pillars. n = 20 traces from 3 cells. Experiment was repeated twice. (b) mCherry-Tmod3 localizes between nascent adhesions (visualised by GFP-β3-integrin) after 15 min of spreading on fibronectin-coated glass coverslips. Experiment was repeated twice.
Supplementary Figure 3 Controls for detection of nanometer-level steps.
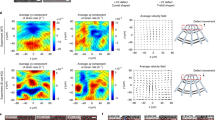
(a) Movement traces of three pillars from the same experiment in which the pillar array was displaced by a piezo-device. The similarities between the traces indicate the reliability of the tracking procedure. (b) Averaged pairwise distance analysis (±s.e.m.) from 23 records of pillars displaced by a piezo-device with 1.2 nm steps at 2 steps/s. The pairwise distance distribution function (PDF) is shown on the left; Fourier transform of this PDF peaks at a spatial frequency of 0.81 nm−1, corresponding to a 1.2 nm step size. Experiment was repeated four times. (c) Left: Pairwise distance analysis for single pillar displacement by a cell. The Fourier transform of the PDF shows a peak at 1 nm−1, corresponding to a 1 nm step size. Right: Averaged pairwise distance analysis from 38 pillar records (from 5 cells). The Fourier transform shows a peak at 0.92 nm−1, corresponding to a periodicity of 1.1 nm. The average amplitude fluctuation around a polynomial fit of the power spectrum was 0.015 ± 0.001 units (mean ± s.e.m.) [AU: please provide and define n numbers HW: n was defined above—38 pillars from 5 cells]; in contrast, the 0.92 nm−1 peak was 0.06 units above the fit. Experiment was repeated four times. (d) Example of real pillar displacement data (left), along with its corresponding negative control data (right) that was produced by adding noise to the polynomial fit of the real data (middle). [HW: there was a mistake in the legend of panel d. I replaced it with the right legend]. (e) Averaged pairwise distance analysis from 38 negative control records (from 5 cells). The 0.92 nm−1 peak observed in the real data (panel c) is absent. Experiment was repeated four times. (f) Examples of fitting negative control and real data with the step-detection algorithm (fits shown in black). In some cases (see top zoom-in), the algorithm detected steps of ∼1.2 nm with much smaller steps immediately adjacent to them. Since it was clear that these small steps were not real, they had to be merged with the adjacent large steps (original fit displayed in red). Therefore, we performed a post-fitting procedure in which a step was defined as a transition from one pillar position to the next, where the transition lasted less than 5 frames, and the second position was held for more than 10 frames. This was only performed in cases where large transition from one position to the next was very clear; whenever there was any doubt (such as in the bottom zoom-in), fitting was left as-is. This helped distinguish between ramps (top zoom-in) and steps (bottom zoom-in).
Supplementary Figure 4 Negative controls to verify that no contribution to the steps came from optical or pillar configuration effects.
(a) Left: Example of a pillar displacement with fitted steps taken from a movie of a MEF spreading on 8.4 pN nm−1 pillars whose tops were labelled with quantum-dots. Right: Step sizes detected in movements of pillars labelled with quantum-dots. Same average step size was detected as when brightfield imaging was used (P-value = 0.8, two-tailed, equal variance t-test; n = 101 steps from 5 pillars). Red line is the median value, the edges of the blue box are the 25th and 75th percentiles, the whiskers extend to the most extreme data points not considered outliers, and outliers (data greater than 3 SD than the median) are plotted individually as red dots. Experiment was repeated twice. (b) Step sizes detected when cells were plated on pillar arrays with a centre-to-centre distance between the pillars of 1 μm (twice the pillar diameter = 2D) and a hexagonal configuration (hex), 3D square configuration, and 3D hexagonal configuration. Same average step size was detected in these configurations as when 2D square configuration was used (Fig. 2 data; p-value >0.5 in all cases, two-tailed, equal variance t-tests; n = 145, 131, 159 steps from at least 8 pillars in each case) box plots definition is the same as in panel b; comparison between boxes is done by the overlap of the notches: if they do not overlap, the conclusion with 95% confidence is that the true medians do differ. Experiment was repeated twice for each case. (c) To test how the change in refractive index due to cellular coverage affects pillar displacement, the movements of the same pillar were analysed twice—once in which the starting position of the pillar was outside the cell (left), and once in which the starting position was when the cell edge had already passed the pillar (right). In both cases, the pillar movement was essentially the same. This was repeated for multiple pillars, and similar results were observed, indicating that the refractive index of the cell did not have a significant effect on the pillar displacement analysis. (d) Examples of the standard deviations (SD) [AU: please provide and define n numbers. HW: there is no n here; this is an example of single pillars—one under the cell and one not under the cell. The SD is of the fluctuations around the average positions of the pillars—we have now added the dotted lines showing the average pillar positions] of the fluctuations around the average positions (dotted lines) of a pillar not in contact with the cell and a pillar underneath the cell during a time that the latter was not being displaced. The SD is equivalent of the noise level.
Supplementary Figure 5 Analyses of steps during early stages of pillar displacements.
(a) Example of traces from two pillars that formed a CU in which simultaneous steps were observed during early pillar displacements. In red are shown steps that did not match. (b) Examples of first force production cycles before the pause on stiff and soft pillars. (c) Histograms of the times between steps before the pause on stiff and soft pillars. (n = 123, 98 holding times between steps).
Supplementary Figure 6
(a) Time traces of pillar displacement, d (solid lines), and maximum fluorescence intensity, Imax (dashed lines), of GFP-α-actinin (top) and GFP-vinculin (bottom). Imax is the maximum intensity within a distance of 0.75D (D, pillar diameter, is 0.5 μm) from the pillar centre (red circles). (b) Mean Pearson’s correlation coefficient as a function of time lag between pillar displacement traces and intensity traces of GFP-α-actinin (top, n = 26 pillars from 3 cells) or GFP-vinculin (bottom, n = 20 pillars from 3 cells). (c) Histograms of the time interval between the first peak in pillar displacement and the first peak in GFP-α-actinin intensity (top) (Δtα, n = 20 pillars from 3 cells) or GFP-vinculin intensity (bottom) (ΔtV, n = 19 pillars from 3 cells). Positive time lag in panel d indicates that protein localization precedes pillar displacement. Results in panels a–c are from an experiment that was repeated twice. (d) Recruitment of α-actinin, but not talin, correlated well with force. Time traces of the recruitment of mCherry-α-actinin and GFP-talin to a pillar upon first contact of the cell. Whereas α-actinin levels rapidly increase in correlation with force production and then rapidly decrease along with the force, talin levels gradually and steadily increase with no apparent correlation to the force. Experiment was repeated twice.
Supplementary Figure 7
(a) YFP-Tpm2.1 localizes to cell edge, but GFP-Tpm1.6 and GFP-Tpm1.7 are concentrated in central regions of cells and are absent from the lamellipodium. Experiment was repeated twice. (b) Western blots and quantifications showing knockdown of Tpm2.1 expression by siRNA and shRNA (two separate target sequences, see Methods) to ∼30% and ∼25% of the original levels (n = 3 separate repeats of KD in each case). Unprocessed original scan of the siRNA blot is shown in Supplementary Fig. 9a. (c) Edge localization is lost when staining Tpm2.1-KD cells with the antibody that detects HMW tropomyosins. Experiment was repeated twice. (d) WT cells and cells transfected with NT-SiRNA are polarized and elongated after 1 h of spreading on 8.4 pN nm−1 pillars, whereas Tpm2.1-KD cells are small and round. The polarization of the cells was measured using circularity measurements in ImageJ; a circularity value of 1 indicates a perfect circle and as the value approaches 0 it indicates an increasingly elongated polygon. Experiment was repeated twice. (e) Force vectors by Tpm2.1-KD cells on pillars show completely disorganized force fields, with no contractile units. (f) Another example to that given in Fig. 7c showing high displacement velocity by a Tpm2.1-DK cell. (g) Higher step sizes produced by Tpm2.1-KD cells. Left: Pairwise distance analysis for Tpm2.1-KD cells (average PDF taken from 25 pillar records from 4 cells). Middle: Fourier transform of PDF showing a peak at 0.39 ± 0.1 nm−1 (corresponding to a periodicity of 2.5 ± 0.3 nm). Right: Boxplot of step sizes produced by Tpm2.1-KD cells (KD by shRNA). Red line is the median value, the edges of the blue box are the 25th and 75th percentiles, the whiskers extend to the most extreme data points not considered outliers, and outliers (data greater than 3 SD than the median) are plotted individually as red dots. n = 155 steps from 7 pillars). Experiment was repeated twice. (h) Knockdown of Tpm1.7 by two different shRNAs (∼80% and ∼65% KD for sh1 and sh2, respectively; n = 3 separate repeats of KD) does not alter the step size during pillar displacements. Unprocessed original scan of the blot is shown in Supplementary Fig. 9b. Box plots definition is the same as in panel g; comparison between boxes is done by the overlap of the notches: if they do not overlap, the conclusion with 95% confidence is that the true medians do differ. n = 120, 133 steps from 6 and 7 pillars. Experiment was repeated twice in each case. (i) CUs (red arrows) are observed in MCF-10A cells, but rarely in MDA-MB-231 cells. (j) Western blot showing ∼75% knockdown of Tpm2.1 expression in MCF-10A cells (n = 3 separate repeats of KD).
Supplementary Figure 8
(a) Model for tropomyosin regulation of rigidity sensing. In this illustration, actin fibres (green) that are polymerized from nascent adhesions on adjacent pillars are locally contracted by myosin-IIA mini-filaments (blue). In WT cells (left), tropomyosin (orange fibres) regulates myosin stepping by restricting the actin monomers which it can attach to1. After a certain number of steps that leads to load of ∼20 pN, there is a pause in the displacement due to high myosin-actin affinity under high loads. This leads to further recruitment of myosin molecules, as well as to reinforcement of the adhesion, possibly due to recruitment of adhesion proteins to binding sites that were exposed by the applied forces2. This then promotes further pulling up to a maximal distance of ∼50 nm. In Tpm2.1-depleted cells (right), myosin can perform larger steps since Tpm restriction is removed. This leads to higher force production at faster rates, eventually resulting in adhesion breakage. Another feature that might contribute to force regulation is the extensibility of the actin filaments3, which increases by ∼50% in the absence of Tpm (ref. 4). This could affect the locations on actin to which myosin can bind5, and therefore the forces produced by each myosin cross-bridge. Whether or not this attribute is (part of) the reason for the higher forces and movement velocities that we observe in the absence of tropomyosin remains to be determined. (b) Force-velocity curve constructed by measuring the displacement velocities of pillars with different rigidities at a displacement level of 15–25 nm.
Supplementary Figure 9
Unprocessed original scans of the blots showing KD of Tpm2.1 and Tpm1.7.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1282 kb)
Live imaging at 1 frame/s of a MEF spreading on 0.5 μm pillars.
The movie begins right after P0 stage (initial attachment). Note the rapid increase in cell area up to ∼3.5 min (P1), after which slower spreading is observed with protrusion/retraction cycles of the cell edge (P2). (AVI 6444 kb)
Live imaging at 100 frames s−1 of a MEF spreading on 0.5 μm pillars; pillars that are part of a CU are marked.
Note that other, more short-lived, CUs also appear throughout the movie. (AVI 17669 kb)
MEF spreading on ultra-soft (0.85 pN nm−1 stiffness) pillars.
Note the frequent breakage of adhesions and return of pillars to zero force position. (AVI 7017 kb)
YFP-Tpm2.1-expressing MEF spreading on fibronectin-coated glass.
Note the gradual accumulation of YFP-Tpm2.1 at the edge, followed by membrane retraction. (AVI 1258 kb)
Rights and permissions
About this article
Cite this article
Wolfenson, H., Meacci, G., Liu, S. et al. Tropomyosin controls sarcomere-like contractions for rigidity sensing and suppressing growth on soft matrices. Nat Cell Biol 18, 33–42 (2016). https://doi.org/10.1038/ncb3277
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3277
This article is cited by
-
Biophysical control of plasticity and patterning in regeneration and cancer
Cellular and Molecular Life Sciences (2024)
-
TPM2 attenuates progression of prostate cancer by blocking PDLIM7-mediated nuclear translocation of YAP1
Cell & Bioscience (2023)
-
The biophysical property of the limbal niche maintains stemness through YAP
Cell Death & Differentiation (2023)
-
The glycocalyx affects the mechanotransductive perception of the topographical microenvironment
Journal of Nanobiotechnology (2022)
-
Correlative cryo-ET identifies actin/tropomyosin filaments that mediate cell–substrate adhesion in cancer cells and mechanosensitivity of cell proliferation
Nature Materials (2022)