Abstract
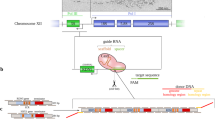
Chromatin structure is thought to play a critical role in gene expression. Using the lac operator/repressor system and two colour variants of green fluorescent protein (GFP), we developed a system to visualize a gene and its protein product directly in living cells, allowing us to examine the spatial organization and timing of gene expression in vivo. Dynamic morphological changes in chromatin structure, from a condensed to an open structure, were observed upon gene activation, and targeting of the gene product, cyan fluorescent protein (CFP) reporter to peroxisomes was visualized directly in living cells. We found that the integrated gene locus was surrounded by a promyelocytic leukaemia (PML) nuclear body. The association was transcription independent but was dependent upon the direct in vivo binding of specific proteins (EYFP/lac repressor, tetracycline receptor/VP16 transactivator) to the locus. The ability to visualize gene expression directly in living cells provides a powerful system with which to study the dynamics of nuclear events such as transcription, RNA processing and DNA repair.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Spector, D. L. Macromolecular domains within the cell nucleus. Annu. Rev. Cell Biol. 9, 265–315 ( 1993).
Lamond, A. I. & Earnshaw, W. C. Structure and function in the nucleus. Science 280, 547– 553 (1998).
Misteli, T. & Spector, D. L. Applications of the green fluorescent protein in cell biology and biotechnology. Nature Biotechnol. 15, 961–964 (1997).
Tsien, R. Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Ellenberg, J., Lippincott-Schwartz, J. & Presley, J. F. Dual-colour imaging with GFP variants . Trends Cell. Biol. 9, 52– 56 (1999).
Belmont, A. S. & Straight, A.F. In vivo visualization of chromosomes using lac operator–repressor binding. Trends Cell. Biol. 8, 121–124 ( 1998).
Robinett, C. C. et al. In vivo localization of DNA sequences and visualization of large-scale chromatin organization using lac operator/repressor recognition . J. Cell Biol. 135, 1685– 1700 (1996).
Li, G., Sudlow, G. & Belmont, A. S. Interphase cell cycle dynamics of a late-replicating, heterochromatic homogeneously staining region: precise choreography of condensation/decondensation and nuclear positioning. J. Cell Biol. 140, 975–989 (1998).
Tumbar, T., Sudlow, G. & Belmont, A. S. Large-scale chromatin unfolding and remodeling induced by VP16 acidic activation domain. J. Cell Biol. 145 , 1341–1354 (1999).
Straight, A. F., Belmont, A. S., Robinett, C. C. & Murray, A. W. GFP tagging of budding yeast chromosomes reveals that protein-protein interactions can mediate sister chromatid cohesion. Curr. Biol. 6, 1599–1608 (1996).
Minshull, J. et al. Protein phosphatase 2A regulates MPF activity and sister chromatid cohesion in budding yeast. Curr. Biol. 6, 1609–1620 (1996).
Miyazawa, S. et al. Peroxisome targeting signal of rat liver acyl-coenzyme A oxidase resides at the carboxy terminus. Mol. Cell. Biol. 9 , 83–91 (1989).
Miller, J. H. & Reznikoff, W. A. The Operon, 17–220 (Cold Spring Harbor Laboratory Press, New York, 1980).
Maul, G. G., Negorev, D., Bell, P. & Ishov, A. M. Properties and assembly mechanisms of ND10, PML bodies, or PODs. J. Struct. Biol. 129, 278–287 ( 2000).
Zhong, S., Salomoni, P. & Pandolfi, P. P. The transcriptional role of PML and the nuclear body . Nature Cell Biol. 2, E85– E90 (2000).
Weintraub, H. & Groudine, M. Chromosomal subunits in active genes have an altered conformation. Science 193, 848–856 (1976).
Paranjape, S. M., Kamakaka, R. T. & Kadonaga, J. T. Role of chromatin structure in the regulation of transcription by RNA polymerase II. Annu. Rev. Biochem. 63, 265–297 (1994).
Armstrong, J. A. & Emerson, B. M. Transcription of chromatin: these are complex times. Curr. Opin. Genet. Dev. 8, 165–172 ( 1998).
Kuo, M. H. & Allis, C. D. Roles of histone acetyltransferases and deacetylases in gene regulation. Bioessays 20, 615–626 (1998).
Pederson, T. Chromatin structure and gene transcription: nucleosomes permit a new synthesis . Int. Rev. Cytol. 55, 1– 21 (1978).
Manuelidis, L. Different central nervous system cell types display distinct and nonrandom arrangements of satellite DNA sequences. Proc. Natl Acad. Sci. USA 81, 3123–3127 ( 1984).
Manuelidis, L. & Borden, J. Reproducible compartmentalization of individual chromosome domains in human CNS cells revealed by in situ hybridization and three-dimensional reconstruction. Chromosoma 96 , 397–410 (1988).
Ferguson, M. & Ward, D. C. Cell cycle dependent chromosomal movement in pre-mitotic human T-lymphocyte nuclei. Chromosoma 101, 557–565 (1992).
Dietzel, S. et al. Three-dimensional distribution of centromeric or paracentromeric heterochromatin of chromosomes 1, 7, 15, and 17 in human lymphocyte nuclei studied with light microscopic axial tomography. Bioimaging 3, 121–133 (1995).
Abney, J. R., Cutler, B., Fillbach, M. L., Axelrod, D. & Scalettar, B. A. Chromatin dynamics in interphase nuclei and its implications for nuclear structure. J. Cell Biol. 137, 1459–1468 (1997).
Marshall, W. F. et al. Interphase chromosomes undergo constrained diffusional motion in living cells. Curr. Biol. 7, 930– 939 (1997).
Manuelidis, L. Individual interphase chromosome domains revealed by in-situ hybridization . Hum. Genet. 71, 288–293 (1985).
Janevski, J., Park, P. C. & De Boni, U. Organization of centromeric domains in hepatocyte nuclei: rearrangement associated with de novo activation of the vitellogenin gene family in Xenopus laevis. Exp. Cell Res. 217 , 227–239 (1995).
Bartholdi, M. F. Nuclear distribution of centromeres during the cell cycle of human diploid fibroblasts. J. Cell Sci. 99, 255– 263 (1991).
Funabiki, H., Hagan, I., Uzawa, S. & Yanagida, M. Cell cycle-dependent specific positioning and clustering of centromeres and telomeres in fission yeast. J. Cell Biol. 121, 961– 976 (1993).
LaSalle, J. M. & Lalande, M. Homologous association of oppositely imprinted chromosomal domains. Science 272, 725–728 (1996).
Borden, J. & Manuelidis, L. Movement of the X chromosome in epilepsy. Science 242, 1687 –1691 (1988).
Guldner, H. H., Szostecki, C., Grotzinger, T. & Will, H. IFN enhance expression of Sp100, an autoantigen in primary biliary cirrhosis . J. Immunol. 149, 4067– 4073 (1992).
Koken, M. H. et al. The t(15;17) translocation alters a nuclear body in a retinoic acid-reversible fashion. EMBO J. 13, 1073 –1083 (1994).
Korioth, F., Gieffers, C., Maul, G. G. & Frey, J. Molecular characterization of NDP52, a novel protein of the nuclear domain 10, which is redistributed upon virus infection and interferon treatment. J. Cell Biol. 130, 1–13 (1995).
Maul, G. G., Yu, E., Ishov, A. M. & Epstein, A. L. Nuclear domain 10 (ND10) associated proteins are also present in nuclear bodies and redistribute to hundreds of nuclear sites after stress. J. Cell. Biochem. 59, 498–513 (1995).
Ishov, A. M. & Maul, G. G. The periphery of nuclear domain 10 (ND10) as site of DNA virus deposition. J. Cell Biol. 134, 815–826 ( 1996).
Maul, G. G., Jensen, D. E., Ishov, A. M., Herlyn, M. & Rauscher, F. J. III Nuclear redistribution of BRCA1 during viral infection. Cell Growth Differ. 9 , 743–755 (1998).
Gongora, C. et al. Molecular cloning of a new interferon-induced PML nuclear body-associated protein. J. Biol. Chem. 272, 19457–19463 (1997).
Chen, C. & Okayama, H. High-efficiency transformation of mammalian cells by plasmid DNA. Mol. Cell. Biol. 7, 2745–2752 (1987).
Mintz, P. J., Patterson, S. D., Neuwald, A. F., Spahr, C. S. & Spector, D. L. Purification and biochemical characterization of interchromatin granule clusters. EMBO J. 18, 4308–4320 ( 1999).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular Cloning (Cold Spring Harbor Laboratory Press, New York, 1989).
Spector, D. L., Goldman, R. D. & Leinwand, L. A. Cells: A Laboratory Manual (Cold Spring Harbor Laboratory Press, New York, 1998).
Acknowledgements
We thank T. Misteli for discussions and P. Sacco-Bubulya and N. Saitoh for reviewing the manuscript. pW7-C3 (=pCFP-C3), which encoded CFP, was provided by R. Tsien, EV-124 was provided by M. Wilkinson and pUHD10-4B was provided by J. Skowronski, and monoclonal antibody 5E10 was obtained from R. van Driel. S.M.J. is supported by an NIH/NCI training grant 5T32CA09311. D.L.S. is funded by a grant from NIGMS (NIH 498100).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Movie 1
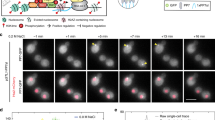
Visualization of gene expression Clone 2 cells were transiently transfected with EYFP/lac repressor and pTet-On and plated onto a coverslip fitted for a FCS2 live-cell chamber (Bioptechs, Butler, Pennsylvania). An image was acquired using OpenLab software (Improvision, Boston, Massachusetts); doxycline then was perfused into the live-cell chamber. Images were acquired every 10 minutes in the YFP channel for the first 2 h, after which images were obtained in bothe the YFP and CFP channels at 20-min intervals. The entire movie was taken over a 7-h period and is played back at 2,880 x. (MOV 3600 kb)
Rights and permissions
About this article
Cite this article
Tsukamoto, T., Hashiguchi, N., Janicki, S. et al. Visualization of gene activity in living cells. Nat Cell Biol 2, 871–878 (2000). https://doi.org/10.1038/35046510
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/35046510
This article is cited by
-
Transcription organizes euchromatin via microphase separation
Nature Communications (2021)
-
Copy number-dependent DNA methylation of the Pyricularia oryzae MAGGY retrotransposon is triggered by DNA damage
Communications Biology (2021)
-
Super‐resolution microscopy of genome organization
Genes & Genomics (2021)
-
Visualizing the genome in high resolution challenges our textbook understanding
Nature Methods (2020)
-
Tunable light and drug induced depletion of target proteins
Nature Communications (2020)