Abstract
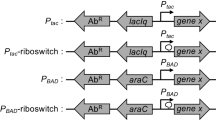
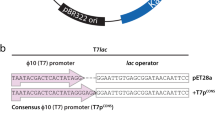
A broad host-range expression plasmid was constructed comprising the incQ rep-licon, the recA promoter from Escherichia coli and the g10-L. ribosome binding site (RBS) derived from bacteriophage T7. The structural genes for porcine soma-totropin (pst) and E. coli β-galactosidase (lacZ) were used to monitor gene expression in a diverse collection of Gram-negative bacterial hosts: Escherichia coli, Pseudomonas aeruginosa, Pseudomonas syringae, Pseudomonas putida, Pseudomonas fluorescens, Pseudomonas testosteroni, Serratia marcescens and Erwinia herbicola. The E. coli recA promoter was functional in this wide range of hosts and was inducible by the addition of nalidixic acid. Moreover, the level of lacZ expression was often at least as high as that observed in E. coli. Previous studies had shown that the g10-L RBS was superior to a simple “consensus” RBS sequence for expression of foreign genes in E. coli. Here we demonstrate a 38 to 70 fold increase in expression in two Pseudomonas hosts using the g10-L RBS, indicating that the translational enhancer present in the g10-L RBS is also functional in other bacteria. The juxtaposition of these transcriptional and translational elements in a broad host-range vector provides a simple way to evaluate alternate hosts for recombinant protein production.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Jeenes, D.J., Soldati, L., Baur, H., Watson, J.M., Mercenier, A., Reimmann, C., Leisinger, T. and Haas, D. 1986. Expression of biosynthetic genes from Pseudomonas aeruginosa and Escherichia coli in the heterologous host. Mol. Gen. Genet. 203: 421–429.
Mergeay, M., Boyen, A., Legrain, C. and Glansdorff, N. 1978. Expression of Escherichia coli K-12 arginine genes in Pseudomonas fluorescens. J. Bacteriol. 136: 1187–1188.
Martin, C., Cami, B., Borne, F., Jeenes, D.J., Haas, D. and Patte, J.C. 1986. Heterologous expression and regulation of the lysA genes of Pseudomonas aeruginosa and Escherichia coli. Mol. Gen. Genet. 203: 430–434.
Nagahari, K., Sano, Y. and Sakaguchi, K. 1977. Derepression of E. coli trp operon on interfamilial transfer. Nature 266: 745–746.
Werneke, J.M., Sligar, S.G. and Schuler, M.A. 1985. Development of broad host-range vectors for expression of cloned genes in Pseudomonas. Gene 38: 73–84.
Bagdasarian, M.M., Amann, E., Lurz, R., Ruckert, B. and Bagdasarian, M. 1983. Activity of the hybrid trp-lac (tac) promoter of E. coli in Pseudomonas putida. Construction of broad-host-range, controlled-expression vectors. Gene 26: 273–282.
Mermod, N., Ramos, J.L., Lehrbach, P.R. and Timmis, K.N. 1986. Vector for regulated expression of cloned genes in a wide range of Gram-negative bacteria. J. Bacteriol. 167: 447–454.
Olins, P.O., Devine, C.S., Rangwala, S.H. and Kavka, K.S. 1988. The T7 phage gene 10 leader RNA, a ribosome-binding site that dramatically enhances the expression of foreign genes in Escherichia coli. Gene 73: 227–235.
Olins, P.O. and Rangwala, S.H. 1989. A novel sequence element derived from bacteriophage T7 mRNA acts as an enhancer of translation of the lacZ gene in Escherichia coli. J. Biol. Chem. 264: 16973–16976.
Horii, T., Ogawa, T. and Ogawa, H. 1980. Organization of the recA gene of Escherichia coli. Proc. Natl. Acad. Sci. USA. 77: 313–317.
Better, M. and Helinski, D.R. 1983. Isolation and characterization of the recA gene of Rhizobium meliloti. J. Bacteriol. 155: 311–316.
Keener, S.L., McNamee, K.P. and McEntee, K. 1984. Cloning and characterization of recA genes from Proteus vulgaris, Erwinia caratovora, Shigella flexneri, and Escherichia coli B/r. J. Bacteriol. 160: 153–160.
Kokjohn, T.A. and Miller, R.V. 1985. Molecular cloning and characterization of the recA gene of Pseudomonas aeruginosa PAO. J. Bacteriol. 163: 568–572.
Zink, R.T., Engwall, J.K., McEvoy, J.L. and Chatterjee, A.K. 1985. recA is required in the induction of pectin lyase and carotovoricin in Erwinia carotovora subsp. carotovora. J. Bacteriol. 164: 390–396.
Goldberg, I. and Mekelanos, J.J. 1986. Cloning of the Vibrio cholerae recA gene and construction of a Vibrio cholerae recA mutant. J. Bacteriol. 165: 715–722.
Hickman, M.J., Orser, C.S., Willis, D.K., Lindow, S.E. and Panopoulos, N.J. 1987. Molecular cloning and biological characterization of the recA gene from Pseudomonas syringae. J. Bacteriol. 169: 2906–2910.
Benbrook, D.M. and Miller, R.V. 1986. Effects of norfloxacin on DNA metabolism in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 29: 1–6.
Messing, J. 1979. A multipurpose cloning system based on single-stranded DNA bacteriophage M13. Recombinant DNA technical bulletin, NIH publication ♯79–99, Vol. 2, No. 2:43–48.
Murray, K., Dugglebey, C.J., Sala-Trepat, J.M. and Williams, P. 1972. The metabolism of benzoate and methylbenzoates via the meta-cleavage pathway by Pseudomonas arvilla mt-2. Eur. J. Biochem. 28: 301–310.
Loper, J.E., Orser, C.S., Panopoulos, N.J. and Schroth, M.N. 1984. Genetic analysis of fluorescent pigment production in Pseudomonas syringae pv. syringae. J. Gen. Microbiol. 130: 1507–1515.
Drahos, D.J., Hemming, B.C. and McPherson, S. 1986. Tracking recombinant organisms in the environment: β-galactosidase as a selectable non-antibiotic marker for fluorescent pseudomonads. Bio/Technology 4: 439–444.
Hirota, Y. 1960. The effects of acridine dyes on mating type functions in Escherichia coli. Proc. Natl. Acad. Sci. USA. 46: 57–64.
Barry, G. 1988. A broad host-range shuttle system for gene insertion into the chromosome of Gram-negative bacteria. Gene 71: 75–84.
European Patent Application ♯193,515.
Dente, L., Cesarini, G. and Cortese, R. 1983. pEMBL: a new family of single-stranded plasmids. Nucleic Acids Res. 11: 1645–1655.
Maniatis, T., Fritsch, E. and Sambrook, J. 1982. Molecular Cloning, a Laboratory Manual. Cold Spring Harbor Laboratory, New York.
Ditta, G., Stanfield, S., Corbin, D. and Helinski, D.R. 1980. Broad host-range DNA cloning system for Gram-negative bacteria: Construction of a gene bank of Rhizobium meliloti. Proc. Natl. Acad. Sci. USA. 77: 7347–7351.
Renart, J., Reiser, J. and Stark, G.R. 1979. Transfer of proteins from gels to diazobenzyloxymethyl paper and detection with antisera: A method for studying antibody specificity and antigen structure. Proc. Natl. Acad. Sci. USA. 76: 3116–3120.
Miller, J.H. 1972. Experiments in Molecular Genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, New York.
Bradford, M.M. 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principal of protein-dye binding. Anal. Biochem. 72: 248–254.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Rangwala, S., Fuchs, R., Drahos, D. et al. Broad Host-Range Vector for Efficient Expression of Foreign Genes in Gram-Negative Bacteria. Nat Biotechnol 9, 477–479 (1991). https://doi.org/10.1038/nbt0591-477
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0591-477