Abstract
The genus Wolbachia is an archetype of maternally inherited intracellular bacteria that infect the germline of numerous invertebrate species worldwide. They can selfishly alter arthropod sex ratios and reproductive strategies to increase the proportion of the infected matriline in the population. The most common reproductive manipulation is cytoplasmic incompatibility, which results in embryonic lethality in crosses between infected males and uninfected females. Females infected with the same Wolbachia strain rescue this lethality. Despite more than 40 years of research1 and relevance to symbiont-induced speciation2,3, as well as control of arbovirus vectors4,5,6 and agricultural pests7, the bacterial genes underlying cytoplasmic incompatibility remain unknown. Here we use comparative and transgenic approaches to demonstrate that two differentially transcribed, co-diverging genes in the eukaryotic association module of prophage WO8 from Wolbachia strain wMel recapitulate and enhance cytoplasmic incompatibility. Dual expression in transgenic, uninfected males of Drosophila melanogaster crossed to uninfected females causes embryonic lethality. Each gene additively augments embryonic lethality in crosses between infected males and uninfected females. Lethality associates with embryonic defects that parallel those of wild-type cytoplasmic incompatibility and is notably rescued by wMel-infected embryos in all cases. The discovery of cytoplasmic incompatibility factor genes cifA and cifB pioneers genetic studies of prophage WO-induced reproductive manipulations and informs the continuing use of Wolbachia to control dengue and Zika virus transmission to humans.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Yen, J. H. & Barr, A. R. New hypothesis of the cause of cytoplasmic incompatibility in Culex pipiens L. Nature 232, 657–658 (1971)
Brucker, R. M. & Bordenstein, S. R. Speciation by symbiosis. Trends Ecol. Evol. 27, 443–451 (2012)
Shropshire, J. D. & Bordenstein, S. R. Speciation by symbiosis: the microbiome and behavior. MBio 7, e01785–15 (2016)
O’Connor, L. et al. Open release of male mosquitoes infected with a Wolbachia biopesticide: field performance and infection containment. PLoS Negl. Trop. Dis. 6, e1797 (2012)
Walker, T. et al. The wMel Wolbachia strain blocks dengue and invades caged Aedes aegypti populations. Nature 476, 450–453 (2011)
Carneiro Dutra, H. L. et al. Wolbachia blocks currently circulating Zika virus isolates in Brazilian Aedes aegypti mosquitoes. Cell Host Microbe 19, 771–774 (2016)
Zabalou, S. et al. Wolbachia-induced cytoplasmic incompatibility as a means for insect pest population control. Proc. Natl Acad. Sci. USA 101, 15042–15045 (2004)
Bordenstein, S. R. & Bordenstein, S. R. Eukaryotic association module in phage WO genomes from Wolbachia . Nature Commun. 7, 13155 (2016)
Ishmael, N. et al. Extensive genomic diversity of closely related Wolbachia strains. Microbiology 155, 2211–2222 (2009)
Beckmann, J. F. & Fallon, A. M. Detection of the Wolbachia protein WPIP0282 in mosquito spermathecae: implications for cytoplasmic incompatibility. Insect Biochem. Mol. Biol. 43, 867–878 (2013)
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes. J. Mol. Biol. 305, 567–580 (2001)
Lorenzen, M. D. et al. The maternal-effect, selfish genetic element Medea is associated with a composite Tc1 transposon. Proc. Natl Acad. Sci. USA 105, 10085–10089 (2008)
Zabalou, S. et al. Multiple rescue factors within a Wolbachia strain. Genetics 178, 2145–2160 (2008)
Poinsot, D., Bourtzis, K., Markakis, G., Savakis, C. & Mercot, H. Wolbachia transfer from Drosophila melanogaster into D. simulans: host effect and cytoplasmic incompatibility relationships. Genetics 150, 227–237 (1998)
Reynolds, K. T. & Hoffmann, A. A. Male age, host effects and the weak expression or non-expression of cytoplasmic incompatibility in Drosophila strains infected by maternally transmitted Wolbachia . Genet. Res. 80, 79–87 (2002)
Gutzwiller, F. et al. Dynamics of Wolbachia pipientis gene expression across the Drosophila melanogaster life cycle. G3 5, 2843–2856 (2015)
Yamada, R., Floate, K. D., Riegler, M. & O’Neill, S. L. Male development time influences the strength of Wolbachia-induced cytoplasmic incompatibility expression in Drosophila melanogaster . Genetics 177, 801–808 (2007)
Rørth, P. Gal4 in the Drosophila female germline. Mech. Dev. 78, 113–118 (1998)
White-Cooper, H. Tissue, cell type and stage-specific ectopic gene expression and RNAi induction in the Drosophila testis. Spermatogenesis 2, 11–22 (2012)
Serbus, L. R., Casper-Lindley, C., Landmann, F. & Sullivan, W. The genetics and cell biology of Wolbachia-host interactions. Annu. Rev. Genet. 42, 683–707 (2008)
Landmann, F., Orsi, G. A., Loppin, B. & Sullivan, W. Wolbachia-mediated cytoplasmic incompatibility is associated with impaired histone deposition in the male pronucleus. PLoS Pathog. 5, e1000343 (2009)
Lassy, C. W. & Karr, T. L. Cytological analysis of fertilization and early embryonic development in incompatible crosses of Drosophila simulans . Mech. Dev. 57, 47–58 (1996)
Callaini, G., Riparbelli, M. G., Giordano, R. & Dallai, R. Mitotic defects associated with cytoplasmic incompatibility in Drosophila simulans . J. Invertebr. Pathol. 67, 55–64 (1996)
Wright, J. D. & Barr, A. R. Wolbachia and the normal and incompatible eggs of Aedes polynesiensis (Diptera: Culicidae). J. Invertebr. Pathol. 38, 409–418 (1981)
Duron, O. & Weill, M. Wolbachia infection influences the development of Culex pipiens embryo in incompatible crosses. Heredity 96, 493–500 (2006)
Foe, V. E., Odell, G. M. & Edgar, B. A. in The Development of Drosophila melanogaster (eds Bate, M. & Martinez-Arias, A. ) Ch. 3 (Cold Spring Harbor Laboratory Press, 1993)
Zug, R. & Hammerstein, P. Still a host of hosts for Wolbachia: analysis of recent data suggests that 40% of terrestrial arthropod species are infected. PLoS ONE 7, e38544 (2012)
Jaenike, J., Dyer, K. A., Cornish, C. & Minhas, M. S. Asymmetrical reinforcement and Wolbachia infection in Drosophila . PLoS Biol. 4, e325 (2006)
Bordenstein, S. R., O’Hara, F. P. & Werren, J. H. Wolbachia-induced incompatibility precedes other hybrid incompatibilities in Nasonia . Nature 409, 707–710 (2001)
Bossan, B., Koehncke, A. & Hammerstein, P. A new model and method for understanding Wolbachia-induced cytoplasmic incompatibility. PLoS ONE 6, e19757 (2011)
Vallenet, D. et al. MicroScope: a platform for microbial genome annotation and comparative genomics. Database 2009, bap021 (2009)
Wu, M. et al. Phylogenomics of the reproductive parasite Wolbachia pipientis wMel: a streamlined genome overrun by mobile genetic elements. PLoS Biol. 2, e69 (2004)
Klasson, L. et al. The mosaic genome structure of the Wolbachia wRi strain infecting Drosophila simulans . Proc. Natl Acad. Sci. USA 106, 5725–5730 (2009)
Klasson, L. et al. Genome evolution of Wolbachia strain wPip from the Culex pipiens group. Mol. Biol. Evol. 25, 1877–1887 (2008)
Metcalf, J. A., Jo, M., Bordenstein, S. R., Jaenike, J. & Bordenstein, S. R. Recent genome reduction of Wolbachia in Drosophila recens targets phage WO and narrows candidates for reproductive parasitism. PeerJ 2, e529 (2014)
Foster, J. et al. The Wolbachia genome of Brugia malayi: endosymbiont evolution within a human pathogenic nematode. PLoS Biol. 3, e121 (2005)
Werren, J. H. & Loehlin, D. W. Rearing Sarcophaga bullata fly hosts for Nasonia (parasitoid wasp). Cold Spring Harb. Protoc. 2009, pdb.prot5308 (2009)
Bordenstein, S. R. & Bordenstein, S. R. Temperature affects the tripartite interactions between bacteriophage WO, Wolbachia, and cytoplasmic incompatibility. PLoS ONE 6, e29106 (2011)
Kearse, M. et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012)
Beckmann, J. F., Markowski, T. W., Witthuhn, B. A. & Fallon, A. M. Detection of the Wolbachia-encoded DNA binding protein, HU beta, in mosquito gonads. Insect Biochem. Mol. Biol. 43, 272–279 (2013)
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res. 36, W5–W9 (2008)
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004)
Hurvich, C. M. & Tsai, C.-L. A corrected Akaike information criterion for vector autoregressive model selection. J. Time Ser. Anal. 14, 271–279 (1993)
Abascal, F., Zardoya, R. & Posada, D. ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21, 2104–2105 (2005)
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 61, 539–542 (2012)
Finn, R. D. et al. The Pfam protein families database: towards a more sustainable future. Nucleic Acids Res. 44, D279–D285 (2016)
Letunic, I., Doerks, T. & Bork, P. SMART 7: recent updates to the protein domain annotation resource. Nucleic Acids Res. 40, D302–D305 (2012)
Casiraghi, M., Anderson, T. J., Bandi, C., Bazzocchi, C. & Genchi, C. A phylogenetic analysis of filarial nematodes: comparison with the phylogeny of Wolbachia endosymbionts. Parasitology 122, 93–103 (2001)
Chatzispyrou, I. A., Held, N. M., Mouchiroud, L., Auwerx, J. & Houtkooper, R. H. Tetracycline antibiotics impair mitochondrial function and its experimental use confounds research. Cancer Res. 75, 4446–4449 (2015)
Ferguson, S. B., Blundon, M. A., Klovstad, M. S. & Schupbach, T. Modulation of gurken translation by insulin and TOR signaling in Drosophila . J. Cell Sci. 125, 1407–1419 (2012)
Groth, A. C., Fish, M., Nusse, R. & Calos, M. P. Construction of transgenic Drosophila by using the site-specific integrase from phage phiC31. Genetics 166, 1775–1782 (2004)
Southall, T. D., Elliott, D. A. & Brand, A. H. The GAL4 system: a versatile toolkit for gene expression in Drosophila . Cold Spring Harb. Protoc. 2008, pdb.top49 (2008)
LePage, D. P., Jernigan, K. K. & Bordenstein, S. R. The relative importance of DNA methylation and Dnmt2-mediated epigenetic regulation on Wolbachia densities and cytoplasmic incompatibility. PeerJ 2, e678 (2014)
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nature Methods 9, 671–675 (2012)
Vizcaíno, J. A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016)
Acknowledgements
This work was supported by National Institutes of Health (NIH) R21 HD086833 and National Science Foundation IOS 1456778 to Seth R.B., National Science Foundation DEB-1501398 and NIH 5T32GM008554 training grant support to D.P.L., NIH T32GM07347 training grant support for J.A.M. to the Vanderbilt Medical Scientist Training Program, and NIH AI081322 to A.M.F. Imaging was performed in part through the use of the Vanderbilt University Medical Center Cell Imaging Shared Resource (supported by NIH grants CA68485, DK20593, DK58404, DK59637, and EY08126). We thank K. Jernigan and P. Snider for help with preliminary studies, and A. Brooks for assistance with figure preparation.
Author information
Authors and Affiliations
Contributions
D.P.L. performed gene expression and hatch rate assays, embryo cytology, and assayed for transgene and infection status of flies. J.A.M. performed comparative genomics analyses, generated transgenic flies, and drafted the manuscript. Sarah R.B. performed evolutionary and bioinformatic analyses. J.O. performed hatch rates, assayed sex ratios, collected flies for all experiments, and assayed for transgene and infection status of flies. J.I.P. conducted younger brother effect experiments and performed embryo cytology. J.D.S. performed hatch rate assays, collected flies for parallel embryo cytology, and assayed for transgene and infection status of flies. E.M.L. collected flies and performed hatch rate assays. L.J.F.-J. obtained the wVitA transcriptome. J.F.B. obtained the wPip proteome. Seth R.B. supervised the work and contributed to all experimental designs, data analysis, and data interpretation. All authors participated in manuscript preparation, editing, and final approval.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks S. L. O’Neill, W. Sullivan and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
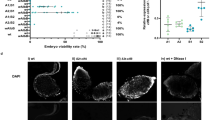
Extended Data Figure 1 CI and the evolution of Wolbachia and prophage WO genes.
a, The effect of parental Wolbachia infection on progeny viability and infection status. CI (embryonic inviability) occurs in crosses between Wolbachia-infected males and uninfected females. Wolbachia-infected females mated to infected males rescue the inviability. b, Bayesian phylogenies based on a 393-aa alignment of WD0723, the wMel ftsZ gene, and its homologues, and (c) a 70-aa alignment of WD0640, the phage WO gpW gene, and its homologues. Trees are based on JTT+G and CpRev+I models of evolution, respectively, and are unrooted. Consensus support values are shown at the nodes. Asterisk indicates that the CI genes are not included in Fig. 1. The WOPip5 homologue is truncated while the WOPip2 and second wAlbB homologues are highly divergent from WD0632.
Extended Data Figure 2 WD0631/WD0632 homologues associate with the eukaryotic association module in prophage WO regions.
CI gene homologues are labelled and coloured pink. Structural modules are labelled as baseplate, head, or tail. The WD0611–WD0621 label highlights a conserved gene cluster that is often associated with the CI genes. Only one phage haplotype is shown per Wolbachia strain when multiple copies of the same type are present.
Extended Data Figure 3 Wolbachia CI patterns correlate with WD0631/WD0632 homologue similarity and copy number.
a, The percentage aa identity between each WD0631/WD0632 homologue correlates with Wolbachia compatibility patterns. The only compatible cross, wMel males × wRi females, features close homology between WOMelB and WORiB. All other crosses are greater than 30% divergent and are bidirectionally incompatible. Each ‘% aa identity’ value is based on the region of query coverage in a 1:1 BLASTp analysis. b, CI strength, protein architecture, and clade type are listed for each of the Wolbachia strains shown in Fig. 1d. Asterisk indicates the proteins are disrupted and not included in comparison analyses.
Extended Data Figure 4 Wolbachia titres, the male age effect, and the younger brother effect.
a, Relative Wolbachia titres in WT lines do not decrease with age. DNA copy number of the wMel groEL gene is shown normalized to D. melanogaster rp49 gene copy number in testes at the indicated ages. b, Absolute Wolbachia titres do not decrease from day 1 to day 7 males. c, d, In wMel-infected males, WD0631 gene expression is equal between older (first day of emergence) and younger (fifth day of emergence) brothers while WD0632 gene expression is slightly higher in early emerging brothers. e, There is no statistical difference in CI penetrance between older and younger brothers. n = 8 for each group in a–d; n = 19–25 for each group in e. Bars, mean ± s.d. *P < 0.05, ***P < 0.001, ****P < 0.0001 by ANOVA with Kruskal–Wallis test and Dunn’s multiple test correction for a, b, and e, and two-tailed Mann–Whitney U-test for c and d. Exact P values are provided in Supplementary Table 7. These experiments were performed once.
Extended Data Figure 5 WD0625 transgene expression does not induce CI-like defects.
Expression of control gene WD0625 in 1-day-old uninfected males does not affect (a) embryo hatch rates or (b) sex ratios. Infection status is designated with filled symbols for a wMel-infected parent or open symbols for an uninfected parent. Transgenic flies are labelled with their transgene to the right of their male/female symbol. Unlabelled symbols represent WT flies. Data points are coloured according to the type of cross: blue, no CI; red, a CI cross; purple, a rescue cross with wMel-infected females. n = 18–47 for each group in a; n = 7–8 for b. Bars, mean ± s.d. *P < 0.05, ***P < 0.001 by ANOVA with Kruskal–Wallis test and Dunn’s multiple test correction. Exact P values are provided in Supplementary Table 7. This experiment was replicated three times.
Extended Data Figure 6 Expression of transgenes does not alter sex ratios.
Graphs correspond to the same crosses as in Fig. 3. Infection status is designated with filled symbols for a wMel-infected parent or open symbols for an uninfected parent. Transgenic flies are labelled with their transgene to the right of their gender symbol. Unlabelled gender symbols represent WT flies. Data points are coloured according to the type of cross: blue, no CI; red, a CI cross; purple, a rescue cross with wMel-infected females. n = 10–36 for each group. Bars, mean ± s.d. Statistics include a Kruskal–Wallis tests and Dunn’s multiple test corrections. The experiment in Extended Data Fig. 6a, c was performed once, while that in Extended Data Fig. 6b was performed twice.
Extended Data Figure 7 Transgenes are expressed in testes.
a, b, WD0508 and WD0625 transgenes are expressed in testes as evident by PCR performed against cDNA generated from dissected males used in Fig. 3a and Extended Data Fig. 5a, respectively. c, d, WD0631 and WD0632 transgenes are expressed in the testes from transgenic males specifically inducing high CI, no CI, or rescued CI. Testes were removed from males used in a replicate of Fig. 3b. n = six pools of six pairs of testes, with representative image shown. This experiment was performed once.
Extended Data Figure 8 Transgenic expression of WD0508, WD0625, and WD0625/WD0632 (cifB) does not enhance or induce CI.
a, The WD0508 transgene alone does not enhance CI in 2- to 4-day-old infected males. b, The WD0625 transgene alone does not enhance CI either; conversely, WD0632 does enhance CI as previously shown in Fig. 3c. The WD0625 transgene together with WD0632 does not enhance CI further than WD0632 alone. c, WD0625/WD0632 dual expression cannot induce CI in uninfected 1-day-old males. Infection status is designated with filled symbols for a wMel-infected parent or open symbols for an uninfected parent. Transgenic flies are labelled with their transgene to the right of their male/female symbol. Unlabelled symbols represent WT flies. Data points are coloured according to the type of cross: blue, no CI; red, a CI cross; purple, a rescue cross with wMel-infected females. n = 12–44 for each group. Bars, mean ± s.d. **P < 0.01, ***P < 0.001, ****P < 0.0001 by ANOVA with Kruskal–Wallis test and Dunn’s multiple test correction. Exact P values are provided in Supplementary Table 7. These experiments were done twice (a, c), three times (b, WD0625, WD0632), or once (b, WD0625/WD0632).
Extended Data Figure 9 Transgenic expression of control genes does not affect sex ratios.
All flies are from same crosses shown in Extended Data Fig. 8, except for c, which comes from a replicate experiment. Infection status is designated with filled symbols for a wMel-infected parent or open symbols for an uninfected parent. Transgenic flies are labelled with their transgene to the right of their male/female symbol. Unlabelled symbols represent WT flies. Data points are coloured according to the type of cross: blue, no CI; red, a CI cross; purple, a rescue cross with wMel-infected females. n = 4–27 for each group. Bars, mean ± s.d. Statistics performed by ANOVA with Kruskal–Wallis test and Dunn’s multiple test correction. These experiments were done twice (b) or once (a, c).
Extended Data Figure 10 There is variation in Wolbachia titres in transgenic lines.
a–c, Relative Wolbachia titres are higher in WD0508, WD0631, and WD0632 (cifB) transgenic lines than in WT lines. This does not occur in the WD0625 transgenic line, nor does there appear to be an additive effect. DNA copy number of the wMel groEL gene is shown normalized to D. melanogaster rp49 gene copy number in testes of the indicated strains. n = 8 independent pools of 15 pairs of testes for each group. Bars, mean ± s.d. *P < 0.05, ***P < 0.001, ****P < 0.0001 for two-tailed Mann–Whitney U-test (a) and Kruskal–Wallis test with Dunn’s multiple test correction (b, c). Exact P values are provided in Supplementary Table 7. These experiments were done once.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-7. (PDF 422 kb)
Source data
Rights and permissions
About this article
Cite this article
LePage, D., Metcalf, J., Bordenstein, S. et al. Prophage WO genes recapitulate and enhance Wolbachia-induced cytoplasmic incompatibility. Nature 543, 243–247 (2017). https://doi.org/10.1038/nature21391
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature21391
This article is cited by
-
Transgenic expression of cif genes from Wolbachia strain wAlbB recapitulates cytoplasmic incompatibility in Aedes aegypti
Nature Communications (2024)
-
Evolution of Wolbachia reproductive and nutritional mutualism: insights from the genomes of two novel strains that double infect the pollinator of dioecious Ficus hirta
BMC Genomics (2023)
-
Ability of a selfish B chromosome to evade genome elimination in the jewel wasp, Nasonia vitripennis
Heredity (2023)
-
Egg provisioning explains the penetrance of symbiont-mediated sex allocation distortion in haplodiploids
Heredity (2023)
-
pWCP is a widely distributed and highly conserved Wolbachia plasmid in Culex pipiens and Culex quinquefasciatus mosquitoes worldwide
ISME Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.