Abstract
The Southern Ocean houses a diverse and productive community of organisms1,2. Unicellular eukaryotic diatoms are the main primary producers in this environment, where photosynthesis is limited by low concentrations of dissolved iron and large seasonal fluctuations in light, temperature and the extent of sea ice3,4,5,6,7. How diatoms have adapted to this extreme environment is largely unknown. Here we present insights into the genome evolution of a cold-adapted diatom from the Southern Ocean, Fragilariopsis cylindrus8,9, based on a comparison with temperate diatoms. We find that approximately 24.7 per cent of the diploid F. cylindrus genome consists of genetic loci with alleles that are highly divergent (15.1 megabases of the total genome size of 61.1 megabases). These divergent alleles were differentially expressed across environmental conditions, including darkness, low iron, freezing, elevated temperature and increased CO2. Alleles with the largest ratio of non-synonymous to synonymous nucleotide substitutions also show the most pronounced condition-dependent expression, suggesting a correlation between diversifying selection and allelic differentiation. Divergent alleles may be involved in adaptation to environmental fluctuations in the Southern Ocean.
Similar content being viewed by others
Main
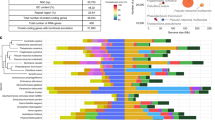
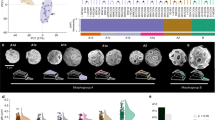
The pennate diatom genus Fragilariopsis is especially successful in the Southern Ocean, with the cold-adapted species F. cylindrus (Fig. 1a) regarded as an indicator species for polar water8,9,10. It is frequently found to form large populations in both the bottom layer of sea ice and the wider sea-ice zone, including open waters9 (Fig. 1b). Sea ice is characterized by temperatures under 0 °C, high salinity and, owing to the semi-enclosed pore system within the ice, low diffusion rates of dissolved gases and exchange of inorganic nutrients11. However, unlike in ice-free surface waters of the Southern Ocean12, dissolved iron is not considered to be limiting to phytoplankton growth within sea ice13. Most phytoplankton in the Southern Ocean face inclusion into sea ice every winter and are released again in summer when most of the sea ice melts14; certain species such as F. cylindrus have therefore evolved adaptations to cope with this drastic environmental change. Thus, comparative analyses of the genome of the psychrophile F. cylindrus with those of diatoms that evolved in temperate oceans provide an opportunity to obtain insights into how this species has adapted to conditions in Southern Ocean surface waters.
a, Scanning electron micrograph of two cells of F. cylindrus. b, Southern Ocean sea ice at the marginal ice-edge zone. The colored underside of the ice floe in the middle indicates dense populations of diatoms including F. cylindrus. c, Relative abundance of iron- and copper-binding proteins in selected eukaryotic genomes. Genomes are arranged according to genome size. d, Expansion of MYND zinc-binding domains as a function of total annotated domains for selected eukaryotic genomes.
We found many loci with highly divergent alleles in the diploid F. cylindrus draft genome sequence. To resolve the divergent alleles from paralogous genes, we independently carried out Sanger and PacBio sequencing and used haplotyped Sanger-finished fosmids to validate the haplotype-resolved genome assemblies (Supplementary Data 1–3). Using complementary approaches, we found that the F. cylindrus genome assembly consists of 15.1 Mb of loci with highly divergent alleles that were assigned to different scaffolds. The remaining 46 Mb of sequence consists of alleles similar enough to be assembled onto the same scaffold (Supplementary Information 2–5). The haplotype assembly size of the genome (61.1 Mb; Extended Data Table 1) was confirmed by quantitative PCR with reverse-transcription (qRT–PCR) (57.9 Mb). The genome completeness according to the Core Eukaryotic Genes Mapping Approach15 is 95.6% and the nuclear scaffold N50/L50 is 16/1.3 Mb, corresponding to assembly size (Extended Data Table 1).
The haplotype-resolved genome contains 21,066 predicted protein-coding genes (Extended Data Table 1) with 6,071 genes (29%) being represented by diverged alleles (Allele sets 1 and 2, Supplementary Data 1). Sequence divergence between alleles was up to 6%, but this was still significantly less (Mann–Whitney, P < 0.001) than that between paralogous genes (Extended Data Fig. 1 and Supplementary Information 4, 5).
We compared the F. cylindrus genome with those of Thalassiosira pseudonana16 and Phaeodactylum tricornutum17 (Extended Data Table 1), both of which live in temperate and neritic marine environments18,19 characterized by higher water temperatures, turbidity and concentrations of dissolved iron. The haploid gene content of F. cylindrus is enriched for two conserved metal-binding protein families (structural classification of proteins (SCOP) fold families; Supplementary Information 6). When accounting for its genome size, it is enriched for copper-binding but not iron-binding proteins (Fig. 1c), and it contains a disproportionate abundance of domains belonging to the plastocyanin/azurin-like family fold (SCOP ID 49504). Copper-containing plastocyanin may facilitate photosynthetic electron transport, reducing the need for iron20. There also appear to be more zinc-binding proteins in the F. cylindrus genome than in the other genomes, with 121 proteins containing zinc-binding myeloid–Nervy–DEAF-1 (MYND) domains, compared to 7 in T. pseudonana and 12 in P. tricornutum (Fig. 1d). MYND domains facilitate protein–protein interactions and are involved in regulatory processes21; most of those in F. cylindrus appear to be lineage-specific (Supplementary Information 6). Evolutionary genetic analysis of MYND-containing proteins suggests that this family has expanded within the last 30 million years (Supplementary Information 7). The relatively high zinc concentration of Southern Ocean surface waters22 may have facilitated the great expansion and functional divergence of zinc-binding MYND domains. The presence of lineage-specific protein families might indicate specific adaptations to the extreme conditions in the Southern Ocean. Some of these protein families appear to have been acquired through horizontal gene transfer from bacteria (Supplementary Information 8). Those proteins include groups of ice-binding proteins23 and proton-pumping proteorhodopsins24 (Supplementary Information 9, 10). There is also an unusually large number of genes for chlorophyll a/c light-harvesting complex (LHC) proteins, including 11 members of the Lhcx clade, which is involved in stress response (Extended Data Fig. 2 and Supplementary information 11).
Gene Ontology analysis show significant enrichment of genes in the categories ‘catalytic activity’, ‘transporter activity’, ‘metabolic process’, ‘transport’ and ‘integral to membrane’ in the group of diverged alleles compared to the non-diverged alleles (Fisher’s exact test, adjusted P < 0.05; Extended Data Table 2 and Supplementary Information 12). We found that similar processes (for example, transport and metabolic process) were enriched in metatranscriptome sequences from Southern Ocean sea ice (Figs 1b, 2a and Supplementary Data 4) with strong homology (BLASTx, E value ≤ 1 × 10−10) to F. cylindrus protein sequences of diverged alleles (Supplementary Information 13). According to these cut-off criteria, 64% of all Bacillariophyta-like metatranscriptome sequences had homology with proteins in F. cylindrus and around 60% of these sequences matched diverged alleles in the genome of F. cylindrus, including sequences from the enriched Gene Ontology categories (Supplementary Information 13).
a, REViGO semantic similarity scatterplot of biological process Gene Ontology terms for F. cylindrus-like sequences (E value ≤ 1 × 10−10) in Southern Ocean metatranscriptome samples. Gene Ontology terms that are overrepresented in the set of diverged alleles compared to non-diverged alleles are shown in bold. b, Hierarchical clustering of 4,030 differentially expressed allelic gene pairs in F. cylindrus (likelihood ratio test, P < 0.001; log2 fold change ≤ −2 or ≥+2) under low iron, freezing temperature (−2 °C), elevated temperature (+11 °C), elevated carbon dioxide (1,000 p.p.m. CO2) and prolonged darkness, relative to optimal growth conditions. Each experimental treatment corresponds to two separate columns for both allelic variants and each single-haplotype gene to a single row. c, Allele set-specific REViGO semantic similarity scatter plots for biological process Gene Ontology terms, showing relative differences between allele 1 and allele 2 in F. cylindrus RNA-seq experiments under low temperatures and low iron relative to optimal growth conditions using the cut-off ≥0.3 for relative difference.
RNA sequencing (RNA-seq) transcriptome profiling under environmentally relevant growth conditions (darkness, low iron, freezing, elevated temperature and CO2) identified stress-specific responses (Fig. 2b). The broadest transcriptome response (approximately 60% of total genes, including divergent alleles) was observed under prolonged darkness, characteristic of polar winters (Supplementary Information 14 and Supplementary Data 5). Placing F. cylindrus in darkness for seven days downregulated genes involved in photosynthesis, light harvesting and photoprotection relative to their expression under continuous light. By contrast, genes involved in starch, sucrose and lipid metabolism were strongly upregulated in the dark (Extended Data Fig. 3), indicating the utilization of chrysolaminarin and fatty acid storage products. Notably, under prolonged darkness, the percentage of RNA-seq reads that did not map to predicted genes (30%) was higher than under any other tested growth condition.
In allele-specific analyses of transcriptomes, approximately 66% (4,030) of diverged alleles showed greater than fourfold significant differential expression (likelihood ratio test, P < 0.001) relative to optimal nutrient-replete growth (Fig. 2b) and approximately 45% (2,730) of divergent alleles showed greater than fourfold unequal bi-allelic expression between allele 1 and allele 2 in at least one RNA-seq experiment (likelihood ratio test, P < 0.001; Supplementary Data 6). Additionally, the functional significance of this unequal bi-allelic expression for metabolism was inferred by an individual analysis of both sets of alleles using Gene Ontology. This demonstrated different metabolic signatures between the groups of divergent alleles (Fig. 2c). The differential expression of divergent alleles in response to environmental stresses suggests that individual alleles may be under different regulatory controls. Generally, variations in allelic expression have been attributed to differences in non-coding DNA sequences and epigenetic regulation25. Notably, nucleotide sequence analysis of gene promoter and coding regions of all diverged allelic pairs revealed a significantly lower (P = 1.0−23) sequence identity in promoter regions (Extended Data Fig. 4 and Supplementary Information 15), which suggests functional diversity in allelic promoter regions26.
To test whether the divergent alleles may be the consequence of adaptive evolution to distinct environmental conditions, we divided the allelic pairs into seven subsets according to their ratio of non-synonymous to synonymous nucleotide substitutions (dN/dS) (Fig. 3a, b). Alleles with an elevated rate of non-synonymous mutations showed a significantly higher maximum fold change in bi-allelic expression during RNA-seq experiments (Fig. 3a; median test, adjusted P < 0.05). The highest median log2 fold-change in bi-allelic expression was 2.73—this was observed for the subset of diverged alleles with dN/dS ≥ 1, which is indicative of positive selection. The lowest median log2 fold-change was 2.01—this was observed for the subset of alleles with dN/dS 0–0.1, the smallest range (Fig. 3a). This suggests that positive selection has a role in driving the evolution of alleles with strong bi-allelic expression. However, most of the alleles with the highest dN/dS had unknown functions (Extended Data Table 3).
a, Relationship between median values for dN/dS subsets and maximum observed differential bi-allelic expression for each diverged allelic pair. The dotted line shows a second-order polynomial regression (F2,4 = 123.6; P < 0.001; R2 = 0.98) with 0.95 confidence levels. A median test (grand median = 2.25) showed significant differences (P < 0.001) between different dN/dS subsets and their associated maximum differential bi-allelic expression. Pairwise multiple comparisons (Median post-hoc test, adjusted P < 0.05) were used to identify significant differences between groups indicated by different letter codes with no significant differences between data with same letter code. b, Frequency distribution of different dN/dS subsets.
If allelic divergence is important for adaptation to a fluctuating environment, one might predict that recombination would be suppressed. We therefore examined the effect of recombination and genetic drift on the allelic variation, studying a natural population of F. cylindrus from Southern Ocean sea ice (Fig. 1b). We analysed around 200 high-quality Sanger sequences from alleles of two genes, the ferrichrome ABC transporter (Joint Genome Institute (JGI) protein ID 240308) and large ribosomal protein L10 (JGI protein ID 267462). Recombination analysis identified various intragenic recombinant alleles consistent with reticulate evolution (Fig. 4a, b and Supplementary Information 16). We then analysed the phylogenetic networks of these alleles and compared the branch lengths and the number of splits to networks of simulated populations. In addition, we compared the alleles of 645 genes to homologous alleles from mate-pairs of the temperate diatom Pseudo-nitzschia multistriata, a closely related sexually reproducing species, showing that the alleles in F. cylindrus have an overall higher allelic diversity than those of P. multistriata (Fig. 4c and Supplementary Information 16). These analyses indicated that the extensive allelic diversity in F. cylindrus is maintained in a vast gene pool with an effective population size Ne ≈ 16.5 × 107 (assuming a base mutation rate μ = 10−9), and that the recombination rate is about five times the mutation rate (Fig. 4d, e). The observed divergence is thus not the result of genetic introgression after hybridization, but simply the consequence of a high mutation-drift parameter (Θ) in conjunction with positive selection. Furthermore, alleles in the genome of F. cylindrus appear to coalesce shortly after the onset of the last glacial period, which began about 110,000 years ago27 (Extended Data Fig. 5). Thus, our studies suggest that the diversification of alleles took place only recently and is maintained in the vast gene pool of the diatom, which allows it to thrive under the highly variable environmental conditions of the Southern Ocean2,3,5,7,13.
a, b, Unrooted phylogenetic networks of a ferrichrome ABC Transporter (a, JGI Protein ID 240308) and the large ribosomal protein L10 (b, Protein ID 267462) from natural F. cylindrus populations with an average branch length close to 10−2, and containing approximately 225 splits. c, Comparison of Ks (number of synonymous mutations per synonymous site) between 645 orthologues of F. cylindrus (Fc) and P. multistriata (Pm). d, Population genetic simulations across a range of θ values (the population mutation parameter, θ = 4Neμ) show that the branch lengths of the networks (a, b) are most consistent with Θ ≈ 0.066, which equates to an effective population size Ne ≈ 16.5 × 107, assuming a base mutation rate μ = 10−9. e, Simulations furthermore indicate that, based on the number of splits in the networks (a, b), the recombination rate is between 1 and 10 times the mutation rate (R/μ ≈ 5).
Methods
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment.
Culture strain and DNA preparation
F. cylindrus (Grunow) Krieger (strain accession CCMP1102) was obtained from the National Centre for Marine Algae and Microbiota. Bacterial contaminants were removed by treatment with ampicillin (50 μg ml−1) and chloramphenicol (1 μg ml−1) and cultures were single cell sorted using flow cytometry. High-molecular-weight DNA for whole-genome sequencing was extracted from an axenic and monoclonal culture as previously described28 with minor modifications (see Supplementary Information 1); genome size was estimated using quantitative real-time PCR as previously described29.
Sanger sequencing
All sequencing reads were collected using standard Sanger sequencing protocols on ABI 3730XL capillary sequencing machines at the US Department of Energy Joint Genome Institute. Three different sized libraries were used as templates for the plasmid subclone sequencing process and both ends were sequenced. We obtained 270,371 reads from the 2.5-kb library, 319,392 reads from the 6.3-kb library, and 81,408 reads from a 35.4-kb fosmid library.
PacBio sequencing
All sequencing reads were collected using standard PacBio single-molecule real-time (SMRT) sequencing protocols. Two different-sized libraries were created and sequenced on a PacBio RSII instrument using the sixth generation of polymerase and the fourth generation of chemistry (P6-C4 chemistry). A 20-kb fragment length library was sequenced using three SMRT cells with total yield of 1.37 Gb of raw data. Additionally, a 4-kb insert size library was sequenced using four SMRT cells with a yield of 3.85 Gb of raw data.
Sanger assembly
A total of 671,171 sequence reads (7.25× final sequence coverage) were assembled using Arachne v.20071016 (ref. 30). The final genome assembly was produced by passing the initial assembly through the Rebuilder module to merge adjacent haplotypes, followed by another complete Arachne assembly process. We obtained 4,622 contigs that were linked into 286 scaffolds, including 105 scaffolds larger than 100 kb. To exclude organelle sequences and contaminating scaffolds from the nuclear genome assembly, each scaffold was screened against bacterial proteins and organelle sequences in the NCBI GenBank database and a set of known microbial proteins using megaBLAST and blastp searches31. Additional scaffolds were removed if they consisted of more than 95% 24-mers that occurred four other times in the scaffolds larger than 50 kb or if the scaffold contained only unanchored RNA sequences. We classified additional scaffolds as one chloroplast scaffold, five mitochondrial scaffolds, two scaffolds of <1 kb length, and seven small repetitive scaffolds. The final nuclear genome assembly contains 4,602 contigs with a contig N50 (that is, the contig size above which 50% of the total length of the sequence is included) of 78.2 kb and 271 scaffolds with a scaffold N50 of 1.3 Mb. The genome completeness was assessed using the Core Eukaryotic Genes Mapping Approach (CEGMA)15. The cumulative haploid genome size was estimated at 61.2 Mb, accounting for 46 Mb genomic scaffolds that were collapsed into a single haplotype, 29.8 Mb of genomic scaffolds that could not be collapsed into a single haplotype (that is, 14.9 Mb for collapsed single haplotype; see below), and 0.3 Mb low-coverage scaffolds. This is consistent with independently estimated genome sizes of 57.9 (± 16.9) Mb and 59.7 Mb using qPCR and PacBio sequence data, respectively. Additionally, we experimentally validated the haplotype-resolved genome assembly from whole-genome shotgun Sanger sequences by sequencing a large-insert fosmid library and aligning it to genomic scaffold sequences, using the contiguity information of fosmids to directly phase ascertained collapsed (homologous) and diverged haplotypes. We then assessed nucleotide alignments between annotated protein sequences from the genome assembly scaffolds and the haplotyped Sanger-finished fosmid clones that were not included in the genome assembly. Finally, under the assumption that gene duplicates are more divergent than alleles, we compared the sequence similarity between predicted diverged alleles and duplicates. We independently validated the sequence similarity for the alleles by Sanger sequencing of diverged F. cylindrus alleles from a natural sea ice population (see ‘Evolutionary genetic analysis’ below).
PacBio assembly
PacBio data from seven SMRT cells were used, and after filtering the shortest reads, we obtained 1,971,632 reads and 3.8 Gb of data which gave 63× coverage. We used the diploid aware PacBio assembler FALCON 0.3.0 (ref. 32), which has recently been used to successfully assemble highly heterozygous genomes33, to assemble the F. cylindrus genome. The cut off for long reads was 2,000 bp. The FALCON assembly consisted of 745 primary contigs with a total length of 59.7 Mb. The N50 of the primary contigs was 245 kb. The assembler also produced alternate contigs, which represent two diverged haplotypes for those regions. There were 288 alternate contigs, with N50 of 42 kb and total length 9.1 Mb. We used the QUIVER algorithm to polish the PacBio assembly using parts of the smrtanalysis 2.3.0p5 (http://www.pacb.com) pipeline. We assessed the accuracy of this assembly using the Sanger finished haplotyped fosmids, which we aligned with bwa 0.7.12 (ref. 34) using the bwa mem –x pacbio command. The polished assembly was highly accurate, ranging from 99.65% to 100%, with all fosmids aligning to it. One of the fosmids aligned perfectly over 43,010 bp. Out of the remaining 13 fosmids, 8 had an accuracy of >99.9%.
Genome annotation
The F. cylindrus genome assembly was annotated using the Joint Genome Institute (JGI) annotation pipeline17. The assembly was masked for repeats using RepeatMasker35 and the RepBase library36, and the most frequent (>150 times) repeats were recognized by RepeatScout37. Protein-coding gene models were predicted using three levels of evidence: ab initio Fgenesh38; homology-based Fgenesh+38 and Genewise39 seeded by BLASTx alignments against the NCBI NR database; and transcriptome-based Fgenesh. For each genomic locus, automated filtering selected the best model based on homology and transcriptome support. tRNAs were predicted using tRNAscan-SE40. All predicted proteins were functionally annotated using SignalP41 for signal sequences, TMHMM42 for transmembrane domains, interProScan43 for integrated collection of functional and structural protein domains, and protein alignments to NCBI NR, SwissProt44, KEGG45 for metabolic pathways, and KOG46 for eukaryotic clusters of orthologues. InterPro and SwissProt hits were used to map Gene Ontology terms47. Additionally, custom analyses were performed on selected protein families.
Analysis of metal-binding protein families
Metal-binding protein families were annotated using the Structural Classification of Proteins (SCOP) database v1.75 (ref. 48), which provides a classification of protein domains published in the Protein Data Bank49 into a hierarchy including classes, folds, fold superfamilies, fold families, and domains. Metal annotations of the SCOP database were built upon those in refs 50, 51. New fold families and fold superfamilies were manually curated according to metal binding and compared to automated annotation of metal binding by SCOP fold families from the PROCOGNATE database52. We used hidden Markov models (HMMs) from the Superfamily database53,54 to annotate protein sequences according to structural composition. Using the Superfamily HMMs and HMMER3 we analysed the F. cylindrus genome in comparison to other phytoplankton and Phytophthora genomes from the PHYTAX database. To perform an evolutionary genetic analysis of zinc-binding MYND protein domains, nucleotide sequences were aligned using Muscle within MEGA5 (ref. 55); DNAsp v5 (ref. 56) was used to obtain measures of nucleotide diversity. We then used BEAST 1.6.2 (ref. 57) to infer tree topology and relative divergence times between sequence clusters using a HKY+G nucleotide substitution model58 under a relaxed molecular clock59 and a Yule tree prior60.
The assembly and annotation were released as a public web portal available at http://genome.jgi-psf.org/Fracy1/Fracy1.home.html.
Identification and analysis of diverged alleles
To produce a non-redundant single haplotype gene set with only one allele of each gene, we aligned the genomic assembly against itself using BLAT61 with thresholds of 95% nucleotide identity and ≥50% alignment coverage for the smaller scaffold. We obtained alignments of 210 smaller scaffolds against larger scaffolds, with a total length of 15.9 Mb and an alignment coverage over the entire length of the smaller scaffold for 74.3% of the alignments. Syntenic scaffolds that were homologous over the entire length were analysed using Mauve genome alignment software62. Gene models of the aligned smaller scaffolds, which formed the best bi-directional blastn hit pairs with corresponding genes on larger scaffolds and >90% nucleotide identity were removed to produce a final non-redundant protein-coding gene model set. We also defined the set of diverged alleles for allele-specific downstream analyses. Diverged alleles on the larger scaffolds were referred to as allele 1 set and alleles on the smaller scaffolds as allele 2 set.
Additionally, the assembly based on PacBio sequencing was used to validate the allelic divergence observed in the assembly based on Sanger sequencing. For this analysis, the 15 longest scaffolds from the PacBio assembly were used, which accounted for 21 Mb of primary sequence and 2 Mb of alternate sequence. After filtering for genes that were only deviant by ±1% from the length of the predicted protein-coding gene models, we identified 305 genes that possessed two diverged alleles and 30 genes that were classified as paralogues.
Functional Gene Ontology enrichment analysis of diverged alleles
An intra-genomic Gene Ontology enrichment analysis was performed to test whether diverged alleles were enriched for functional Gene Ontology classes in comparison to non-diverged alleles. We compared the proportion of diverged allelic pairs associated with a Gene Ontology class in the total set of Gene Ontology annotated diverged allelic pairs against the same proportion calculated for the set of non-diverged alleles using Fisher’s exact test and adjusted P values63.
Promoter analysis of diverged alleles
Putative promoter nucleotide sequences were collected by extracting good quality sequences from different regions relative to the transcription start site (TSS). The collected sequences were divided into promoters (before TSS) and transcripts (after TSS) and ClustalW64 alignments of both sets were parsed with custom scripts using Bioperl65. We calculated the average identity of the alignments and the average percentage identity in 10-bp intervals using a sliding window approach.
Environmental metatranscriptome signature of F. cylindrus
Sequences from a Southern Ocean, eukaryote-targeted metatranscriptome were quality filtered, clustered and taxonomically classified using PhymmBL66 with a custom reference database6. To identify F. cylindrus-like transcripts a BLASTx search (E value ≤ 1 × 10−10) of sequences classified as Bacillariophyta (PhymmBL confidence score ≥ 0.9) was performed against all predicted gene models in the genome assembly, including gene models for diverged alleles on scaffolds that could not be collapsed into a single haplotype. The total number of hits was then calculated for all genes, including genes present as diverged alleles. We used a functional Gene Ontology analysis to assess the abundance of metatranscriptome reads associated to diverged alleles and investigate the environmental significance of bi-allelic transcript abundance for diverged alleles. Results were visualized with REViGO67.
Transcriptome sequencing
F. cylindrus batch cultures were grown in three biological replicates under optimal growth conditions (+4 °C, nutrient-replete Aquil68, 24 h light at 35 μmol photons m−2 s−1), freezing temperatures (−2 °C), elevated temperatures (+11 °C), elevated carbon dioxide (+4 °C, 1,000 p.p.m. CO2), low iron (+4 °C, iron-depleted Aquil), and prolonged darkness (+4 °C, 7 days darkness). Total RNA was extracted using an adaptation of the acid guanidinium thiocyanate–phenol–chloroform method69. cDNA libraries for RNA-sequencing were constructed from total RNA using random hexamer primers and sequenced in a single-flow cell lane using multiplex DNA barcodes on an Illumina HiSeq 2000 instrument at Edinburgh Genomics to generate paired-end reads of 101 bases in length. A total of 68,832,506 RNA-seq reads were aligned to the F. cylindrus genome assembly using GSNAP70, and HTSeq71 was used to count unique fragments mapping in each genomic feature.
Differential expression analysis
Data were analysed using edgeR72 for differential expression and goseq73 for functional Gene Ontology analysis. Results were visualized using REViGO67 and graphical packages of R statistical software74. Statistical testing for genes that were differentially expressed between an experimental treatment and optimal growth reference condition was performed using the generalized linear model (glm) functionality implemented in edgeR. After estimating genewise (tagwise) dispersions and fitting negative binomial models, the glm likelihood ratio test75 was applied to test for differentially expressed genes and P values adjusted63. Testing for differentially expressed diverged alleles was performed analogously. Differential bi-allelic expression was analysed comparing the expression of diverged alleles between experimental treatment and optimal growth reference conditions, and within diverged allelic pairs for each single growth condition.
Bi-allelic expression relative to allelic divergence
To test the role of adaptive evolution in allelic divergence we investigated the relationship between the dN/dS and the degree of bi-allelic expression under all experimental conditions (maximum differential expression within diverged allelic pairs). To calculate the dN/dS ratios for diverged allelic pairs their nucleotide transcript sequences were translated into amino acids and aligned with ClustalW2 (ref. 64). Amino acid alignments were mapped back over nucleotide sequences to ensure that nucleotide sequences contained full codons and were in frame. After realignment of adjusted nucleotide sequences the dN/dS was calculated for each allelic pair using codeml (pairwise mode) within PAML76. Outlier genes showing abnormally high dN/dS > 10 were discarded for subsequent analysis. The diverged allelic pairs were divided into subsets according to their associated dN/dS ratios and differences in the maximum differential bi-allelic expression between groups were compared using nonparametric statistical testing.
Sequencing of diverged F. cylindrus alleles from a natural sea ice population
A total of 196 sequences for alleles of two genes encoding a ferrichrome ABC transporter (JGI protein ID 240308) and the large ribosomal protein L10 (JGI protein ID 267462) were amplified from environmental samples using species-specific PCR primers. Purified PCR fragments were cloned using a TOPO cloning strategy, sequenced using capillary sequencing technology (Sanger method), and manually inspected for sequence quality. The sequence divergence between the two allelic pairs was assessed and compared to the sequence divergence of all alleles and duplicates as predicted for haplotype-resolved genome assemblies based on Sanger and PacBio sequencing.
Recombination analysis
We developed a novel approach to identify triplets in the diverged allelic sequences that show evidence of homologous recombination (Supplementary Information 16). Additionally, we used the R package HybridCheck77 and pairwise homoplasy index testing78 to identify sites of recombination.
Phylogenetic network analysis
The evolutionary relationships between the diverged alleles encoding for the ferrichrome ABC transporter and the large ribosomal protein L10 was visualized in splits graphs using SplitsTree4 (ref. 79), and branch lengths and number of splits in the observed phylogenetic networks were compared with those from simulated populations using simuPOP80. Population genetic simulations assuming a base mutation rate μ = 10−9 were performed across a range of values for the population mutation parameter theta (θ = 4Neμ) and the recombination rate (R) to estimate effective population size (Ne) and recombination rates relative to the mutation rate (R/μ). To minimize the effects of selection, the substitution patterns at the third codon position were studied only. DNAsp56 and LAMARC81 were used to estimate theta from sequences of the diverged alleles.
Comparative analysis of allelic nucleotide divergence between a polar and temperate diatom
The nucleotide divergence of alleles for 645 F. cylindrus genes was compared to homologous alleles from RNA-seq transcriptomes of mate pairs of the sexually reproducing temperate diatom P. multistriata. Alleles in P. multistriata were identified using best reciprocal blastn (≥90% overall identity, ≥75% coverage of both sequences) searches of the two mate pair strains. Homologous alleles between both diatoms were identified using reciprocal tBLASTx (≥30% overall identity, ≥ 50% coverage of the query sequence) searches of the theoretical six frame translations of the sequences. The divergence of allelic pairs was calculated as described above using PAML76.
Coalescence time estimates of diverged F. cylindrus alleles
The coalescence time between the two independently sequenced diverged F. cylindrus alleles from a natural population (ferrichrome ABC transporter and large ribosomal protein L10) were estimated and compared to the coalescence time of diverged allelic pairs of the approximately 6,000 genes in the genome assembly. Coalescence time was estimated using the algorithm implemented in HybridCheck77. The coalescence time estimate returned by the algorithm is expressed in terms of generations and was converted into years using an estimated division rate of 12.5 per year for F. cylindrus.
Data Availability
The Sanger genome assembly and annotations used in this study are available via the JGI genome portal at http://genome.jgi.doe.gov/Fragilariopsis_cylindrus and the Whole Genome Shotgun Project has been deposited to DDBJ/EMBL/GenBank under accession number LFJG00000000 (BioProject PRJNA32761). ESTs have been deposited to DDBJ/EMBL/GenBank under accession number GW070125. The PacBio genome assembly has been deposited at DDBJ/EMBL/GenBank under accession number PRJEB15040. F. cylindrus RNA-seq data are available in the ArrayExpress database (http://www.ebi.ac.uk/arrayexpress) under accession number E-MTAB-5024. Metatranscriptome sequences have been deposited at the Sequence Read Archive under accession number SRR1752079. P. multistriata RNA-sequencing data has been deposited at DDBJ/EMBL/GenBank under accession number PRJNA80045 and is available at http://www.ebi.ac.uk/ena/data/view/SRS190381 (strain Sy373) and http://www.ebi.ac.uk/ena/data/view/SRS190382 (strain Sy379). All other data are available from the author upon reasonable request.
Accession codes
Primary accessions
ArrayExpress
BioProject
NCBI Reference Sequence
References
Rogers, A. D. Evolution and biodiversity of Antarctic organisms: a molecular perspective. Phil. Trans. R. Soc. B 362, 2191–2214 (2007)
Goldman, J. A. et al. Gross and net production during the spring bloom along the Western Antarctic Peninsula. New Phytol. 205, 182–191 (2015)
Strzepek, R. F. et al. Iron–light interactions differ in Southern Ocean phytoplankton. Limnol. Oceanogr. 57, 1182–1200 (2012)
Bertrand, E. M. et al. Iron limitation of a springtime bacterial and phytoplankton community in the ross sea: implications for vitamin B12 nutrition. Front. Microbiol. 2, 160 (2011)
Tagliabue, A. et al. Surface-water iron supplies in the Southern Ocean sustained by deep winter mixing. Nat. Geosci. 7, 314–320 (2014)
Toseland, A. et al. The impact of temperature on marine phytoplankton resource allocation and metabolism. Nat. Clim. Chang. 3, 979–984 (2013)
Parkinson, C. L. & Cavalieri, D. J. Antarctic sea ice variability and trends, 1979–2010. Cryosphere 6, 871–880 (2012)
Fiala, M. & Oriol, L. Light–temperature interactions on the growth of Antarctic diatoms. Polar Biol. 10, 629–636 (1990)
Kang, S.-H. & Fryxell, G. A. Fragilariopsis cylindrus (Grunow) Krieger: The most abundant diatom in water column assemblages of the Antarctic marginal ice-edge zones. Polar Biol. 12, 609–627 (1992)
von Quillfeld, C. H. The diatom Fragilariopsis cylindrus and its potential as an indicator species for cold water rather than for sea ice. Vie Milieu 54, 137–143 (2004)
Thomas, D. N. & Dieckmann, G. S. Antarctic Sea ice—a habitat for extremophiles. Science 295, 641–644 (2002)
Smetacek, V. et al. Deep carbon export from a Southern Ocean iron-fertilized diatom bloom. Nature 487, 313–319 (2012)
Wang, S. et al. Impact of sea ice on the marine iron cycle and phytoplankton productivity. Biogeosciences 11, 4713–4731 (2014)
Vancoppenolle, M. et al. Role of sea ice in global biogeochemical cycles: emerging views and challenges. Quat. Sci. Rev. 79, 207–230 (2013)
Parra, G., Bradnam, K. & Korf, I. CEGMA: a pipeline to accurately annotate core genes in eukaryotic genomes. Bioinformatics 23, 1061–1067 (2007)
Armbrust, E. V. et al. The genome of the diatom Thalassiosira pseudonana: ecology, evolution, and metabolism. Science 306, 79–86 (2004)
Bowler, C. et al. The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456, 239–244 (2008)
Alverson, A. J., Beszteri, B., Julius, M. L. & Theriot, E. C. The model marine diatom Thalassiosira pseudonana likely descended from a freshwater ancestor in the genus Cyclotella. BMC Evol. Biol. 11, 125 (2011)
De Martino, A., Meichenin, A., Shi, J., Pan, K. & Bowler, C. Genetic and phenotypic characterization of Phaeodactylum tricornutum (Bacillariophyceae) accessions. J. Phycol. 43, 992–1009 (2007)
Peers, G. & Price, N. M. Copper-containing plastocyanin used for electron transport by an oceanic diatom. Nature 441, 341–344 (2006)
Gamsjaeger, R., Liew, C. K., Loughlin, F. E., Crossley, M. & Mackay, J. P. Sticky fingers: zinc-fingers as protein-recognition motifs. Trends Biochem. Sci. 32, 63–70 (2007)
Croot, P. L., Baars, O. & Streu, P. The distribution of dissolved zinc in the Atlantic sector of the Southern Ocean. Deep Sea Res. Part II Top. Stud. Oceanogr. 58, 2707–2719 (2011)
Raymond, J. A. & Kim, H. J. Possible role of horizontal gene transfer in the colonization of sea ice by algae. PLoS One 7, e35968 (2012)
Marchetti, A. et al. Comparative metatranscriptomics identifies molecular bases for the physiological responses of phytoplankton to varying iron availability. Proc. Natl Acad. Sci. USA 109, E317–E325 (2012)
Knight, J. C. Allele-specific gene expression uncovered. Trends Genet. 20, 113–116 (2004)
Guo, M. et al. Allelic variation of gene expression in maize hybrids. Plant Cell 16, 1707–1716 (2004)
Blunier, T. & Brook, E. J. Timing of millennial-scale climate change in Antarctica and Greenland during the last glacial period. Science 291, 109–112 (2001)
Doyle, J. J. & Doyle, J. L. Isolation of plant DNA from fresh tissue. Focus 12, 13–15 (1990)
Wilhelm, J., Pingoud, A. & Hahn, M. Real-time PCR-based method for the estimation of genome sizes. Nucleic Acids Res. 31, e56 (2003)
Jaffe, D. B. et al. Whole-genome sequence assembly for mammalian genomes: Arachne 2. Genome Res. 13, 91–96 (2003)
Wheeler, D. L. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 35, D5–D12 (2007)
Pendleton, M. et al. Assembly and diploid architecture of an individual human genome via single-molecule technologies. Nat. Methods 12, 780–786 (2015)
Chin, C.-S. et al. Phased diploid genome assembly with single-molecule real-time sequencing. Nat. Methods 13, 1050–1054 (2016)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Smit, A. F., Hubley, R. & Green, P. RepeatMasker Open-3.0 (1996–2010) http://www.repeatmasker.org
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005)
Price, A. L., Jones, N. C. & Pevzner, P. A. De novo identification of repeat families in large genomes. Bioinformatics 21 (Suppl. 1), i351–i358 (2005)
Salamov, A. A. & Solovyev, V. V. Ab initio gene finding in Drosophila genomic DNA. Genome Res. 10, 516–522 (2000)
Birney, E. & Durbin, R. Using GeneWise in the Drosophila annotation experiment. Genome Res. 10, 547–548 (2000)
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997)
Nielsen, H., Engelbrecht, J., Brunak, S. & von Heijne, G. Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng. 10, 1–6 (1997)
Melén, K., Krogh, A. & von Heijne, G. Reliability measures for membrane protein topology prediction algorithms. J. Mol. Biol. 327, 735–744 (2003)
Quevillon, E. et al. InterProScan: protein domains identifier. Nucleic Acids Res. 33, W116–W120 (2005)
UniProt Consortium. Activities at the Universal Protein Resource (UniProt). Nucleic Acids Res. 42, D191–D198 (2014)
Kanehisa, M. et al. KEGG for linking genomes to life and the environment. Nucleic Acids Res. 36, D480–D484 (2007)
Koonin, E. V. et al. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 5, R7 (2004)
The Gene Ontology Consortium. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000)
Murzin, A. G., Brenner, S. E., Hubbard, T. & Chothia, C. SCOP: a structural classification of proteins database for the investigation of sequences and structures. J. Mol. Biol. 247, 536–540 (1995)
Rose, P. W. et al. The RCSB Protein Data Bank: redesigned web site and web services. Nucleic Acids Res. 39, D392–D401 (2011)
Dupont, C. L., Butcher, A., Valas, R. E., Bourne, P. E. & Caetano-Anollés, G. History of biological metal utilization inferred through phylogenomic analysis of protein structures. Proc. Natl Acad. Sci. USA 107, 10567–10572 (2010)
Dupont, C. L., Yang, S., Palenik, B. & Bourne, P. E. Modern proteomes contain putative imprints of ancient shifts in trace metal geochemistry. Proc. Natl Acad. Sci. USA 103, 17822–17827 (2006)
Bashton, M., Nobeli, I. & Thornton, J. M. PROCOGNATE: a cognate ligand domain mapping for enzymes. Nucleic Acids Res. 36, D618–D622 (2007)
Gough, J. Genomic scale sub-family assignment of protein domains. Nucleic Acids Res. 34, 3625–3633 (2006)
Gough, J., Karplus, K., Hughey, R. & Chothia, C. Assignment of homology to genome sequences using a library of hidden Markov models that represent all proteins of known structure. J. Mol. Biol. 313, 903–919 (2001)
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011)
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009)
Drummond, A. J. & Rambaut, A. BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol. Biol. 7, 214 (2007)
Hasegawa, M., Kishino, H. & Yano, T. Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J. Mol. Evol. 22, 160–174 (1985)
Drummond, A. J., Ho, S. Y. W., Phillips, M. J. & Rambaut, A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 4, e88 (2006)
Yule, G. U. A mathematical theory of evolution. Based on the conclusions of Dr. J. C. Willis, F.R.S. Phil. Trans. R. Soc. B 213, 21–87 (1925)
Kent, W. J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002)
Darling, A. E., Mau, B. & Perna, N. T. progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5, e11147 (2010)
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995)
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007)
Stajich, J. E. et al. The Bioperl toolkit: Perl modules for the life sciences. Genome Res. 12, 1611–1618 (2002)
Brady, A. & Salzberg, S. L. Phymm and PhymmBL: metagenomic phylogenetic classification with interpolated Markov models. Nat. Methods 6, 673–676 (2009)
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One 6, e21800 (2011)
Price, N. M. et al. Preparation and chemistry of the artificial algal culture medium Aquil. Biol. Oceanogr. 6, 443–461 (1988/89)
Chomczynski, P. & Sacchi, N. The single-step method of RNA isolation by acid guanidinium thiocyanate–phenol–chloroform extraction: twenty-something years on. Nat. Protocols 1, 581–585 (2006)
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010)
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015)
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol. 11, R14 (2010)
R Development Core Team. R: A language and environment for statistical computing (2015) http://www.R-project.org
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012)
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol. 24, 1586–1591 (2007)
Ward, B. J. & van Oosterhout, C. HYBRIDCHECK: software for the rapid detection, visualization and dating of recombinant regions in genome sequence data. Mol. Ecol. Resour. 16, 534–539 (2016)
Bruen, T. C., Philippe, H. & Bryant, D. A simple and robust statistical test for detecting the presence of recombination. Genetics 172, 2665–2681 (2006)
Huson, D. H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006)
Peng, B. & Kimmel, M. simuPOP: a forward-time population genetics simulation environment. Bioinformatics 21, 3686–3687 (2005)
Kuhner, M. K. LAMARC 2.0: maximum likelihood and Bayesian estimation of population parameters. Bioinformatics 22, 768–770 (2006)
Acknowledgements
We thank A. Stecher and K. Schmidt for extracting and providing environmental DNA samples and the Natural Environment Research Council UK (NERC) Biomolecular Analysis Facility (NBAF) for conducting transcriptome sequencing and providing bioinformatics support. C.B. acknowledges funding from the ERC Advanced Grant ERC-2011-ADG (Diatomite). The work conducted by the U.S. Department of Energy Joint Genome Institute, a DOE Office of Science User Facility, was supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC02-05CH11231. PacBio sequencing and library construction was delivered via the BBSRC National Capability in Genomics (BB/J010375/1) at the Earlham Institute (formerly The Genome Analysis Centre, Norwich), by members of the Platforms and Pipelines Group, PacBio assembly and sequence analysis was strategically funded by the BBSRC, Institute Strategic Programme Grant (BB/J004669/1). Additional funding for this work was provided by NERC under grants NE/I001751/1, NE/K004530/1, MGF (NBAF) grant 197, The Royal Society grant RG090774 and the Earth & Life Systems Alliance in Norwich.
Author information
Authors and Affiliations
Contributions
T.M. conceived and coordinated the project. T.M. extracted DNA and J.St. performed genome size estimation with qRT–PCR. R.P.O. and A.S. performed large-scale genome annotation, identification of diverged alleles. J.St. peformed transcriptome profiling and gene annotations. M.M., P.P., J.Sc., A.S., R.S., A.T. and B.J.W. contributed equally with data on Sanger genome sequencing, assembly and annotation (E.A.L., J.Sc., A.S.), population genomics (M.M., P.P., B.J.W.), promoter analysis (R.S.) and metatranscriptomics (A.T., V.M.). A.E.A., C.L.D., S.F., F.M., A.V. and T.W. also contributed equally with large-scale analyses on gene families (A.E.A., C.L.D., S.F., A.V.), repeats (F.M., F.L., C.M.) and population genetics (T.W.). K.W.B and I.V.G. coordinated work at the DOE Joint Genome Institute, including Sanger sequencing. P.P. coordinated and performed PacBio sequencing and assembly at the Earlham Institute including identification of alleles and paralogues with contributions from C.v.O., T.M., M.M. and M.D.C. C.v.O. coordinated the evolutionary genetic analyses. M.I.F. and R.S. built and gave access to Pseudo-nitzschia multistriata transcriptome data. Other authors (A.F., A.E.F., G.G., A.G., R.H., M.G.J., P.G.K., B.R.L., J.M., C.M., M.P., H.Q., J.A.R., C.U., R.E.V., K.U.V., C.B., E.V.A., B.R.G., A.Z.W.) contributed as members of the Fragilariopsis genome sequencing consortium by annotation of individual genes and gene families. T.M., C.v.O. and J.St. wrote and revised the paper. All named authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
M.D.C. owns shares in Pacific Biosciences of California.
Additional information
Reviewer Information Nature thanks P. Boyd, M. Montresor, P. Wincker and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Comparison of sequence similarities of diverged alleles and paralogous genes.
Notched box plots showing sequence similarity (in percentage of shared nucleotides) of 5,430 diverged allelic pairs and 2,426 paralogous genes identified in the F. cylindrus ARACHNE genome assembly (Sanger), and 305 diverged allelic pairs and 30 paralogous genes identified in the F. cylindrus FALCON assembly. The sequence divergence between alleles is significantly smaller than the divergence between paralogous genes (Mann–Whitney U-test, ***P < 10−4).
Extended Data Figure 2 Light-harvesting proteins across eukaryotic algal genomes.
Bar and line chart showing total numbers of annotated Chlorophyll a/c light-harvesting complex (LHC) domains and Lhcx proteins involved in stress response are shown. Genomes are arranged according to genome size.
Extended Data Figure 3 Expression of genes involved in mitochondrial and peroxisomal fatty acid β-oxidation in F. cylindrus.
Metabolic pathway showing expression values in fragments per kilobase of transcript per million mapped reads from six transcriptome sequencing experiments for annotated genes encoding isoenzymes (red rectangles) and associated JGI protein IDs. Colour scale represents expression values on a relative scale per gene. The direct oxidation of acyl-CoA using oxygen takes place in peroxisomes while the FAD-dependent oxidation takes places in mitochondria. ACSL, long-chain acyl-CoA synthetase; ACOX, acyl-CoA oxidase; ACD, acyl-CoA dehydrogenase; ECH, enoyl-CoA hydratase; HADH, 3-hydroxyacyl-CoA dehydrogenase; ACAT, acetyl-CoA acetyltransferase.
Extended Data Figure 4 Promoter analysis for diverged allelic pairs.
a, Average identity of allelic pairs in 10-bp windows in the interval from −1,000 to +500 bp, with respect to the transcription start sites (TSSs). The chart shows two regular trends of conservation linked by a small decrease close to the TSS. On average, the promoter regions are less conserved than the transcribed ones. b, Box plots showing the distributions of percentage identities between allelic pairs in 500-bp intervals built around the TSSs. The chart clearly shows that the transcribed regions are significantly more conserved than the promoters.
Extended Data Figure 5 Coalescence time estimates of diverged alleles.
Density graph of coalescence time estimates of alleles of two Sanger sequenced genes (gene encoding the ferrichrome ABC transporter in red and that encoding the large ribosomal subunit L10 in green) and the coalescence time of diverged allelic pairs identified in the genome sequence (blue).
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Discussions, Supplementary Figures 1-40, Supplementary Tables 1-14 and one Supplementary Note. The Supplementary Information describes full details of data processing and analyses. (PDF 7122 kb)
Supplementary Data
This zipped file contains Supplementary Data 1-6 and a Supplementary Data guide. (ZIP 10513 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International (CC BY 4.0) licence. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons licence, users will need to obtain permission from the licence holder to reproduce the material. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mock, T., Otillar, R., Strauss, J. et al. Evolutionary genomics of the cold-adapted diatom Fragilariopsis cylindrus. Nature 541, 536–540 (2017). https://doi.org/10.1038/nature20803
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature20803
This article is cited by
-
Plastid-localized xanthorhodopsin increases diatom biomass and ecosystem productivity in iron-limited surface oceans
Nature Microbiology (2023)
-
Genome analysis of Parmales, the sister group of diatoms, reveals the evolutionary specialization of diatoms from phago-mixotrophs to photoautotrophs
Communications Biology (2023)
-
MarFERReT, an open-source, version-controlled reference library of marine microbial eukaryote functional genes
Scientific Data (2023)
-
Iron-limitation light switch
Nature Microbiology (2023)
-
Gene duplication and functional divergence of new genes contributed to the polar acclimation of Antarctic green algae
Marine Life Science & Technology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.