Abstract
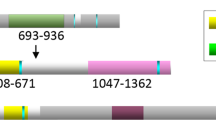
Breast cancer is the most common malignancy among women, and mutations in the BRCA genes produce increased susceptibility to these malignancies in certain families. Here we identify BRCA1-IRIS as a 1,399-amino-acid BRCA1 gene product encoded by an uninterrupted open reading frame that extends from codon 1 of the known BRCA1 open reading frame to a termination point 34 triplets into intron 11. Unlike full-length BRCA1 (p220), BRCA1-IRIS is exclusively chromatin-associated, fails to interact with BARD1 in vivo or in vitro and exhibits unique nuclear immunostaining. Unlike BRCA1FL (or p220), BRCA1-IRIS also co-immunoprecipitated with DNA-replication-licensing proteins and with known replication initiation sites. Suppression of BRCA1-IRIS expression hindered the normal departure of geminin from pre-replication complexes, and depressed the rate of cellular DNA replication and possibly initiation-related synthesis. In contrast, BRCA1-IRIS overexpression stimulated DNA replication. These data imply that endogenous BRCA1-IRIS positively influences the DNA replication initiation machinery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Hill, A.D., Doyle, J.M., McDermott, E.W. & O'Higgins, N.J. Hereditary breast cancer. Br. J. Surg. 84, 1334–1339 (1997).
Miki, Y. et al. A strong candidate for the breast and ovarian cancer susceptibility gene BRCA1. Science 266, 66–71 (1994).
Smith, S.A., Easton, D.F., Evans, D.G. & Ponder, B.A. Allele losses in the region 17q12-21 in familial breast and ovarian cancer involve the wild-type chromosome. Nature Genet. 2, 128–131 (1992).
Rice, J.C., Ozcelik, H., Maxeiner, P., Andrulis, I. & Futscher, B.W. Methylation of the BRCA1 promoter is associated with decreased BRCA1 mRNA levels in clinical breast cancer specimens. Carcinogenesis 9, 1761–1765 (2000).
Chen, Y. et al. BRCA1 is a 220-kDa nuclear phosphoprotein that is expressed and phosphorylated in a cell cycle-dependent manner. Cancer Res. 56, 3168–3172 (1996).
Wilson, C.A. et al. Differential subcellular localization, expression and biological toxicity of BRCA1 and the splice variant BRCA1-Δ11b. Oncogene 14, 1–16 (1997).
Lee, E.Y. BRCA1 and Chk1 in G2/M checkpoint: a new order of regulation. Cell Cycle. 1, 178–180 (2002).
Deng, C.X. & Scott, F. Role of the tumor suppressor gene Brca1 in genetic stability and mammary gland tumor formation. Oncogene 19, 1059–1064 (2000).
Hakem, R. et al. The tumor suppressor gene Brca1 is required for embryonic cellular proliferation in the mouse. Cell 85, 1009–1023 (1996).
Gudas, J.M. et al. Cell-cycle regulation of BRCA1 messenger RNA in human breast epithelial cells. Cell Growth Differ. 7, 717–723 (1996).
Chodosh, L.A. et al. Mammary gland development, reproductive history, and breast cancer risk. Cancer Res 59, 1765–1771s; discussion 1771s–1772s (1999).
Magdinier, F., Dalla Venezia, N., Lenoir, G.M., Frappart, L. & Dante, R. BRCA1 expression during prenatal development of the human mammary gland. Oncogene 18, 4039–4043 (1999).
Meza, J.E., Brzovic, P.S., King, M.C. & Klevit, R.E. Mapping the functional domains of BRCA1. Interaction of the ring finger domains of BRCA1 and BARD1. J. Biol. Chem. 274, 5659–5665 (1999).
Bochar, D.A. et al. BRCA1 is associated with a human SWI/SNF-related complex: linking chromatin remodeling to breast cancer. Cell. 102, 257–265 (2000).
Yarden, R.I. & Brody, L.C. BRCA1 interacts with components of the histone deacetylase complex. Proc. Natl Acad. Sci. USA 96, 4983–4988 (1999).
Scully, R. Dynamic changes of BRCA1 subnuclear location and phosphorylation state are initiated by DNA damage. Cell. 90, 425–435 (1997).
Cohen, S.M., Brylawski, B.P., Cordeiro-Stone, M. & Kaufman, D.G. Mapping of an origin of DNA replication near the transcriptional promoter of the human HPRT gene. J. Cell. Biochem. 85, 346–356 (2002).
Abdurashidova, G. et al. Start sites of bidirectional DNA synthesis at the human lamin B2 origin. Science. 287, 2023–2026 (2000).
Tao, L., Dong, Z., Leffak, M., Zannis-Hadjopoulos, M. & Price, G. Major DNA replication initiation sites in the c-Myc locus in human cells. J. Cell Biochem. 78, 442–457 (2000).
Yoon, Y., Sanchez, J.A., Brun, C. & Huberman, J.A. Mapping of replication initiation sites in human ribosomal DNA by nascent-strand abundance analysis. Mol. Cell. Biol. 15, 2482–2489 (1995).
Taira, T., Iguchi-Ariga, S.M. & Ariga, H. A novel DNA replication origin identified in the human heat shock protein 70 gene promoter. Mol. Cell. Biol. 14, 6386–6397 (1994).
Kamath, S. & Leffak, M. Multiple sites of replication initiation in the human β-globin gene locus. Nucleic Acids Res. 29, 809–817 (2001).
Avni, D. et al. Active localization of the retinoblastoma protein in chromatin and its response to S phase DNA damage. Mol. Cell 12, 735–746 (2003).
Roy, I. et al. Efficient translocation and apoptosis induction by adenovirus encoded VP22-p53 fusion protein in human tumor cells in vitro. Anticancer Res. 22, 3185–3189 (2002).
Zender, L. et al. Gene therapy by intrahepatic and intratumoral trafficking of p53–VP22 induces regression of liver tumors. Gastroenterology 123, 608–618 (2002).
Ebihara, K. et al. Intron retention generates a novel isoform of the murine vitamin D receptor that acts in a dominant-negative way on the vitamin D signaling pathway. Mol. Cell. Biol. 16, 3393–3400 (1996).
Solomon, D. et al. Cyclin D1 Splice Varients. Differential effects on localization, Rb phosphorylation, and cultural transformation. J. Biol. Chem. 278, 30339–30347 (2003).
Kolpakova, E., Rusten, T.E. & Olsnes, S. Characterization and tissue expression of acidic fibroblast growth factor binding protein homologue in Drosophila melanogaster. Gene 310, 185–191 (2003).
Burkhead, J.L., Abdel-Ghany, S.E., Morrill, J.M., Pilon-Smits, E.A. & Pilon, M. The Arabidopsis thaliana CUTA gene encodes an evolutionarily conserved copper binding chloroplast protein. Plant J. 34, 856–867 (2003).
van Doorn, R. et al. A novel splice variant of the Fas gene in patients with cutaneous T-cell lymphoma. Cancer Res. 62, 5389–5392 (2002).
Bloor, B.K., Rajarajan, A., Jaafary-Haghighat, K. & Odell, E.W. Transcription and expression of CD44 variant exons by oro-pharyngeal squamous cell carcinomas. Int. J. Oncol. 21, 907–913 (2002).
Le Hir, H., Charlet-Berguerand, N., de Franciscis, V. & Thermes, C. 5′-End RET splicing: absence of variants in normal tissues and intron retention in pheochromocytomas. Oncology. 63, 84–91 (2002).
Liu, J.W., Chandra, D., Tang, S.H., Chopra, D. & Tang, D.G. Identification and characterization of Bimgamma, a novel proapoptotic BH3-only splice variant of Bim. Cancer Res. 62, 2976–2981 (2002).
Flowers, J.M., Powell, J.F., Leigh, P.N., Andersen, P. & Shaw, C.E. Intron 7 retention and exon 9 skipping EAAT2 mRNA variants are not associated with amyotrophic lateral sclerosis. Ann. Neurol. 49, 643–649 (2001).
Emmert, S., Schneider, T.D., Khan, S.G. & Kraemer, K.H. The human XPG gene: gene architecture, alternative splicing and single nucleotide polymorphisms. Nucleic Acids Res. 29, 1443–1452 (2001).
Lejeune, F., Cavaloc, Y. & Stevenin, J. Alternative splicing of intron 3 of the serine/arginine-rich protein 9G8 gene. Identification of flanking exonic splicing enhancers and involvement of 9G8 as a trans-acting factor. J. Biol. Chem. 276, 7850–7858 (2000).
Celebi, J.T., Wanner, M., Ping, X.L., Zhang, H. & Peacocke, M. Association of splicing defects in PTEN leading to exon skipping or partial intron retention in Cowden syndrome. Hum. Genet. 107, 234–238 (2000).
Lindahl, M., Timmusk, T., Rossi, J., Saarma, M. & Airaksinen, M.S. Expression and alternative splicing of mouse Gfra4 suggest roles in endocrine cell development. Mol. Cell. Neurosci. 15, 522–533 (2000).
Williams R. et al. Detection of protein folding defects caused by BRCA1–BRCT truncation and missense mutations. J. Biol. Chem. 278, 53007–53016 (2003).
Shao, N., Chai, Y.L., Shyam, E., Reddy, P. & Rao, V.N. Induction of apoptosis by the tumor suppressor protein BRCA1. Oncogene. 13, 1–7 (1996).
Holt, J.T. et al. Growth retardation and tumour inhibition by BRCA1. Nature Genet. 12, 298–302 (1996).
Gavin, K.A., Hidaka, M. & Stillman, B. Conserved initiator proteins in eukaryotes. Science. 270, 1667–1671 (1995).
Tada, S. et al. Repression of origin assembly in metaphase depends on inhibition of RLF-B/Cdt1 by geminin. Nature Cell Biol. 3, 107–113 (2001).
Wohlschlegel, J.A. et al. Inhibition of eukaryotic DNA replication by geminin binding to Cdt1. Science. 290, 2309–2312 (2000).
Tada, S., Li, A., Maiorano, D., Mechali, M., Blow, J.J. Repression of origin assembly in metaphase depends on inhibition of RLF-B/Cdt1 by geminin. Nature Cell Biol. 3, 107–113 (2001).
Bermejo, R., Vilaboa, N. & Cales, C. Regulation of CDC6, geminin, and CDT1 in human cells that undergo polyploidization. Mol. Biol. Cell 13, 3989–4000 (2002).
Hodgson, B., Li, A., Tada, S. & Blow, J.J. Geminin becomes activated as an inhibitor of Cdt1/RLF-B following nuclear import. Curr. Biol. 16, 678–683 (2002).
Magdinier, F., Ribieras, S., Lenoir, G.M., Frappart, L. & Dante, R. Down-regulation of BRCA1 in human sporadic breast cancer; analysis of DNA methylation patterns of the putative promoter region. Oncogene 17, 3169–3176 (1998).
Gayther, S. et al. Germline mutations of the BRCA1 gene in breast and ovarian cancer families provide evidence for a genotype-phenotype correlation. Nature Genet. 11, 428–433 (1995).
Harlow, E. & Lane, D. Antibodies, a Laboratory Manual. (Cold Spring Harbor Laboratory, New York, 1988).
Xu, C.F. et al. Distinct transcription start sites generate two forms of BRCA1 mRNA. Hum. Mol. Genet. 4, 2259–2264 (1995).
Coffman, F.D. et al. In vitro replication of plasmids containing human ribosomal gene sequence: Origin localization and dependence on an Aprotinin-binding cytosolic protein. Exp. Cell Res. 209, 123–132 (1993).
Zink, D. et al. Structure and dynamics of human interphase chromosome territories in vivo. Hum. Genet. 102, 241–251 (1998).
Acknowledgements
We wish to thank R. Laskey for his comments following the reading of this manuscript. We are also grateful to V. Joukov for sharing his IMR90 cDNA library and D. Avni, for sharing the results of his unpublished data with us. We also thank J. Lane for technical assistance. This work was supported by grants from the National Cancer Institute (N.C.I.). W.E. was supported, in part, by a European Molecular Biology Organization (EMBO) Fellowship and in part by a “Massachusetts Department of Public Health Breast Cancer Research Grant”, number 30481126109. W.E. is currently supported by a grant from the N.C.I. Specialized Program of Research Excellence (SPORE) in breast cancer research at the Dana-Farber/Harvard Cancer Center.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information, Figures
Fig. S1, Fig. S2, Fig. S3, Fig. S4 and Table 1 (PDF 1223 kb)
Rights and permissions
About this article
Cite this article
ElShamy, W., Livingston, D. Identification of BRCA1-IRIS, a BRCA1 locus product. Nat Cell Biol 6, 954–967 (2004). https://doi.org/10.1038/ncb1171
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1171
This article is cited by
-
The pro- and anti-tumor roles of mesenchymal stem cells toward BRCA1-IRIS-overexpressing TNBC cells
Breast Cancer Research (2019)
-
Premature polyadenylation of MAGI3 is associated with diminished N6-methyladenosine in its large internal exon
Scientific Reports (2018)
-
The role of BRCA1-IRIS in the development and progression of triple negative breast cancers in Egypt: possible link to disease early lesion
BMC Cancer (2017)
-
Detecting splicing patterns in genes involved in hereditary breast and ovarian cancer
European Journal of Human Genetics (2017)
-
BRCA1-IRIS inactivation overcomes paclitaxel resistance in triple negative breast cancers
Breast Cancer Research (2015)