Abstract
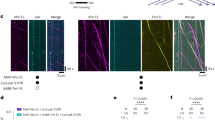
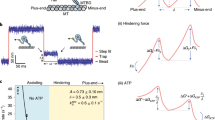
Subcellular localization of mRNAs by cytoskeletal motors plays critical roles in the spatial control of protein function1. However, optical limitations of studying mRNA transport in vivo mean that there is little mechanistic insight into how transcripts are packaged and linked to motors, and how the movement of mRNA–motor complexes on the cytoskeleton is orchestrated. Here, we have reconstituted transport of mRNPs containing specific RNAs in vitro. We show directly that mRNAs that are either apically localized or non-localized in Drosophila melanogaster embryos associate with the dynein motor and move bidirectionally on individual microtubules, with localizing mRNPs exhibiting a strong minus-end-directed bias. Single-molecule fluorescence measurements reveal that RNA localization signals increase the average number of dynein and dynactin components recruited to individual mRNPs. We find that, surprisingly, individual RNA molecules are present in motile mRNPs in vitro and provide evidence that this is also the case in vivo. Thus, RNA oligomerization is not obligatory for transport. Our findings lead to a model in which RNA localization signals produce highly polarized distributions of transcript populations through modest changes in motor copy number on single mRNA molecules.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Holt, C. E. & Bullock, S. L. Subcellular mRNA localization in animal cells and why it matters. Science 326, 1212–1216 (2009).
Wilkie, G. S. & Davis, I. Drosophila wingless and pair-rule transcripts localize apically by dynein-mediated transport of RNA particles. Cell 105, 209–219 (2001).
Welte, M. A., Gross, S. P., Postner, M., Block, S. M. & Wieschaus, E. F. Developmental regulation of vesicle transport in Drosophila embryos: forces and kinetics. Cell 92, 547–557 (1998).
Dienstbier, M., Boehl, F., Li, X. & Bullock, S. L. Egalitarian is a selective RNA-binding protein linking mRNA localization signals to the dynein motor. Genes Dev. 23, 1546–1558 (2009).
Navarro, C., Puthalakath, H., Adams, J. M., Strasser, A. & Lehmann, R. Egalitarian binds dynein light chain to establish oocyte polarity and maintain oocyte fate. Nat. Cell Biol. 6, 427–435 (2004).
Bullock, S. L. & Ish-Horowicz, D. Conserved signals and machinery for RNA transport in Drosophila oogenesis and embryogenesis. Nature 414, 611–616 (2001).
Hoogenraad, C. C. et al. Mammalian Golgi-associated Bicaudal-D2 functions in the dynein–dynactin pathway by interacting with these complexes. EMBO J. 20, 4041–4054 (2001).
Bullock, S. L., Nicol, A., Gross, S. P. & Zicha, D. Guidance of bidirectional motor complexes by mRNA cargoes through control of dynein number and activity. Curr. Biol. 16, 1447–1452 (2006).
Vendra, G., Hamilton, R. S. & Davis, I. Dynactin suppresses the retrograde movement of apically localized mRNA in Drosophila blastoderm embryos. RNA 13, 1860–1867 (2007).
Serano, T. L. & Cohen, R. S. A small predicted stem-loop structure mediates oocyte localization of Drosophila K10 mRNA. Development 121, 3809–3818 (1995).
Bullock, S. L., Ringel, I., Ish-Horowicz, D. & Lukavsky, P. J. A’-form RNA helices are required for cytoplasmic mRNA transport in Drosophila. Nat. Struct. Mol. Biol. 17, 703–709 (2010).
Ingham, P. W., Howard, K. R. & Ish-Horowicz, D. Transcriptional pattern of the Drosophila segmentation gene hairy. Nature 318, 439–445 (1985).
Fusco, D. et al. Single mRNA molecules demonstrate probabilistic movement in living mammalian cells. Curr. Biol. 13, 161–167 (2003).
Larsen, K. S., Xu, J., Cermelli, S., Shu, Z. & Gross, S. P. BicaudalD actively regulates microtubule motor activity in lipid droplet transport. PLoS One 3, e3763 (2008).
Shubeita, G. T. et al. Consequences of motor copy number on the intracellular transport of kinesin-1-driven lipid droplets. Cell 135, 1098–1107 (2008).
Vale, R. D. et al. Direct observation of single kinesin molecules moving along microtubules. Nature 380, 451–453 (1996).
Ross, J. L., Wallace, K., Shuman, H., Goldman, Y. E. & Holzbaur, E. L. Processive bidirectional motion of dynein–dynactin complexes in vitro. Nat. Cell Biol. 8, 562–570 (2006).
Shu, D., Zhang, H., Jin, J. & Guo, P. Counting of six pRNAs of phi29 DNA-packaging motor with customized single-molecule dual-view system. EMBO J. 26, 527–537 (2007).
Kardon, J. R., Reck-Peterson, S. L. & Vale, R. D. Regulation of the processivity and intracellular localization of Saccharomyces cerevisiae dynein by dynactin. Proc. Natl Acad. Sci. USA 106, 5669–5674 (2009).
Schuster, M., Lipowsky, R., Assmann, M. A., Lenz, P. & Steinberg, G. Transient binding of dynein controls bidirectional long-range motility of early endosomes. Proc. Natl. Acad. Sci. USA 108, 3618–3623 (2011).
Coy, D. L., Hancock, W. O., Wagenbach, M. & Howard, J. Kinesin’s tail domain is an inhibitory regulator of the motor domain. Nat. Cell Biol. 1, 288–292 (1999).
Mallik, R., Petrov, D., Lex, S. A., King, S. J. & Gross, S. P. Building complexity: an in vitro study of cytoplasmic Dynein with in vivo implications. Curr. Biol. 15, 2075–2085 (2005).
Kiebler, M. A. & Bassell, G. J. Neuronal RNA granules: movers and makers. Neuron 51, 685–690 (2006).
Mhlanga, M. M. et al. In vivo colocalisation of oskar mRNA and trans-acting proteins revealed by quantitative imaging of the Drosophila oocyte. PLoS One 4, e6241 (2009).
Delanoue, R., Herpers, B., Soetaert, J., Davis, I. & Rabouille, C. Drosophila Squid/hnRNP helps Dynein switch from a gurken mRNA transport motor to an ultrastructural static anchor in sponge bodies. Dev. Cell. 13, 523–538 (2007).
Lange, S. et al. Simultaneous transport of different localized mRNA species revealed by live-cell imaging. Traffic 9, 1256–1267 (2008).
Ferrandon, D., Koch, I., Westhof, E. & Nusslein-Volhard, C. RNA–RNA interaction is required for the formation of specific bicoid mRNA 3′ UTR-STAUFEN ribonucleoprotein particles. EMBO J. 16, 1751–1758 (1997).
Paré, A. et al. Visualization of individual Scr mRNAs during Drosophila embryogenesis yields evidence for transcriptional bursting. Curr. Biol. 19, 2037–2042 (2009).
Wilkie, G. S., Shermoen, A. W., O’Farrell, P. H. & Davis, I. Transcribed genes are localized according to chromosomal position within polarized Drosophila embryonic nuclei. Curr. Biol. 9, 1263–1266 (1999).
Nasiadka, A., Dietrich, B. H. & Krause, H. M. Anterior–posterior patterning in the Drosophila embryo. Adv. Dev. Biol. Biochem. 12, 155–204 (2002).
Mikl, M., Vendra, G. & Kiebler, M. A. Independent localization of MAP2, CaMKIIα and β-actin RNAs in low copy numbers. EMBO Rep. 12, 1077–1084 (2011).
Zimyanin, V. L. et al. In vivo imaging of oskar mRNA transport reveals the mechanism of posterior localization. Cell 134, 843–853 (2008).
Kerssemakers, J. W. et al. Assembly dynamics of microtubules at molecular resolution. Nature 442, 709–712 (2006).
Loiseau, P., Davies, T., Williams, L. S., Mishima, M. & Palacios, I. M. Drosophila PAT1 is required for Kinesin-1 to transport cargo and to maximize its motility. Development 137, 2763–2772 (2010).
Pandey, R., Heeger, S. & Lehner, C. F. Rapid effects of acute anoxia on spindle kinetochore interactions activate the mitotic spindle checkpoint. J. Cell Sci. 120, 2807–2818 (2007).
Januschke, J. et al. Polar transport in the Drosophila oocyte requires Dynein and Kinesin I cooperation. Curr. Biol. 12, 1971–1981 (2002).
Cole, K., Truong, V., Barone, D. & McGall, G. Direct labeling of RNA with multiple biotins allows sensitive expression profiling of acute leukemia class predictor genes. Nucleic Acids Res. 32, e86 (2004).
Ross, J. L., Shuman, H., Holzbaur, E. L. & Goldman, Y. E. Kinesin and dynein–dynactin at intersecting microtubules: motor density affects dynein function. Biophys. J. 94, 3115–3125 (2008).
Duncan, J. E. & Warrior, R. The cytoplasmic dynein and kinesin motors have interdependent roles in patterning the Drosophila oocyte. Curr. Biol. 12, 1982–1991 (2002).
Mische, S. et al. Dynein light intermediate chain: an essential subunit that contributes to spindle checkpoint inactivation. Mol. Biol. Cell 19, 4918–4929 (2008).
Mach, J. M. & Lehmann, R. An Egalitarian-BicaudalD complex is essential for oocyte specification and axis determination in Drosophila. Genes Dev. 11, 423–435 (1997).
Suter, B. & Steward, R. Requirement for phosphorylation and localizationof the Bicaudal-D protein in Drosophila oocyte differentiation. Cell 67, 917–926 (1991).
Sharp, D. J. et al. Functional coordination of three mitotic motors in Drosophila embryos. Mol. Biol. Cell 11, 241–253 (2000).
Lecuyer, E., Parthasarathy, N. & Krause, H. M. Fluorescent in situ hybridization protocols in Drosophila embryos and tissues. Methods Mol. Biol. 420, 289–302 (2008).
Roth, P. et al. The Drosophila nucleoporin DNup88 localizes DNup214 and CRM1 on the nuclear envelope and attenuates NES-mediated nuclear export. J. Cell Biol. 163, 701–706 (2003).
Acknowledgements
This work was financially supported by the UK Medical Research Council (project U105178790) and a Lister Institute prize fellowship (to S.L.B.). We are very grateful to A. Carter, A. Guichet, T. Hays, R. Lehmann, E. Miska, C. Navarro, I. Palacios, J. Raff, C. Samakovlis, B. Suter and R. Warrior for providing reagents, A. Paré for advice on in situ hybridization, N. Barry for advice on microscopy, A. Nicol and D. Zicha for help with mRNP tracking software, M. Dogterom for the Stepfinder algorithm, and members of the Bullock laboratory, D. St Johnston and A. Carter for their valuable insights.
Author information
Authors and Affiliations
Contributions
M.A-N. and S.L.B. conceived the project. M.A-N. developed the in vitro assays, carried out all in vitro experiments and analysed the data. S.L.B. carried out the embryo injections and in situ hybridizations and analysed the data. M.A-N. and S.L.B. interpreted the data. S.L.B. and M.A-N. wrote and edited the manuscript, respectively.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 585 kb)
Supplementary Movie 1
Supplementary Information (AVI 450 kb)
Supplementary Movie 2
Supplementary Information (AVI 357 kb)
Supplementary Movie 3
Supplementary Information (AVI 158 kb)
Supplementary Movie 4
Supplementary Information (AVI 166 kb)
Rights and permissions
About this article
Cite this article
Amrute-Nayak, M., Bullock, S. Single-molecule assays reveal that RNA localization signals regulate dynein–dynactin copy number on individual transcript cargoes. Nat Cell Biol 14, 416–423 (2012). https://doi.org/10.1038/ncb2446
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2446
This article is cited by
-
In the right place at the right time: visualizing and understanding mRNA localization
Nature Reviews Molecular Cell Biology (2015)
-
Independent and coordinate trafficking of single Drosophila germ plasm mRNAs
Nature Cell Biology (2015)
-
Nervous translation, do you get the message? A review of mRNPs, mRNA–protein interactions and translational control within cells of the nervous system
Cellular and Molecular Life Sciences (2014)
-
Single-molecule reconstitution of mRNA transport by a class V myosin
Nature Structural & Molecular Biology (2013)
-
A numbers game underpins cytoplasmic mRNA transport
Nature Cell Biology (2012)