Abstract
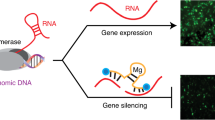
RNA interference (RNAi) is the process of sequence-specific post-transcriptional gene silencing triggered by double-stranded RNAs1,2,3. In attempts to identify RNAi triggers that effectively function at lower concentrations, we found that synthetic RNA duplexes 25–30 nucleotides in length can be up to 100-fold more potent than corresponding conventional 21-mer small interfering RNAs (siRNAs). Some sites that are refractory to silencing by 21-mer siRNAs can be effectively targeted by 27-mer duplexes, with silencing lasting up to 10 d. Notably, the 27-mers do not induce interferon or activate protein kinase R (PKR). The enhanced potency of the longer duplexes is attributed to the fact that they are substrates of the Dicer endonuclease, directly linking the production of siRNAs to incorporation in the RNA-induced silencing complex. These results provide an alternative strategy for eliciting RNAi-mediated target cleavage using low concentrations of synthetic RNA as substrates for cellular Dicer-mediated cleavage.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Hannon, G.J. RNA interference. Nature 418, 244–251 (2002).
Hutvagner, G. & Zamore, P.D. RNAi: nature abhors a double-strand. Curr. Opin. Genet. Dev. 12, 225–232 (2002).
Sharp, P.A. RNAi and double-strand RNA. Genes Dev. 13, 139–141 (1999).
Napoli, C., Lemieux, C. & Jorgensen, R. Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 2, 279–289 (1990).
Romano, N. & Macino, G. Quelling: transient inactivation of gene expression in Neurospora crassa by transformation with homologous sequences. Mol. Microbiol. 6, 3343–3353 (1992).
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806–811 (1998).
Stark, G.R., Kerr, I.M., Williams, B.R., Silverman, R.H. & Schreiber, R.D. How cells respond to interferons. Annu. Rev. Biochem. 67, 227–264 (1998).
Minks, M.A., Benvin, S., Maroney, P.A. & Baglioni, C. Synthesis of 2′–5′-oligo(A) in extracts of interferon-treated HeLa cells. J. Biol. Chem. 254, 5058–5064 (1979).
Elbashir, S.M., Lendeckel, W. & Tuschl, T. RNA interference is mediated by 21- and 22-nucleotide RNAs. Genes Dev. 15, 188–200 (2001).
Zamore, P.D., Tuschl, T., Sharp, P.A. & Bartel, D.P. RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 101, 25–33 (2000).
Bernstein, E., Caudy, A.A., Hammond, S.M. & Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363–366 (2001).
Elbashir, S.M. et al. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494–498 (2001).
Elbashir, S.M., Martinez, J., Patkaniowska, A., Lendeckel, W. & Tuschl, T. Functional anatomy of siRNAs for mediating efficient RNAi in Drosophila melanogaster embryo lysate. EMBO J. 20, 6877–6888 (2001).
Kim, D.H. et al. Interferon induction by siRNAs and ssRNAs synthesized by phage polymerase. Nat. Biotechnol. 22, 321–325 (2004).
Kim, D.H. & Rossi, J.J. Coupling of RNAi-mediated target downregulation with gene replacement. Antisense Nucleic Acid Drug Dev. 13, 151–155 (2003).
Persengiev, S.P., Zhu, X. & Green, M.R. Nonspecific, concentration-dependent stimulation and repression of mammalian gene expression by small interfering RNAs (siRNAs). RNA 10, 12–18 (2004).
Reynolds, A. et al. Rational siRNA design for RNA interference. Nat. Biotechnol. 22, 326–330 (2004).
Scherer, L. & Rossi, J.J. RNAi applications in mammalian cells. Biotechniques 36, 557–561 (2004).
Ui-Tei, K. et al. Guidelines for the selection of highly effective siRNA sequences for mammalian and chick RNA interference. Nucleic Acids Res. 32, 936–948 (2004).
Amarzguioui, M. & Prydz, H. An algorithm for selection of functional siRNA sequences. Biochem. Biophys. Res. Commun. 316, 1050–1058 (2004).
Markovtsov, V. et al. Cooperative assembly of an hnRNP complex induced by a tissue-specific homolog of polypyrimidine tract binding protein. Mol. Cell. Biol. 20, 7463–7479 (2000).
Wolin, S.L. & Cedervall, T. The La protein. Annu. Rev. Biochem. 71, 375–403 (2002).
Manche, L., Green, S.R., Schmedt, C. & Mathews, M.B. Interactions between double-stranded RNA regulators and the protein kinase DAI. Mol. Cell. Biol. 12, 5238–5248 (1992).
Gunnery, S. & Mathews, M.B. RNA binding and modulation of PKR activity. Methods 15, 189–198 (1998).
Jackson, A.L. et al. Expression profiling reveals off-target gene regulation by RNAi. Nat. Biotechnol. 21, 635–637 (2003).
Bohula, E.A. et al. The efficacy of small interfering RNAs targeted to the type 1 insulin-like growth factor receptor (IGF1R) is influenced by secondary structure in the IGF1R transcript. J Biol. Chem. 278, 15991–15997 (2003).
Caplen, N.J., Parrish, S., Imani, F., Fire, A. & Morgan, R.A. Specific inhibition of gene expression by small double-stranded RNAs in invertebrate and vertebrate systems. Proc. Natl. Acad. Sci. USA 98, 9742–9747 (2001).
Siolas, D. et al. Synthetic shRNAs as potent RNAi triggers. Nat. Biotechnol. 23, in the press (2005).
Acknowledgements
D.K. is a Beckman Fellow. This work was supported by a grant from the Arnold and Mabel Beckman Foundation and the US National Institutes of Health (AI29329 and AI42552, and HL074704 to J.J.R.). The authors wish to dedicate this work to the memory of Arnold Beckman, who recently passed away.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
M.A.B. and S.D.R. are employed by Integrated DNA Technologies, an institution that may gain or lose financially as a result of the publication of this article.
Supplementary information
Supplementary Fig. 2
ESI mass spectra of the 27mer duplex EGFPS1 27+0 before (top) and after (bottom) incubation with Dicer are shown. (PDF 125 kb)
Supplementary Fig. 3
Sequence specificity of Dicer substrate 27mer dsRNAs. (PDF 156 kb)
Supplementary Fig. 4
SiRNAs and Dicer substrate dsRNAs do not induce interferons or activate PKR or generate specific “off target effects”. (PDF 253 kb)
Supplementary Table 1
Summary of oligonucleotide reagents. (PDF 113 kb)
Supplementary Table 2
Molecular weights of possible 21mer duplexes derived from the 27mer duplex by Dicer processing. (PDF 107 kb)
Rights and permissions
About this article
Cite this article
Kim, DH., Behlke, M., Rose, S. et al. Synthetic dsRNA Dicer substrates enhance RNAi potency and efficacy. Nat Biotechnol 23, 222–226 (2005). https://doi.org/10.1038/nbt1051
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1051
This article is cited by
-
RNA interference in the era of nucleic acid therapeutics
Nature Biotechnology (2024)
-
Oligonucleotide therapeutics and their chemical modification strategies for clinical applications
Journal of Pharmaceutical Investigation (2024)
-
Nucleic acid drug vectors for diagnosis and treatment of brain diseases
Signal Transduction and Targeted Therapy (2023)
-
Sequence determinant of small RNA production by DICER
Nature (2023)
-
Innate immune regulations and various siRNA modalities
Drug Delivery and Translational Research (2023)