Abstract
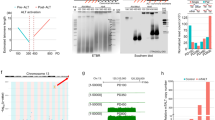
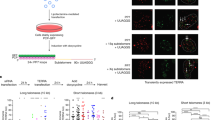
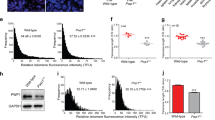
Mammalian telomeres consist of non-coding TTAGGG repeats that are bound by the multi-protein complex 'shelterin', thus protecting chromosome ends from DNA repair mechanisms and degradation1. Mammalian telomeric chromatin is enriched for the constitutive heterochromatin marks H3K9me3, H4K20me3 and HP1 (refs 2–7). Similar to pericentric heterochromatin, telomeric heterochromatin is thought to be fundamental for the maintenance of chromosomal integrity8. Here, we report that telomeric repeats are transcribed by DNA-dependent RNA polymerase II, which, in turn, interacts with the TRF1 shelterin protein. Telomeric RNAs (TelRNAs) contain UUAGGG repeats, are polyadenylated and are transcribed from the telomeric C-rich strand. Transcription of mammalian telomeres is regulated by several mechanisms, including developmental status, telomere length, cellular stress, tumour stage and chromatin structure. Using RNA-flourescent in situ hybridization (FISH), we show that TelRNAs are novel structural components of telomeric chromatin. Importantly, we provide evidence that TelRNAs block the activity of telomerase in vitro, suggesting that TelRNAs may regulate telomerase activity at chromosome ends. Our results indicate that TelRNAs are novel components of mammalian telomeres, which are anticipated to be fundamental for understanding telomere biology and telomere-related diseases, such as cancer and ageing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
de Lange, T. Shelterin: the protein complex that shapes and safeguards human telomeres. Genes Dev. 19, 2100–2110 (2005).
Gonzalo, S. et al. DNA methyltransferases control telomere length and telomere recombination in mammalian cells. Nature Cell Biol. 8, 416–424 (2006).
Gonzalo, S. et al. Role of the RB1 family in stabilizing histone methylation at constitutive heterochromatin. Nature Cell Biol. 7, 420–428 (2005).
Garcia-Cao, M., Gonzalo, S., Dean, D. & Blasco, M. A. A role for the Rb family of proteins in controlling telomere length. Nature Genet. 32, 415–419 (2002).
Garcia-Cao, M., O'Sullivan, R., Peters, A. H., Jenuwein, T. & Blasco, M. A. Epigenetic regulation of telomere length in mammalian cells by the Suv39h1 and Suv39h2 histone methyltransferases. Nature Genet. 36, 94–99 (2004).
Benetti, R., Garcia-Cao, M. & Blasco, M. A. Telomere length regulates the epigenetic status of mammalian telomeres and subtelomeres. Nature Genet. 39, 243–250 (2007).
Benetti, R. et al. Suv4-20h deficiency results in telomere elongation and derepression of telomere recombination. J. Cell Biol. 178, 925–936 (2007).
Blasco, M. A. The epigenetic regulation of mammalian telomeres. Nature Rev. Genet. 8, 299–309 (2007).
Gottschling, D. E., Aparicio, O. M., Billington, B. L. & Zakian, V. A. Position effect at S. cerevisiae telomeres: reversible repression of Pol II transcription. Cell 63, 751–762 (1990).
Levis, R., Hazelrigg, T. & Rubin, G. M. Effects of genomic position on the expression of transduced copies of the white gene of Drosophila. Science 229, 558–561 (1985).
Nimmo, E. R., Cranston, G. & Allshire, R. C. Telomere-associated chromosome breakage in fission yeast results in variegated expression of adjacent genes. EMBO J. 13, 3801–3811 (1994).
Baur, J. A, Zou, Y., Shay, J. W. & Wright, W. E. Telomere position effect in human cells. Science 292, 2075–2077 (2001)
Koering, C. E. et al. Human telomeric position effect is determined by chromosomal context and telomeric chromatin integrity. EMBO Rep 3, 1055–1061 (2002).
Bernstein, E. & Allis, C. D. RNA meets chromatin. Genes Dev. 19, 1635–1655 (2005).
Blasco, M. A. et al. Telomere shortening and tumor formation by mouse cells lacking telomerase RNA. Cell 91, 25–34 (1997).
Mattick, J. S. & Makunin, I. V. Non-coding RNA. Hum. Mol. Genet. 15, R17–R29 (2006).
Valgardsdottir, R. et al. Structural and functional characterization of noncoding repetitive RNAs transcribed in stressed human cells. Mol. Biol. Cell 16, 2597–2604 (2005).
Rizzi, N. et al. Transcriptional activation of a constitutive heterochromatic domain of the human genome in response to heat shock. Mol. Biol. Cell 15, 543–551 (2004).
Jolly, C. et al. Stress-induced transcription of satellite III repeats. J. Cell Biol. 164, 25–33 (2004).
Masui, O. & Heard, E. RNA and protein actors in X-chromosome inactivation. Cold Spring Harb. Symp. Quant. Biol. 71, 419–428 (2006).
Franke, A. & Baker, B. S. The rox1 and rox2 RNAs are essential components of the compensasome, which mediates dosage compensation in Drosophila. Mol. Cell 4, 117–122 (1999).
Kelley, R. L. et al. Epigenetic spreading of the Drosophila dosage compensation complex from roX RNA genes into flanking chromatin. Cell 98, 513–522 (1999).
Cohen, H. R. & Panning, B. XIST RNA exhibits nuclear retention and exhibits reduced association with the export factor TAP/NXF1. Chromosoma 116, 373–383 (2007).
Chan, S. W. & Blackburn, E. H. New ways not to make ends meet: telomerase, DNA damage proteins and heterochromatin. Oncogene 21, 553–563 (2002).
Teixeira, M. T., Arneric, M., Sperisen, P. & Lingner, J. Telomere length homeostasis is achieved via a switch between telomerase-extendible and-nonextendible states. Cell 117, 323–335 (2004).
Ancelin, K. et al. Targeting assay to study the cis functions of human telomeric proteins: evidence for inhibition of telomerase by TRF1 and for activation of telomere degradation by TRF2. Mol. Cell Biol. 22, 3474–3487 (2002).
Azzalin, C. M., Reichenbach, P., Khoriauli, L., Giulotto, E. & Lingner, J. Telomeric repeat containing RNA and RNA surveillance factors at mammalian chromosome ends. Science 318, 798–801 (2007).
Peters, A. H. et al. Loss of the Suv39h histone methyltransferases impairs mammalian heterochromatin and genome stability. Cell 107, 323–337 (2001).
Murchison, E. P., Partridge, J. F., Tam, O. H., Cheloufi, S. & Hannon, G. J. Characterization of Dicer-deficient murine embryonic stem cells. Proc. Natl Acad. Sci. USA 102, 12135–12140 (2005).
Li, E., Bestor, T. H. & Jaenisch, R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69, 915–926 (1992).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
Wutz, A. & Jaenisch, R. A shift from reversible to irreversible X inactivation is triggered during ES cell differentiation. Mol. Cell 5, 695–705 (2000).
Benetti, R. et al. The death substrate Gas2 binds m-calpain and increases susceptibility to p53-dependent apoptosis. EMBO J. 20, 2702–2714 (2001).
Acknowledgements
We are indebted to T. Jenuwein for the Suv39h and Suv4-20 cells and to G. Hannon for the Dicer-null cells. We thank M. L. Cayuela for Danio rerio RNA. Thanks to R. Benetti, P. Klatt and A. Canela for helpful discussions and help with the preparation of the manuscript and to R. Serrano for mouse care and genotyping. S.S. is a postdoctoral fellow funded by EMBO. M.A.B's laboratory is funded by the Spanish Ministry of Education and Culture (SAF2001-1869, GEN2001-4856-C13-08), by the Regional Government of Madrid (08.1/0054/01), the European Union (TELOSENS FIGH-CT-2002-00217, INTACT LSHC-CT-2003-506803, ZINCAGE FOOD-CT-2003-506850, RISC-RAD FI6R-CT-2003-508842, MOL CANCER MED LSHC-CT-2004-502943) and the Josef Steiner Award 2003.
Author information
Authors and Affiliations
Contributions
S.S. and M.A.B designed the experiments and S.S. carried out the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary figures S1, S2, S3, S4 and S5 (PDF 740 kb)
Rights and permissions
About this article
Cite this article
Schoeftner, S., Blasco, M. Developmentally regulated transcription of mammalian telomeres by DNA-dependent RNA polymerase II. Nat Cell Biol 10, 228–236 (2008). https://doi.org/10.1038/ncb1685
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1685
This article is cited by
-
Telomeres and aging: on and off the planet!
Biogerontology (2024)
-
Chromosome ends and the theory of marginotomy: implications for reproduction
Biogerontology (2024)
-
TERRA G-quadruplex stabilization as a new therapeutic strategy for multiple myeloma
Journal of Experimental & Clinical Cancer Research (2023)
-
Sexual Dimorphism in Telomere Length in Childhood Autism
Journal of Autism and Developmental Disorders (2023)
-
Global and transcription-coupled repair of 8-oxoG is initiated by nucleotide excision repair proteins
Nature Communications (2022)