Abstract
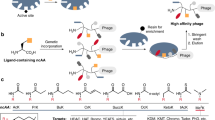
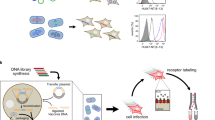
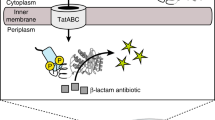
Phage display libraries are widely used for selection and optimization of polypeptide ligands or protease substrates. Because they are expressed and amplified in bacterial hosts, phage are not ideal for displaying eukaryotic polypeptldes or for probing mammalian cells. As retroviruses do not suffer from these limitations we constructed plasmids encoding replication-competent murine leukemia viruses displaying a virally encoded epidermal growth factor (EGF) domain at the N-terminus of the envelope glycoprotein. The EGF-displaying viruses replicated freely on EGF receptor-poor cells without deleting the displayed EGF domain but did not propagate on EGF receptor-rich cells because they were sequestered by the EGF receptors. A retrovirus display library was then generated by diversifying the seven-residue linker between the envelope glycoprotein and the displayed EGF domain. Selective pressure for loss of EGF receptor-binding activity was applied to the library by serial passage on EGF receptor-rich HT1080 cells. The selected viruses propagated on these cells with wild-type efficiencies, a phenotype that was conferred by intracellular cleavage of their displayed linker sequences. The selected linker sequences invariably presented arginine-rich motifs matching the consensus cleavage signal for furin-like proteases. Retrovirus display libraries can be used for the selection of polypepHdes interacting with components of living mammalian cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Little, M., Fuchs, P., Breitling, F., and Dübel, S. 1993. Bacterial surface presentation of proteins and peptides: an alternative to phage technology? Trends in Biotechnology 11: 3–5.
Stahl, S. and Uhlen, M. 1997. Bacterial surface display: trends and progress. Trends in Biotechnology 15: 185–192.
Boder, E.T. and Withrup, K.D. 1997. Yeast surface display for screening combinatorial polypeptlde libraries. Nat. Biotechnol. 15: 553–557.
O'Neil, K.T. and Hoess, R.H. 1995. Phage display: protein engineering by direct evolution. Curr. Opln. Struct. Blot. 5: 443–449.
Winter, G., Grifiths, A.D., Hawkins, R.E., and Hoogenboom, H.R. 1994. Making antibodies by phage display technology. Annu. Rev. Immunol. 12: 433–455.
Matthews, D.J. and Wells, J.A. 1993. JA Substrate phage: selection of protease substrates by monovalent phage display. Science 260: 11–13.
Russell, S.J., Hawkins, R.E. and Winter, G. 1993. Retroviral vectors displaying functional antibody fragments. Nucleic Acids Res. 21: 1081–1085.
Cosset F-L and Russell, S.J., 1996. Targeting retrovirus entry.Gene Ther 3: 946–956.
Nilson, B.H.K., Morilng, F.J., and Russell, S.J. 1996. Targeting of retroviral vectors through protease-substrate interactions. Gene Ther. 3: 280–286.
Peng, K-W., Morling, F.J., Cosset, F-L, Murphy, G., and Russell,, S.J. 1997. A gene delivery system activatable by disease-associated matrix metalloproteinases. Hum. Gene Ther. 8: 729–738.
Cosset, F-L, Morilng, F.J., Takeuchi, Y., Weiss, R.A., Collins, M.K.L, and Russell, S.J. 1995. Retroviral retargeting by envelopes expressing an N-terminal binding domain. J. Virol. 69: 6314–6322.
Nakayama,K 1997. Furin: arriammalian subtilisin/Kex2p-like endoprotease involved in processing of a wide variety of precursor proteins. Biochemical Journal 327: 625–635.
Seidah, N.G. and Chretien, M. 1997. Eukaryotic protein processing: endoproteolysis of precursor proteins. Curr. Opln. Biotechnol. 8: 602–607.
Temin, H.M. 1993. Retrovirus variation and reverse transcription: abnormal strand transfers result in retrovirus genetic variation. Proc. Natl. Acad. Sci. USA 90: 900–6903.
Colicelli, J. and Goff, S.P. 1986. Sequence and spacing requirements of a retrovirus integration site. J. Mol. Biol. 199: 47–59.
Shinnick, T.M., Lerner, R.A., and Sutoliffe, G. 1981. Nucleotide sequence of Moloney murine leukaemia virus. Nature 293: 543–548.
Bell, G.I., Fong, N.M., Stempien, M.M., Wornsted MA, Caput, D., Ku, L.L. et al. 1986. Human epidermal growth factor precursor: cDNA sequence expression In vitro and gene organisation. Nucleic Acids Res. 11: 8427–8446.
Coombs, S.G., Dang, AT., Madison, E.L, and Corey, D.R. 1996. Distinct mechanisms contribute to stringent substrate specificity of tissue-plasminogen activator. J. Biol. Chem. 271: 4461–4467.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Buchholz, C., Peng, KW., Morling, F. et al. In vivo selection of protease cleavage sites from retrovirus display libraries. Nat Biotechnol 16, 951–954 (1998). https://doi.org/10.1038/nbt1098-951
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt1098-951
This article is cited by
-
RNAi-mediated gene silencing in tumour tissue using replication-competent retroviral vectors
Gene Therapy (2011)
-
Cell entry targeting restricts biodistribution of replication-competent retroviruses to tumour tissue
Gene Therapy (2008)
-
Targeted Cell Entry of Lentiviral Vectors
Molecular Therapy (2008)
-
Replicating retroviral vectors mediating continuous production and secretion of therapeutic gene products from cancer cells
Cancer Gene Therapy (2005)
-
Chronic gene delivery of interferon-inducible protein 10 through replication-competent retrovirus vectors suppresses tumor growth
Cancer Gene Therapy (2005)