Abstract
During mouse development, core planar cell polarity (PCP) proteins become polarized in the epidermal plane to guide angling/morphogenesis of hair follicles. How PCP is established is poorly understood. Here, we identify a key role for Wdr1 (also known as Aip1), an F-actin-binding protein that enhances cofilin/destrin-mediated F-actin disassembly. We show that cofilin and destrin function redundantly in developing epidermis, but their combined depletion perturbs cell adhesion, cytokinesis, apicobasal polarity and PCP. Although Wdr1 depletion accentuates single-loss-of-cofilin/destrin phenotypes, alone it resembles core PCP mutations. Seeking a mechanism, we find that Wdr1 and cofilin/destrin-mediated actomyosin remodelling are essential for generating or maintaining cortical tension within the developing epidermal sheet and driving the cell shape and planar orientation changes that accompany establishment of PCP in mammalian epidermis. Our findings suggest intriguing evolutionary parallels but mechanistic modifications to the distal wing hinge-mediated mechanical forces that drive cell shape change and orient PCP in the Drosophila wing disc.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Eaton, S. & Jülicher, F. Cell flow and tissue polarity patterns. Curr. Opin. Genet. Dev. 21, 747–752 (2011).
Goodrich, L. V. & Strutt, D. Principles of planar polarity in animal development. Development 138, 1877–1892 (2011).
McNeill, H. Planar cell polarity: keeping hairs straight is not so simple. Cold Spring Harb. Perspect. Biol. 2, a003376 (2010).
Devenport, D. & Fuchs, E. Planar polarization in embryonic epidermis orchestrates global asymmetric morphogenesis of hair follicles. Nat. Cell Biol. 10, 1257–1268 (2008).
Guo, N., Hawkins, C. & Nathans, J. Frizzled6 controls hair patterning in mice. Proc. Natl Acad. Sci. USA 101, 9277–9281 (2004).
Aigouy, B. et al. Cell flow reorients the axis of planar polarity in the wing epithelium of Drosophila. Cell 142, 773–786 (2010).
Matis, M. & Axelrod, J. D. Regulation of PCP by the Fat signaling pathway. Genes Dev. 27, 2207–2220 (2013).
Luxenburg, C., Pasolli, H. A., Williams, S. E. & Fuchs, E. Developmental roles for Srf, cortical cytoskeleton and cell shape in epidermal spindle orientation. Nat. Cell Biol. 13, 203–214 (2011).
Fujibuchi, T. et al. AIP1/WDR1 supports mitotic cell rounding. Biochem. Biophys. Res. Commun. 327, 268–275 (2005).
Chen, Q., Courtemanche, N. & Pollard, T. D. Aip1 promotes actin filament severing by cofilin and regulates the constriction of the cytokinetic contractile ring. J. Biol. Chem. 290, 2289–2300 (2014).
Aizawa, H. et al. Hyperosmotic stress-induced reorganization of actin bundles in Dictyostelium cells over-expressing cofilin. Genes Cells 4, 311–324 (1999).
Augustine, R. C., Pattavina, K. A., Tuzel, E., Vidali, L. & Bezanilla, M. Actin interacting protein1 and actin depolymerizing factor drive rapid actin dynamics in Physcomitrella patens. Plant Cell 23, 3696–3710 (2011).
Chu, D. et al. AIP1 acts with cofilin to control actin dynamics during epithelial morphogenesis. Development 139, 3561–3571 (2012).
Lin, M-C., Galletta, B. J., Sept, D. & Cooper, J. A. Overlapping and distinct functions for cofilin, coronin and Aip1 in actin dynamics in vivo. J. Cell Sci. 123, 1329–1342 (2010).
Mohri, K. & Ono, S. Actin filament disassembling activity of Caenorhabditis elegans actin-interacting protein 1 (UNC-78) is dependent on filament binding by a specific ADF/cofilin isoform. J. Cell Sci. 116, 4107–4118 (2003).
Okada, K., Obinata, T. & Abe, H. XAIP1: a Xenopus homologue of yeast actin interacting protein 1 (AIP1), which induces disassembly of actin filaments cooperatively with ADF/cofilin family proteins. J. Cell Sci. 112 (Pt 10), 1553–1565 (1999).
Rodal, A. A., Tetreault, J. W., Lappalainen, P., Drubin, D. G. & Amberg, D. C. Aip1p interacts with cofilin to disassemble actin filaments. J. Cell Biol. 145, 1251–1264 (1999).
Kile, B. T. et al. Mutations in the cofilin partner Aip1/Wdr1 cause autoinflammatory disease and macrothrombocytopenia. Blood 110, 2371–2380 (2007).
Beronja, S., Livshits, G., Williams, S. & Fuchs, E. Rapid functional dissection of genetic networks via tissue-specific transduction and RNAi in mouse embryos. Nat. Med. 16, 821–827 (2010).
Curtin, J. A. et al. Mutation of celsr1 disrupts planar polarity of inner ear hair cells and causes severe neural tube defects in the mouse. Curr. Biol. 13, 1129–1133 (2003).
Kibar, Z. et al. Ltap, a mammalian homolog of Drosophila Strabismus/Van Gogh, is altered in the mouse neural tube mutant Loop-tail. Nat. Genet. 28, 251–255 (2001).
Chen, X. & Macara, I. G. Par-3 mediates the inhibition of LIM kinase 2 to regulate cofilin phosphorylation and tight junction assembly. J. Cell Biol. 172, 671–678 (2006).
Michelot, A. et al. Actin filament elongation in Arp2/3-derived networks is controlled by three distinct mechanisms. Dev. Cell 24, 182–195 (2013).
Nadkarni, A. V. & Brieher, W. M. Aip1 destabilizes cofilin-saturated actin filaments by severing and accelerating monomer dissociation from ends. Curr. Biol. 24, 2749–2757 (2014).
Ikeda, S. et al. Aberrant actin cytoskeleton leads to accelerated proliferation of corneal epithelial cells in mice deficient for destrin (actin depolymerizing factor). Hum. Mol. Genet. 12, 1029–1037 (2003).
Smith, R. S. et al. Corn1: a mouse model for corneal surface disease and neovascularization. Invest. Ophthalmol. Vis. Sci. 37, 397–404 (1996).
Agnew, B. J., Minamide, L. S. & Bamburg, J. R. Reactivation of phosphorylated actin depolymerizing factor and identification of the regulatory site. J. Biol. Chem. 270, 17582–17587 (1995).
Nagaoka, R., Abe, H. & Obinata, T. Site-directed mutagenesis of the phosphorylation site of cofilin: its role in cofilin-actin interaction and cytoplasmic localization. Cell. Motil. Cytoskeleton 35, 200–209 (1996).
Kiuchi, T., Ohashi, K., Kurita, S. & Mizuno, K. Cofilin promotes stimulus-induced lamellipodium formation by generating an abundant supply of actin monomers. J. Cell Biol. 177, 465–476 (2007).
Mahaffey, J. P., Grego-Bessa, J., Liem, K. F. & Anderson, K. V. Cofilin and Vangl2 cooperate in the initiation of planar cell polarity in the mouse embryo. Development 140, 1262–1271 (2013).
Devenport, D., Oristian, D., Heller, E. & Fuchs, E. Mitotic internalization of planar cell polarity proteins preserves tissue polarity. Nat. Cell Biol. 13, 893–902 (2011).
Ren, N., Charlton, J. & Adler, P. N. The flare gene, which encodes the AIP1 protein of Drosophila, functions to regulate F-actin disassembly in pupal epidermal cells. Genetics 176, 2223–2234 (2007).
Shimada, Y., Yonemura, S., Ohkura, H., Strutt, D. & Uemura, T. Polarized transport of Frizzled along the planar microtubule arrays in Drosophila wing epithelium. Dev. Cell 10, 209–222 (2006).
Ross, A. J. et al. Disruption of Bardet-Biedl syndrome ciliary proteins perturbs planar cell polarity in vertebrates. Nat. Genet. 37, 1135–1140 (2005).
Reymann, A. C. et al. Actin network architecture can determine myosin motor activity. Science 336, 1310–1314 (2012).
Ono, K. & Ono, S. Two actin-interacting protein 1 isoforms function redundantly in the somatic gonad and are essential for reproduction in Caenorhabditis elegans. Cytoskeleton (Hoboken) 71, 36–45 (2014).
Yuan, B. et al. A cardiomyocyte-specific Wdr1 knockout demonstrates essential functional roles for actin disassembly during myocardial growth and maintenance in mice. Am. J. Pathol. 184, 1967–1980 (2014).
Rhee, J. M. et al. In vivo imaging and differential localization of lipid-modified GFP-variant fusions in embryonic stem cells and mice. Genesis 44, 202–218 (2006).
Hutson, M. S. et al. Forces for morphogenesis investigated with laser microsurgery and quantitative modeling. Science 300, 145–149 (2003).
Farhadifar, R., Röper, J-C., Aigouy, B., Eaton, S. & Jülicher, F. The influence of cell mechanics, cell–cell interactions, and proliferation on epithelial packing. Curr. Biol. 17, 2095–2104 (2007).
Fernandez-Gonzalez, R., Simoes, S. D. E. M., Röper, J. C., Eaton, S. & Zallen, J. A. Myosin II dynamics are regulated by tension in intercalating cells. Dev. Cell 17, 736–743 (2009).
Straight, A. F. et al. Dissecting temporal and spatial control of cytokinesis with a myosin II inhibitor. Science 299, 1743–1747 (2003).
Ishizaki, T. et al. Pharmacological properties of Y-27632, a specific inhibitor of rho-associated kinases. Mol. Pharmacol. 57, 976–983 (2000).
Bubb, M. R., Senderowicz, A. M., Sausville, E. A., Duncan, K. L. & Korn, E. D. Jasplakinolide, a cytotoxic natural product, induces actin polymerization and competitively inhibits the binding of phalloidin to F-actin. J. Biol. Chem. 269, 14869–14871 (1994).
Heisenberg, C. P. & Bellaïche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013).
Guillot, C. & Lecuit, T. Mechanics of epithelial tissue homeostasis and morphogenesis. Science 340, 1185–1189 (2013).
Arber, S. et al. Regulation of actin dynamics through phosphorylation of cofilin by LIM-kinase. Nature 393, 805–809 (1998).
Yang, N. et al. Cofilin phosphorylation by LIM-kinase 1 and its role in Rac-mediated actin reorganization. Nature 393, 809–812 (1998).
Moriyama, K. & Yahara, I. The actin-severing activity of cofilin is exerted by the interplay of three distinct sites on cofilin and essential for cell viability. Biochem. J. 365, 147–155 (2002).
Wiggan, O., Shaw, A. E., DeLuca, J. G. & Bamburg, J. R. ADF/cofilin regulates actomyosin assembly through competitive inhibition of myosin II binding to F-actin. Dev. Cell 22, 530–543 (2012).
Acknowledgements
We thank D. Devenport, S. Williams, S. Beronja, A. R. Folgueras, D. Schramek, I. Matos and E. Ezratty for intellectual input; D. Oristian and A. Aldeguer as mouse specialists; Comparative Bioscience Center (AAALAC accredited) for care of mice in accordance with National Institutes of Health (NIH) guidelines; Bioimaging Center (A. North, director) for advice; Flow Cytometry facility (S. Mazel, director) for FACS sorting. Cfl–GFP was a generous gift from J. Condeelis (Albert Einstein college of Medicine, New York, USA); E.F. is an Investigator of the Howard Hughes Medical Institute. This research was supported by a grant from the NIH (R37-AR27883, E.F.), a Starr Stem Cell Postdoctoral Fellowship (C.L.) and a Genetics Training Grant by the NIH (E.H.).
Author information
Authors and Affiliations
Contributions
C.L. and E.F. conceived the study. C.L., E.H. and E.F. designed the experiments. C.L. and E.H. carried out the experiments and analysed the data. H.A.P. performed the ultrastructural analyses (Supplementary Fig. 1S). S.C. made the Wdr1-rescue construct, C.L. and N.S. performed the in utero injections. C.L. and M.N. prepared high-titre viruses. C.L., E.H. and E.F. wrote the paper. All authors provided intellectual input, vetted and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 2 Wdr1 knockdown epidermis display normal intercellular adhesion and attachment to the basement membrane.
Correlative light and electron microscopy of E16.5 headskin in WT and Wdr1-368. Whole mount skin was immunolabeled for GFP (and subsequently embedded for electron microscopy. (a) and (b) Semithin sections (1 μm) were stained with toluidine blue and examined for the presence of GFP positive cells (brown nuclei). Dotted line indicated dermo-epidermal boundary. (b and b′) consecutive ultrathin sections were examined at the electron microscope. For Wdr1-368, the same exact cells that were GFP+ by immunohistochemistry were identified (pseudocolored in green) and photographed at high resolution. Red box enlarged in (c) and yellow box enlarged in (e). (c) Epidermal cells in Wdr1-368 display normal adhesion, with intact desmosomes (de) shown in boxed area enlarged in (d) No gaps were observed between the cells. Some inevitable cytoplasmic damage (asterisks) is due to detergent permeabilization necessary for immunolabelling. (e) Electron microscopic analysis showed intact basal lamina (bl) and hemidesmosomes (hd). Bars in (a) and (c), 2 μm, also valid for a′, b and b′. Bars in (d) and (e), 500 nm.
Supplementary Figure 3 Normal adhesion and apicobasal polarity in Wdr1 knockdown epidermis.
10 μm sagittal sections of E16.5 backskins were labelled for: (a) E-cadherin (E-cad) or ZO1 and co-labelled for integrin β4 (β4) and Laminin 5 (Lam5) (b) ABP markers Par3 and pericentrin. DAPI was used to label chromatin. Dotted line denotes the dermal-epidermal border. Inserts denote infected cells (H2B-GFP+ nuclei). Scale bar, 10 μm.
Supplementary Figure 4 Wdr1 knockdown by a second hairpin (Wdr1-1622) results in PCP defects analogous to Wdr1-368 and validation of Vangl2 shRNAs.
(a) 10 μm sagittal sections of E15.5 backskins were labelled for Par3 or Pericentrin (PC) or co-labelled for Keratin 5 (K5) and Keratin 10 (K10). (b) Whole-mount immunofluorescence of E15.5 backskins co-labelled for E-cadherin (E-cad) and Celsr1 and imaged in the mid-plane of the basal layer. (c) Whole-mount immunofluorescence of E18.5 HFs labelled for E-cadherin. (d) Quantifications of data shown in (c), n = 36 hair follicles from 3 embryos. Note similarities to data compiled on similarly aged Wdr1-368 knockdown embryonic epidermis (see Figs 1 and 2). Green arrows denote normal HF orientation, red arrows denote abnormal HF orientation. Red circles denote perpendicularly oriented HFs. Insets denote transduced cells (H2B-RFP+ nuclei). (e) In vitro Knockdown efficiencies of shRNAs against Vangl2 to achieve specific loss of PCP in the skin, n = 3 independent measurements/hairpin. Error bars indicate mean ± s.e.m. Scale bars, 10 μm.
Supplementary Figure 5 Generation and expression of a hairpin-resistant Wdr1-GFP lentiviral transgene.
(a) Wdr1 coding sequence recognized by Wdr1-368 hairpin and a mutant, shown in red, that renders it refactory to the hairpin. (b,c) Western blots of protein extracts from WT cells or cells transduced with lentiviruses that contain either Wdr1-368 shRNA, Scrambled control shRNA or the Wdr1-GFP hairpin-resistant mutant cDNA expression vector. Blots were probed with WDR1 antibodies (Ab) (b) or GFP Ab (c). Note the ∼93 kDa WDR1-GFP protein (endogenous WDR1 = 66 kDa, GFP = 27 kDa) detected with both Abs. Glyceraldehyde dehydrogenase (GAPDH) and hypoxanthine-guanine phosphoribosyltransferase (HPRT) Abs were used as loading controls.
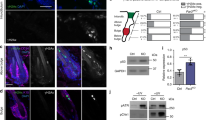
Supplementary Figure 6 Normal localization of Cofilin and Destrin in Wdr1-368 KD epidermis.
(a) 10 μm sagittal sections of E15.5 control and WDR1-368 KD backskins were labelled for Cofilin (Cfl) or Destrin (Dstn). Inserts denote transduced cells (H2B-GFP+ nuclei). DAPI in blue. (c) Western blot analyses of protein extracts from control (Scr) and Wdr1-368 KD 10MK were probed with antibodies against: WDR1, Cofilin (Cfl) phospho-serine 3-cofilin (p-Cfl) and α-tubulin (αTub, loading control). (d) qPCR of mRNAs were from control and Cfl1KD/Dstn-null (KD/KO) 10MK. n = 3 samples/condition Cfl1 levels in Ctrl versus KD/KO p = 0.0026; Dstn levels in Ctrl. versus KD/KO p = 0.009. Dotted line denotes the dermal-epidermal border. Insets show transduced cells (H2B-GFP+ nuclei). Scale bars, 10 μm.
Supplementary Figure 7 Quantification of F-actin content, actin severing, and endogeneous levels of cofilin in Wdr1-KD and cofilin-overexpressing keratinocytes.
(a) Uncropped Western blots used to evaluate Wdr1 knockdown and endogeneous levels of actin in Fig. 1a. (b) Uncropped Western blots used to evalulate endogeneous cofilin and phosph-cofilin levels in transduced keratinocytes in Fig. 5a, b. Immunoblot for GFP indicates overexpression of histone 2B (H2B)-GFP (∼51 kDa), used as a marker of lentiviral transduction in scramble- and Wdr1-KD keratinocytes, and overexpression of GFP-CflS3A (∼52 kDa) in the others. Relative levels were quantified by fluorescence intensity on a LiCor Odyssey CLx imager and normalized to GAPDH levels. (c) Uncropped Western blots used to quantify F/G actin ratios. G-actin standards were loaded at the indicated quantities and used to construct a calibration curve. Actin levels were normalized to the cytoplasmic HPRT signal in each lysate.
Supplementary Figure 8 Defects in mitotic rounding in Wdr1-368 KD epidermis.
(a) Whole-mount immunofluorescence of E15.5 backskin labelled for Ecadherin (E-cad) and imaged at the basal layer plane. (b) Quantifications of data from (a). Control: 1.14 ± 0.088; Wdr1 KD: 1.39 ± 0.27, p = 5.27 × 10−5 (unpaired t-test), n = 23 cells (WT); n = 29 cells (Wdr1 KD) from 3 embryos per condition. Box and whiskers plot: line = median, box = 50% range, whiskers = 100% range. Arrow denotes early mitotic cells. Inserts denote infected cells (H2B-GFP+ nuclei). Scale bar, 10 μm.
Supplementary information
Supplementary Information
Supplementary Information (PDF 921 kb)
Rights and permissions
About this article
Cite this article
Luxenburg, C., Heller, E., Pasolli, H. et al. Wdr1-mediated cell shape dynamics and cortical tension are essential for epidermal planar cell polarity. Nat Cell Biol 17, 592–604 (2015). https://doi.org/10.1038/ncb3146
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3146
This article is cited by
-
Wnt4 and ephrinB2 instruct apical constriction via Dishevelled and non-canonical signaling
Nature Communications (2023)
-
Anillin governs mitotic rounding during early epidermal development
BMC Biology (2022)
-
Symmetry breaking of tissue mechanics in wound induced hair follicle regeneration of laboratory and spiny mice
Nature Communications (2021)
-
WD repeat-containing protein 1 maintains β-Catenin activity to promote pancreatic cancer aggressiveness
British Journal of Cancer (2020)
-
Mechanics of a multilayer epithelium instruct tumour architecture and function
Nature (2020)