Abstract
Medulloblastoma is a highly malignant paediatric brain tumour, often inflicting devastating consequences on the developing child. Genomic studies have revealed four distinct molecular subgroups with divergent biology and clinical behaviour. An understanding of the regulatory circuitry governing the transcriptional landscapes of medulloblastoma subgroups, and how this relates to their respective developmental origins, is lacking. Here, using H3K27ac and BRD4 chromatin immunoprecipitation followed by sequencing (ChIP-seq) coupled with tissue-matched DNA methylation and transcriptome data, we describe the active cis-regulatory landscape across 28 primary medulloblastoma specimens. Analysis of differentially regulated enhancers and super-enhancers reinforced inter-subgroup heterogeneity and revealed novel, clinically relevant insights into medulloblastoma biology. Computational reconstruction of core regulatory circuitry identified a master set of transcription factors, validated by ChIP-seq, that is responsible for subgroup divergence, and implicates candidate cells of origin for Group 4. Our integrated analysis of enhancer elements in a large series of primary tumour samples reveals insights into cis-regulatory architecture, unrecognized dependencies, and cellular origins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
Data deposits
Short-read sequencing data have been deposited at the European Genome-Phenome Archive (EGA, http://www.ebi.ac.uk/ega/) hosted by the EBI, under accession number EGAS00001000215. The data generated in this study are also available at the St Jude PeCan Data Portal (https://pecan.stjude.org/dataset/northcott).
References
Northcott, P. A., Korshunov, A., Pfister, S. M. & Taylor, M. D. The clinical implications of medulloblastoma subgroups. Nat. Rev. Neurol. 8, 340–351 (2012)
Jones, D. T. et al. Dissecting the genomic complexity underlying medulloblastoma. Nature 488, 100–105 (2012)
Northcott, P. A. et al. Enhancer hijacking activates GFI1 family oncogenes in medulloblastoma. Nature 511, 428–434 (2014)
Northcott, P. A. et al. Subgroup-specific structural variation across 1,000 medulloblastoma genomes. Nature 488, 49–56 (2012)
Northcott, P. A. et al. Medulloblastomics: the end of the beginning. Nature Rev. Cancer 12, 818–834 (2012)
Hovestadt, V. et al. Decoding the regulatory landscape of medulloblastoma using DNA methylation sequencing. Nature 510, 537–541 (2014)
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nature Rev. Genet. 15, 272–286 (2014)
Consortium, E. P.; ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012)
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012)
Kundaje, A. et al.; Roadmap Epigenomics Consortium. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015)
Chapuy, B. et al. Discovery and characterization of super-enhancer-associated dependencies in diffuse large B cell lymphoma. Cancer Cell 24, 777–790 (2013)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Lovén, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013)
Kim, T. K. et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 465, 182–187 (2010)
Cho, Y. J. et al. Integrative genomic analysis of medulloblastoma identifies a molecular subgroup that drives poor clinical outcome. J. Clin. Oncol. 29, 1424–1430 (2011)
Northcott, P. A. et al. Medulloblastoma comprises four distinct molecular variants. J. Clin. Oncol. 29, 1408–1414 (2011)
Jin, F. et al. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 503, 290–294 (2013)
Pope, B. D. et al. Topologically associating domains are stable units of replication-timing regulation. Nature 515, 402–405 (2014)
Gröschel, S. et al. A single oncogenic enhancer rearrangement causes concomitant EVI1 and GATA2 deregulation in leukemia. Cell 157, 369–381 (2014)
Milde, T. et al. HD-MB03 is a novel Group 3 medulloblastoma model demonstrating sensitivity to histone deacetylase inhibitor treatment. J. Neurooncol. 110, 335–348 (2012)
Hallberg, B. & Palmer, R. H. Mechanistic insight into ALK receptor tyrosine kinase in human cancer biology. Nature Rev. Cancer 13, 685–700 (2013)
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013)
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013)
Aruga, J. et al. Mouse Zic1 is involved in cerebellar development. J. Neurosci. 18, 284–293 (1998)
Rada-Iglesias, A. et al. A unique chromatin signature uncovers early developmental enhancers in humans. Nature 470, 279–283 (2011)
Gibson, P. et al. Subtypes of medulloblastoma have distinct developmental origins. Nature 468, 1095–1099 (2010)
Schüller, U. et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh-induced medulloblastoma. Cancer Cell 14, 123–134 (2008)
Yang, Z. J. et al. Medulloblastoma can be initiated by deletion of Patched in lineage-restricted progenitors or stem cells. Cancer Cell 14, 135–145 (2008)
Fink, A. J. et al. Development of the deep cerebellar nuclei: transcription factors and cell migration from the rhombic lip. J. Neurosci. 26, 3066–3076 (2006)
Chizhikov, V. V. et al. Lmx1a regulates fates and location of cells originating from the cerebellar rhombic lip and telencephalic cortical hem. Proc. Natl Acad. Sci. USA 107, 10725–10730 (2010)
Millonig, J. H., Millen, K. J. & Hatten, M. E. The mouse Dreher gene Lmx1a controls formation of the roof plate in the vertebrate CNS. Nature 403, 764–769 (2000)
Gilbertson, R. J. Mapping cancer origins. Cell 145, 25–29 (2011)
Byrd, J. C. et al. Targeting BTK with ibrutinib in relapsed chronic lymphocytic leukemia. N. Engl. J. Med. 369, 32–42 (2013)
Hale, G. et al. Remission induction in non-Hodgkin lymphoma with reshaped human monoclonal antibody CAMPATH-1H. Lancet 332, 1394–1399 (1988)
Ramsey, S. A. et al. Genome-wide histone acetylation data improve prediction of mammalian transcription factor binding sites. Bioinformatics 26, 2071–2075 (2010)
Ni, T. T. et al. Conditional control of gene function by an invertible gene trap in zebrafish. Proc. Natl Acad. Sci. USA 109, 15389–15394 (2012)
Robinson, G. et al. Novel mutations target distinct subgroups of medulloblastoma. Nature 488, 43–48 (2012)
Haldipur, P. et al. Expression of Sonic hedgehog during cell proliferation in the human cerebellum. Stem Cells Dev. 21, 1059–1068 (2012)
Hovestadt, V. et al. Robust molecular subgrouping and copy-number profiling of medulloblastoma from small amounts of archival tumour material using high-density DNA methylation arrays. Acta Neuropathol. 125, 913–916 (2013)
Borodina, T. & Adjaye, J. & Sultan, M. A strand-specific library preparation protocol for RNA sequencing. Methods Enzymol. 500, 79–98 (2011)
Sultan, M. et al. A simple strand-specific RNA-Seq library preparation protocol combining the Illumina TruSeq RNA and the dUTP methods. Biochem. Biophys. Res. Commun. 422, 643–646 (2012)
van de Werken, H. J. et al. 4C technology: protocols and data analysis. Methods Enzymol. 513, 89–112 (2012)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
John, S. et al. Chromatin accessibility pre-determines glucocorticoid receptor binding patterns. Nature Genet. 43, 264–268 (2011)
Romanoski, C. E., Glass, C. K., Stunnenberg, H. G., Wilson, L. & Almouzni, G. Epigenomics: Roadmap for regulation. Nature 518, 314–316 (2015)
Skipper, M. et al. Presenting the epigenome roadmap. Nature 518, 313 (2015)
Waszak, S. M. et al. Population variation and genetic control of modular chromatin architecture in humans. Cell 162, 1039–1050 (2015)
Vaquerizas, J. M., Kummerfeld, S. K., Teichmann, S. A. & Luscombe, N. M. A census of human transcription factors: function, expression and evolution. Nature Rev. Genet. 10, 252–263 (2009)
Bindea, G. et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091–1093 (2009)
Hothorn, T., Bretz, F. & Westfall, P. Simultaneous inference in general parametric models. Biom. J. 50, 346–363 (2008)
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006)
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011)
Walter, C., Schuetzmann, D., Rosenbauer, F. & Dugas, M. Basic4Cseq: an R/Bioconductor package for analyzing 4C-seq data. Bioinformatics 30, 3268–3269 (2014)
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009)
Franceschini, A. et al. STRING v9.1: protein-protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 41, D808–D815 (2013)
Acknowledgements
S.E. is a recipient of Human Frontiers Science Program long-term postdoctoral fellowship (LT000432/2014). S.M.W. received funding through a SNSF Early Postdoc Mobility Fellowship (P2ELP3_155365) and an EMBO Long-Term Fellowship (ALTF 755-2014). C.Y.L. is supported by a US Department of Defense CDMRP CA120184 postdoctoral fellowship. P.A.N. is a V Foundation V Scholar in Childhood Cancer Research. We thank Creative Science Studios (http://www.creativesciencestudios.com/) for assistance with artwork. We acknowledge J. Zhang and X. Zhou for their assistance in making this data accessible via the St. Jude PeCan Data Portal (https://pecan.stjude.org/dataset/northcott).
Author information
Authors and Affiliations
Contributions
P.A.N., J.E.B., and S.M.P. conceived and co-led the study. C.Y.L. and S.E. performed all bioinformatics related to the analysis of medulloblastoma enhancers and super-enhancers. Y.T., L.Y., D.K., B.C.W., B.J., and W.C. validated subgroup-specific enhancers in vivo. C.Y.L. and A.J.F. constructed medulloblastoma regulatory circuitry networks. M.Z., S.M.W., R.Z., D.R.P., M.S.-W., D.T.W.J., M.K., V.H., I.B., and L.C. provided informatics and general scientific support. P.H., V.V.C., and K.J.M. performed the developmental studies with dreher and WT mouse embryonic cerebella. T.R., H.-J.W., V.A., H.L., and M.-L.Y. conducted RNA-seq data generation and enhancer RNA analysis. B.A.O. performed ALK staining on medulloblastoma TMAs. L.S., P.J., and S.G. performed 4C-seq experiments. M.R. and A.K. provided medulloblastoma tissue samples. R.E., P.L., J.O.K., S.M.P., J.E.B., and P.A.N. provided institutional support and project supervision. C.Y.L., S.E., S.M.P., J.E.B., and P.A.N. prepared the figures and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Enhancer landscape of primary medulloblastoma.
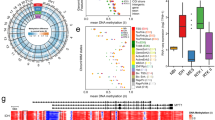
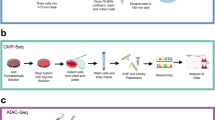
This figure accompanies Fig. 1. a, Experimental workflow for studying enhancers and super-enhancers in primary medulloblastomas. b, H3K27ac ChIP-seq data showing a highly active enhancer at the NEUROD1 locus across all 28 primary medulloblastoma samples from our series. c, Scatter plots showing Pearson correlation of H3K27ac peaks called using either sample-matched WGS or whole-cell extract (WCE) sequences as background for two samples from our series. d, Saturation analysis showing the number of discreet enhancer regions identified as a function of increasing sample number (top), or the fraction of newly gained discreet enhancer regions as a function of increasing sample number (bottom). Error bars represent 95% confidence intervals obtained from 1,000 permutations of sample order. e, Pie chart showing the genomic distribution of enhancer elements in medulloblastoma. f, Heat maps of ChIP-seq data showing the scaled read densities for H3K27ac, BRD4, H3K4me1, and H3K27me3 in regions located ± 5 kb from Group 3-specific H3K27ac (top panel) and H3K27me3 peak midpoints (bottom panel). g, Histograms showing the fractional overlap of enhancers with focal amplifications (top) or focal deletions (bottom) in Group 3 and Group 4 medulloblastoma samples. The blue distributions represent expected fractional overlap generated from 10,000 random simulations. The red line depicts the actual observed fractional overlap with empirical P-value noted. h, Scatter plot correlating average H3K27ac enrichment in Group 3 cell lines with average H3K27ac enrichment in Group 3 primary medulloblastomas. Enrichments are calculated for peaks called in primary Group 3 samples. i, Venn diagram showing the overlap between H3K27ac peaks called for primary Group 3 medulloblastomas and Group 3 medulloblastoma cell lines.
Extended Data Figure 2 Enhancer/gene assignments in medulloblastoma.
This figure accompanies Fig. 2. a, Meta H3K27ac ChIP-seq tracks of the Group 3-specific enhancers (E1 and E2) in the TAD containing ATP10A, GABRB3, and GABRA5. b, Zoom in meta H3K27ac ChIP-seq tracks of enhancer E1 from a. c–e, Scatter plots correlating sample-matched gene expression (log2 RPKM, x-axis) of ATP10A (c), GABRB3 (d), and GABRA5 (e) with H3K27ac enrichment (log2; y-axis) for the Group 3-specific enhancer shown in b. f, Zoom in meta H3K27ac ChIP-seq tracks of enhancers E2 from a. g–i, Scatter plots correlating sample-matched gene expression (log2 RPKM, x-axis) of ATP10A (g), GABRB3 (h), and GABRA5 (i) with H3K27ac enrichment (log2; y-axis) for the Group 3-specific enhancer shown in f. j, k, 4C-seq validation of TGFBR1 (j) and SMAD9 (k) enhancer/promoter interactions in a Group 3 cell line (HD-MB03).
Extended Data Figure 3 Enhancer-driven TGF-β activity in Group 3 medulloblastoma.
This figure accompanies Fig. 2. a, Functional annotation of target genes assigned to subgroup-specific enhancers based on their significant overlap with gene sets annotated in Gene Ontology (GO Biological Process) and pathway databases (KEGG, Reactome). b, Waterfall plot discriminating the top 1,000 Group 3 and Group 4 subgroup-specific enhancers as defined by total H3K27ac signal. The distribution of assigned targets in Group 3, Group 4, and shared Group 3-4 targets are shown below the waterfall. c, Convergence of Group 3-specific enhancers on TGF-β pathway genes. Subgroup-specific enhancers are summarized as nodes according to their respective medulloblastoma enhancer class—Group 3, Group 4, and shared Group 3/Group 4—with edges representing individual enhancer/TGF-β pathway gene assignments. d, Amplification of the TGF-β type II receptor, ACVR2A, in a Group 3 medulloblastoma from the ChIP-seq cohort (MB-4M23). Log2 read depth data (tumour versus matched germline) derived from WGS data for this case is shown (upper panel). Highly active H3K27ac enhancer peaks overlapping the amplified ACVR2A locus are shown for the same case (lower panel). e, Bar plot showing the difference in H3K27ac enhancer signal between MB-4M23 (ACVR2A-amplified Group 3 sample) and all other Group 3 samples. Bar plot shows H3K27ac log2 fold change at all enhancers regulating TGF-β component genes. Enhancers are ranked by increasing change in H3K27ac. Error bars represent standard error of the mean fold change.
Extended Data Figure 4 Features of medulloblastoma super-enhancers.
This figure accompanies Fig. 3. a, Unsupervised hierarchical clustering of primary medulloblastomas and cell lines using H3K27ac signal calculated at all SEs identified in each individual sample. b, Meta tracks of H3K27ac ChIP-seq signal for the ZIC1/ZIC4 SE locus. Expression (mean RPKM) for both ZIC4 (left) and ZIC1 (right) is displayed as bar graphs to the right of each H3K27ac track with error bars representing s.d. of the mean (n = 140 samples). c, Line plot showing the enhancer rank for the ZIC1/ZIC4 SE locus across all samples according to subgroup. d, Heat map showing the SE association of known medulloblastoma driver genes and chromatin modifiers. Genes with called differential SEs are shaded black, whereas genes with proximal SEs (within 100 kb of TSS) are shaded grey, according to their respective subgroup. e, Bar plot showing the number of SE regions assigned to individual enhancer classes in medulloblastoma. f, Bar plot of enhancer signal cross sample variance (y-axis) displayed as a fraction of the mean for SE enhancer constituents (left, black) or TE enhancer constituents (right, grey) identified in each medulloblastoma subgroup. g, Box plots of H3K27ac (left, blue) or BRD4 (right, red) enhancer signal at SEs or typical enhancers (TE) in their active group-specific context or in their inactive group context (for example, for SEs or TEs present in Group 3, active group context includes all Group 3 samples and inactive group context includes all other samples). Differences in the means of the distributions is quantified by a Welch’s two-tailed t-test (***P < 1 × 10−9). h, Dot plots of average H3K27ac enhancer signal in the constituents of SEs (left) or TEs (right) for enhancer constituents identified in WNT, SHH, Group 3, or Group 4 samples, respectively. Error bars represent standard deviation of the mean across all samples in a subgroup.
Extended Data Figure 5 In vivo validation of Group 3 and Group 4 medulloblastoma super-enhancers.
This figure accompanies Fig. 4. a, Summary of zebrafish reporter assays. b, Pie chart showing the fraction of all tested medulloblastoma enhancer regions that demonstrate any central nervous system localized reporter activity. c–l, Representative bright-field and fluorescence images of embryos (1 dpf) injected with individual enhancer-containing Tol2 vectors. Lateral views (60×) show GFP reporter expression in the whole body and dorsal views show GFP expression in the central nervous system (120×). White arrows indicate the locations of GFP signal. CNS, central nervous system; HB, hindbrain; MB, midbrain; CB, cerebellum; TC, telencephalon; RE, retina; OP, olfactory placode; TG, trigeminal ganglion. For each tested enhancer, meta tracks of H3K27ac ChIP-seq signal across medulloblastoma subgroups for the cloned regulatory element are shown. m, Heat map showing H3K27ac enrichment at the ±250 kb region flanking the medulloblastoma MYC SE described in Fig. 4 (SE #2; panels f, h–j) across 77 Epigenome Roadmap tissues. Each row represents a single tissue. Each column represents a region of the MYC gene desert locus. Black shaded regions indicate the presence of H3K27ac enrichment. The samples are ordered by similarity of H3K27ac enrichment pattern. Notable clusters of mesoderm (MESO.), epithelial (EPI.), blood, brain, or GI lineage derived samples are noted. The cloned enhancer reporter region described in Fig. 4 (panels f, h–j) is depicted as a vertical line and shows overlap with only 4/77 H3K27ac Epigenome Roadmap samples.
Extended Data Figure 6 Pathways regulated by super-enhancer associated transcription factors in medulloblastoma.
This figure accompanies Fig. 5. a, Functional pathways regulated by SE-associated TFs in medulloblastoma. b, Heat map of select subgroup-specific TFs showing their expression (left columns) and enhancer motif enrichment (right columns). Enhancer motif enrichment was calculated at differential enhancer elements in the respective enhancer classes.
Extended Data Figure 7 Medulloblastoma subgroup-specific transcription factors and their associated target genes.
This figure accompanies Fig. 5. a–d, Network of subgroup-specific TFs and their predicted target genes for WNT (a), SHH (b), Group 3 (c) and Group 4 (d) subgroups. Nodes represent subgroup-specific TFs. In each subgroup, node size is scaled and shaded according to the expression level of the TF and node font is scaled and shaded according to the number of inferred target genes (that is, OUT degree). TF target genes are shown in red font scaled according to the number of TFs predicted to target that gene (that is, IN degree).
Extended Data Figure 8 Super-enhancers define medulloblastoma regulatory circuitry.
This figure accompanies Fig. 5. a–d, Scatter plots of IN (x-axis) and OUT (y-axis) regulatory degree for SE-associated TFs in each medulloblastoma subgroup. e–h, TF interaction networks for each medulloblastoma subgroup. Nodes represent the top 50% of SE-associated TFs in each subgroup as ranked by total degree (counter clockwise). Each node is coloured by total degree and predicted binding interactions with other TF SEs are shown as edges. For Group 3 and Group 4 networks, edges validated by TF ChIP-seq binding are coloured. i–l, Position weight matrices showing the top statistically enriched motif identified for each transcription factor at the top 10,000 bound enhancers in each subgroup. m, Pie charts showing the fraction of predicted edges in each Group 3 and Group 4 TF networks that are validated by the presence of the respective TF ChIP-seq binding at the enhancer. n, Medulloblastoma subgroup distribution of shared, co-bound peaks for master regulatory TFs analysed by ChIP-seq. TF binding is quantified as area under curve per peak (AUC/peak) in units of rpm. Differences in the means of the distributions is quantified by a Welch’s two-tailed t-test (N.S. P > 0.1, **P < 1 × 10−6). o, Box plot of protein–protein interaction frequency (y-axis) calculated from STRING database for pairs of SE-associated TFs showing patterns of subgroup-specific SE co-regulation (left) or randomized pairs (right).
Extended Data Figure 9 LMX1A, EOMES, and LHX2 are master transcriptional regulators of Group 4 medulloblastoma.
This figure accompanies Fig. 5. a, Subgroup-specific regulatory circuitry. Nodes are TFs associated with an SE in a subgroup-specific context. Edges indicate co-regulating TFs as defined by enrichment of TF binding motifs in respective regulatory regions. Edges validated by TF ChIP-seq are coloured according to their respective subgroup association. b, Network involving LHX2, LMX1A, and EOMES TFs and target genes inferred based on the presence of the respective TF motifs in Group 4-specific enhancers. Target genes are coloured according to their validation status based on LMX1A and LHX2 ChIP-seq, with genes arranged in the centre of the network inferred to be targeted by all three master TFs. For visualization purposes, these common targets are displayed with a larger font size compared to the genes in the surrounding network.
Supplementary information
Supplementary Table 1
List of primary medulloblastoma samples included in the study, including their subgroup affiliation, patient demographic and clinical information, and availability of supporting NGS data. (XLSX 14 kb)
Supplementary Table 2
Genomic coordinates of common and differentially-regulated enhancers identified in medulloblastoma. (XLSX 1530 kb)
Supplementary Table 3
Target gene assignments of differentially-regulated enhancers and their overlap with SEs, cancer gene and drug target databases, and published medulloblastoma signature genes. (XLSX 1353 kb)
Supplementary Table 4
Genomic coordinates of medulloblastoma SEs. (XLSX 298 kb)
Supplementary Table 5
SE-associated target genes and their overlap with multiple databases. (XLSX 49 kb)
Supplementary Table 6
Genomic coordinates of all medulloblastoma SEs and their subgroup-specificity. (XLSX 1139 kb)
Rights and permissions
About this article
Cite this article
Lin, C., Erkek, S., Tong, Y. et al. Active medulloblastoma enhancers reveal subgroup-specific cellular origins. Nature 530, 57–62 (2016). https://doi.org/10.1038/nature16546
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature16546
This article is cited by
-
FLI1 and FRA1 transcription factors drive the transcriptional regulatory networks characterizing muscle invasive bladder cancer
Communications Biology (2023)
-
Super-enhancers and the super-enhancer reader BRD4: tumorigenic factors and therapeutic targets
Cell Death Discovery (2023)
-
Superenhancers as master gene regulators and novel therapeutic targets in brain tumors
Experimental & Molecular Medicine (2023)
-
Epigenomic charting and functional annotation of risk loci in renal cell carcinoma
Nature Communications (2023)
-
A neurodevelopmental epigenetic programme mediated by SMARCD3–DAB1–Reelin signalling is hijacked to promote medulloblastoma metastasis
Nature Cell Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.