Abstract
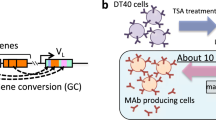
We show that iterative antigen-mediated selection of B-cell lines that constitutively hypermutate their immunoglobulin V genes during culture can be exploited to generate antibodies in vitro. From Ramos, a hypermutating human B-cell line expressing IgM of unknown specificity, we derived descendants that exhibit stepwise improved binding to streptavidin. Binding is initially conferred by mutations in complementarity-determining regions (CDRs), but maturation is due to strategic framework mutations. A more powerful system is provided by a hypermutating chicken B-lymphoma line, owing to its rapid proliferation, high rate of mutation accumulation, and genetic tractability. Starting from a single cell, we selected parallel lineages of derivatives, making mutated antibodies of increasing affinity to independent test antigens. Selection is initiated at an exceedingly low affinity threshold, but antibodies can be delivered with nanomolar affinities. The strategy could prove useful for in vitro generation of antigen-specific monoclonal antibodies and may be extendable to the maturation of other protein–ligand interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rajewsky, K. Clonal selection and learning in the antibody system. Nature 381, 751–758 (1996).
Parham, P. (ed.) Somatic hypermutation of immunoglobulin genes. Immunol. Rev. vol. 162 (1998).
Wagner, S.D., Milstein, C. & Neuberger, M.S. Codon bias targets mutation. Nature 376, 732 (1995).
Jolly, C.J. et al. The targeting of somatic hypermutation. Semin. Immunol. 8, 159–168 (1996).
Winter, G., Griffiths, A.D., Hawkins, R.E. & Hoogenboom, H.R. Making antibodies by phage display technology. Annu. Rev. Immunol. 12, 433–455 (1994).
Arnold, F.H. (ed.). Evolutionary Protein Design (Academic Press, San Diego, 2000) (Advances in Protein Chemistry, Vol. 55).
Gram, H. et al. In vitro selection and affinity maturation of antibodies from a naive combinatorial immunoglobulin library. Proc. Natl. Acad. Sci. USA 89, 3576–3580 (1992).
Low, N.M., Holliger, P. & Winter, G. Mimicking somatic hypermutation: affinity maturation of antibodies displayed on bacteriophage using a bacterial mutator strain. J. Mol. Biol. 260, 359–368 (1996).
Chowdhury, P.S. & Pastan, I. Improving antibody affinity by mimicking somatic hypermutation in vitro. Nat. Biotechnol. 17, 568–572 (1999).
Hanes J., Schaffitzel, C., Knappik, A. & Pluckthun, A. Picomolar affinity antibodies from a fully synthetic naive library selected and evolved by ribosome display. Nat. Biotechnol. 18, 1287–1292 (2000).
Boder, E.T., Midelfort, K.S. & Wittrup, K.D. Directed evolution of antibody fragments with monovalent femtomolar antigen-binding affinity. Proc. Natl. Acad. Sci. USA 97, 10701–10715 (2000).
Schier, R. et al. Isolation of picomolar affinity anti-c-erbB2 single-chain Fv by molecular evolution of the complementarity determining regions in the center of the antibody binding site. J. Mol. Biol. 263, 551–567 (1996).
Sale, J.E. & Neuberger, M.S. TdT-accessible breaks are scattered over the immunoglobulin V domain in a constitutively hypermutating B cell line. Immunity 9, 859–869 (1998).
Sale, J.E., Calandrini, D.M., Takata, M., Takeda, S. & Neuberger, M.S. Ablation of XRCC2/3 transforms immunoglobulin V gene conversion into somatic hypermutation. Nature 412, 921–926 (2001).
Yin, J. et al. A comparative analysis of the immunological evolution of antibody 28B4. Biochemistry 40, 10764–10773 (2001).
Betz, A.G., Rada, C., Pannell, R., Milstein, C. & Neuberger, M.S. Passenger transgenes reveal intrinsic specificity of the antibody hypermutation mechanism: clustering, polarity, and specific hot spots. Proc. Natl. Acad. Sci. USA 90, 2385–2388 (1993).
Foote, J. & Milstein, C. Conformational isomerism and the diversity of antibodies. Proc. Natl. Acad. Sci. USA 91, 10370–10374 (1994).
Baba, T.W., Giroir, B.P. & Humphries, E.H. Cell lines derived from avian lymphomas exhibit two distinct phenotypes. Virology 144, 139–151 (1985).
Buerstedde, J.M. & Takeda, S. Increased ratio of targeted to random integration after transfection of chicken B cell lines. Cell 67, 179–188 (1991).
Gulley, M.L., Ogata, L.C., Thorson, J.A., Dailey, M.O. & Kemp, J.D. Identification of a murine pan-T cell antigen which is also expressed during the terminal phases of B cell differentiation. J. Immunol. 140, 3751–3757 (1988).
Casali, P. & Schettino, E.W. Structure and function of natural antibodies. Curr. Top. Microbiol. Immunol. 10, 167–179 (1996).
Bouvet, J.P. & Dighiero, G. From natural polyreactive autoantibodies to a la carte monoreactive antibodies to infectious agents: is it a small world after all? Infect. Immun. 66, 1–4 (1998).
Bruggemann, M. et al. A repertoire of monoclonal antibodies with human heavy chains from transgenic mice. Proc. Natl. Acad. Sci. USA 86, 6709–6713 (1989).
Wagner, S.D. et al. The diversity of antigen-specific monoclonal antibodies from transgenic mice bearing human immunoglobulin gene miniloci. Eur. J. Immunol. 24, 2672–2681 (1994).
Lonberg, N. et al. Antigen-specific human antibodies from mice comprising four distinct genetic modifications. Nature 368, 856–859 (1994).
Fishwild, D.M. et al. High-avidity human IgGκ monoclonal antibodies from a novel strain of minilocus transgenic mice. Nat. Biotechnol. 14, 845–851 (1996).
Xu, J.L. & Davis, M.M. Diversity in the CDR3 region of V(H) is sufficient for most antibody specificities. Immunity 13, 37–45 (2000).
Yélamos, J. et al. Targeting of non-Ig sequences in place of the V segment by somatic hypermutation. Nature 376, 225–229 (1995).
Hulme, E.C. in Receptor Biochemistry: A Practical Approach (ed. Hulme, E.C.) 303–326 (IRL Press, Oxford, UK, 1990).
Morea, V., Lesk, A.M. & Tramontano, A. Antibody modelling: implications for engineering and design. Methods 20, 267–279 (2000).
Acknowledgements
We thank Veronica Morea for the model of the Ramos structure, Andy Riddell and Andy Johnson for help with flow cytometry, Alan Weeds for advice, and the late César Milstein for many inspiring discussions. S.L.D. and F.D.B. were in part supported by grants from the Leukaemia Research Fund and Arthritis Research Campaign, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Cumbers, S., Williams, G., Davies, S. et al. Generation and iterative affinity maturation of antibodies in vitro using hypermutating B-cell lines. Nat Biotechnol 20, 1129–1134 (2002). https://doi.org/10.1038/nbt752
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt752
This article is cited by
-
Methods and cell-based strategies to produce antibody libraries: current state
Applied Microbiology and Biotechnology (2021)
-
Ex vivo evolution of human antibodies by CRISPR-X: from a naive B cell repertoire to affinity matured antibodies
BMC Biotechnology (2019)
-
Synthetic evolution
Nature Biotechnology (2019)
-
A Novel DT40 Antibody Library for the Generation of Monoclonal Antibodies
Virologica Sinica (2019)
-
Transient AID expression for in situ mutagenesis with improved cellular fitness
Scientific Reports (2018)