Abstract
Ever since Stephen Paget’s 1889 hypothesis, metastatic organotropism has remained one of cancer’s greatest mysteries. Here we demonstrate that exosomes from mouse and human lung-, liver- and brain-tropic tumour cells fuse preferentially with resident cells at their predicted destination, namely lung fibroblasts and epithelial cells, liver Kupffer cells and brain endothelial cells. We show that tumour-derived exosomes uptaken by organ-specific cells prepare the pre-metastatic niche. Treatment with exosomes from lung-tropic models redirected the metastasis of bone-tropic tumour cells. Exosome proteomics revealed distinct integrin expression patterns, in which the exosomal integrins α6β4 and α6β1 were associated with lung metastasis, while exosomal integrin αvβ5 was linked to liver metastasis. Targeting the integrins α6β4 and αvβ5 decreased exosome uptake, as well as lung and liver metastasis, respectively. We demonstrate that exosome integrin uptake by resident cells activates Src phosphorylation and pro-inflammatory S100 gene expression. Finally, our clinical data indicate that exosomal integrins could be used to predict organ-specific metastasis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Gene Expression Omnibus
Data deposits
The raw data for quantitative mass spectrometry analysis of lung-tropic (4173 and 4175), liver-tropic (HPAF-II and HCT116) and brain-tropic (831 and 231BR) exosomes (Fig. 2a and Extended Data Fig. 3a) are available at http://dx.doi.org/10.6084/m9.figshare.1569781. The raw sequencing data for human Kupffer cells treated in vitro with BxPC-3 or BxPC-3 ITGβ5KD exosomes have been deposited in the Gene Expression Omnibus (GEO) under accession number GSE68919.
References
Paget, S. The distribution of secondary growths in cancer of the breast. 1889. Cancer Metastasis Rev. 8, 98–101 (1989)
Hart, I. R. & Fidler, I. J. Role of organ selectivity in the determination of metastatic patterns of B16 melanoma. Cancer Res. 40, 2281–2287 (1980)
Müller, A. et al. Involvement of chemokine receptors in breast cancer metastasis. Nature 410, 50–56 (2001)
Weilbaecher, K. N., Guise, T. A. & McCauley, L. K. Cancer to bone: a fatal attraction. Nature Rev. Cancer 11, 411–425 (2011)
Zhou, W. et al. Cancer-secreted miR-105 destroys vascular endothelial barriers to promote metastasis. Cancer Cell 25, 501–515 (2014)
Chang, Q. et al. The IL-6/JAK/Stat3 feed-forward loop drives tumorigenesis and metastasis. Neoplasia 15, 848–862 (2013)
Lu, X. & Kang, Y. Organotropism of breast cancer metastasis. J. Mammary Gland Biol. Neoplasia 12, 153–162 (2007)
Cox, T. R. et al. The hypoxic cancer secretome induces pre-metastatic bone lesions through lysyl oxidase. Nature 522, 106–110 (2015)
Kaplan, R. N. et al. VEGFR1-positive haematopoietic bone marrow progenitors initiate the pre-metastatic niche. Nature 438, 820–827 (2005)
Hiratsuka, S. et al. MMP9 induction by vascular endothelial growth factor receptor-1 is involved in lung-specific metastasis. Cancer Cell 2, 289–300 (2002)
Peinado, H. et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nature Med. 18, 883–891 (2012)
Balaj, L. et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nature Commun. 2, 180 (2011)
Skog, J. et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nature Cell Biol. 10, 1470–1476 (2008)
Théry, C., Ostrowski, M. & Segura, E. Membrane vesicles as conveyors of immune responses. Nature Rev. Immunol. 9, 581–593 (2009)
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell Biol. 200, 373–383 (2013)
Peinado, H., Lavotshkin, S. & Lyden, D. The secreted factors responsible for pre-metastatic niche formation: old sayings and new thoughts. Semin. Cancer Biol. 21, 139–146 (2011)
Choi, D. S., Kim, D. K., Kim, Y. K. & Gho, Y. S. Proteomics, transcriptomics and lipidomics of exosomes and ectosomes. Proteomics 13, 1554–1571 (2013)
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nature Cell Biol. 9, 654–659 (2007)
Thakur, B. K. et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res. 24, 766–769 (2014)
Costa-Silva, B. et al. Pancreatic cancer exosomes initiate pre-metastatic niche formation in the liver. Nature Cell Biol. 17, 816–826 (2015)
Kang, Y. et al. A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 3, 537–549 (2003)
Gupta, G. P. et al. Identifying site-specific metastasis genes and functions. Cold Spring Harb. Symp. Quant. Biol. 70, 149–158 (2005)
Minn, A. J. et al. Genes that mediate breast cancer metastasis to lung. Nature 436, 518–524 (2005)
Bos, P. D. et al. Genes that mediate breast cancer metastasis to the brain. Nature 459, 1005–1009 (2009)
Desgrosellier, J. S. & Cheresh, D. A. Integrins in cancer: biological implications and therapeutic opportunities. Nature Rev. Cancer 10, 9–22 (2010)
Sroka, T. C., Marik, J., Pennington, M. E., Lam, K. S. & Cress, A. E. The minimum element of a synthetic peptide required to block prostate tumor cell migration. Cancer Biol. Ther. 5, 1556–1562 (2006)
Ruoslahti, E. & Pierschbacher, M. D. Arg-Gly-Asp: a versatile cell recognition signal. Cell 44, 517–518 (1986)
Grum-Schwensen, B. et al. Suppression of tumor development and metastasis formation in mice lacking the S100A4(mts1) gene. Cancer Res. 65, 3772–3780 (2005)
Lukanidin, E. & Sleeman, J. P. Building the niche: the role of the S100 proteins in metastatic growth. Semin. Cancer Biol. 22, 216–225 (2012)
Kim, T. H., Kim, H. I., Soung, Y. H., Shaw, L. A. & Chung, J. Integrin (α6β4) signals through Src to increase expression of S100A4, a metastasis-promoting factor: implications for cancer cell invasion. Mol. Cancer Res. 7, 1605–1612 (2009)
Abdel-Ghany, M., Cheng, H. C., Elble, R. C. & Pauli, B. U. Focal adhesion kinase activated by β4 integrin ligation to mCLCA1 mediates early metastatic growth. J. Biol. Chem. 277, 34391–34400 (2002)
Mainiero, F. et al. p38 MAPK is a critical regulator of the constitutive and the beta4 integrin-regulated expression of IL-6 in human normal thymic epithelial cells. Eur. J. Immunol. 33, 3038–3048 (2003)
Weaver, V. M. et al. β4 integrin-dependent formation of polarized three-dimensional architecture confers resistance to apoptosis in normal and malignant mammary epithelium. Cancer Cell 2, 205–216 (2002)
Nikolopoulos, S. N. et al. Targeted deletion of the integrin beta4 signaling domain suppresses laminin-5-dependent nuclear entry of mitogen-activated protein kinases and NF-κB, causing defects in epidermal growth and migration. Mol. Cell. Biol. 25, 6090–6102 (2005)
Minn, A. J. et al. Distinct organ-specific metastatic potential of individual breast cancer cells and primary tumors. J. Clin. Invest. 115, 44–55 (2005)
Oskarsson, T. et al. Breast cancer cells produce tenascin C as a metastatic niche component to colonize the lungs. Nature Med. 17, 867–874 (2011)
Fukuda, K. et al. Periostin is a key niche component for wound metastasis of melanoma. PLoS ONE 10, e0129704 (2015)
Radisky, D., Muschler, J. & Bissell, M. J. Order and disorder: the role of extracellular matrix in epithelial cancer. Cancer Invest. 20, 139–153 (2002)
Weaver, V. M. et al. Reversion of the malignant phenotype of human breast cells in three-dimensional culture and in vivo by integrin blocking antibodies. J. Cell Biol. 137, 231–245 (1997)
Grum-Schwensen, B. et al. Lung metastasis fails in MMTV-PyMT oncomice lacking S100A4 due to a T-cell deficiency in primary tumors. Cancer Res. 70, 936–947 (2010)
Chen, M., Sinha, M., Luxon, B. A., Bresnick, A. R. & O’Connor, K. L. Integrin α6β4 controls the expression of genes associated with cell motility, invasion, and metastasis, including S100A4/metastasin. J. Biol. Chem. 284, 1484–1494 (2009)
Bäuerle, T. et al. Cilengitide inhibits progression of experimental breast cancer bone metastases as imaged noninvasively using VCT, MRI and DCE-MRI in a longitudinal in vivo study. Int. J. Cancer 128, 2453–2462 (2011)
Wu, Y. J. et al. Targeting αV-integrins decreased metastasis and increased survival in a nude rat breast cancer brain metastasis model. J. Neurooncol. 110, 27–36 (2012)
Zhao, Y. et al. Tumor αvβ3 integrin is a therapeutic target for breast cancer bone metastases. Cancer Res. 67, 5821–5830 (2007)
Tominaga, N. et al. Brain metastatic cancer cells release microRNA-181c-containing extracellular vesicles capable of destructing blood-brain barrier. Nature Commun. 6, 6716 (2015)
Peinado, H. et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nature Med. 18, 883–891 (2012)
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003)
Bunkenborg, J., Garcia, G. E., Paz, M. I., Andersen, J. S. & Molina, H. The minotaur proteome: avoiding cross-species identifications deriving from bovine serum in cell culture models. Proteomics 10, 3040–3044 (2010)
Käll, L., Canterbury, J. D., Weston, J., Noble, W. S. & Mac Coss, M. J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nature Methods 4, 923–925 (2007)
Silva, J. C., Gorenstein, M. V., Li, G. Z., Vissers, J. P. & Geromanos, S. J. Absolute quantification of proteins by LCMSE: a virtue of parallel MS acquisition. Mol. Cell. Proteomics 5, 144–156 (2006)
Cox, J. et al. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol. Cell. Proteomics 13, 2513–2526 (2014)
Acknowledgements
We thank S. Rudchenko at the Hospital for Special Surgery Flow Cytometry Core Facility. We acknowledge the MSK Cancer Center Support Grant/Core Grant (P30 CA008748). Our work is supported by grants from National Cancer Institute (U01-CA169538, D.L. and M.S.B.), National Institutes of Health (R01-CA169416, D.L. and H.P.), United States Department of Defense (W81XWH-13-10249, D.L.), W81XWH-13-1-0425 (D.L., J.B., B.A.G. and Y.K.), Melanoma Research Alliance (H.P.), Sohn Conference Foundation (H.P. and H.Z.), the Children’s Cancer and Blood Foundation (H.P. and D.L.), The Manning Foundation (D.L.), The Hartwell Foundation (D.L.), Fundação para a Ciência e a Técnologia (D.L.), The Nancy C. and Daniel P. Paduano Foundation (H.P. and D.L.), The Feldstein Foundation (H.P.), The Starr Cancer Consortium (H.P. and D.L.), The Mary Kay Foundation (D.L.), Pediatric Oncology Experimental Therapeutic Investigator Consortium (POETIC, D.L. and H.P.), James Paduano Foundation (D.L. and H.P.), Beth Tortolani Foundation (D.L. and J.B.), Malcolm Hewitt Weiner Foundation (D.L.), Theodore A. Rapp Foundation (D.L.), American Hellenic Educational Progressive Association 5th District Cancer Research Foundation (D.L., A.H.), Charles and Marjorie Holloway Foundation (J.B.), Sussman Family Fund (J.B.), Lerner Foundation (J.B.), Breast Cancer Alliance (J.B.), Manhasset Women’s Coalition Against Breast Cancer (J.B.), Ministry of Science and Technology Taiwan (101-2918-I-002-016, T.-L.S.), The JSPS Postdoctoral Fellowships for Research Abroad and Susan G. Komen Postdoctoral Fellowship (A.H.).
Author information
Authors and Affiliations
Contributions
A.H. designed the experimental approach, performed the experimental work, analysed the data, coordinated the project and wrote the manuscript. B.C.-S. designed experiments investigating liver metastasis and performed the experimental work. T.-L.S. performed ECM studies. G.R. analysed brain tropic exosome distribution. A.H. performed western blot analysis. M.T.M. and H.M. performed and analysed exosome mass spectrometry. S.K. prepared overexpression vectors. S.S. and L.B. performed tissue processing and staining. S.C. designed and illustrated Fig. 4c. A.D.G., S.C., V.D.D.-C., Y.A. and C.W. received and processed human samples. N.S. and K.U. performed electron microscopy. A.E.D. performed animal surgeries and contributed to data interpretation and discussion. T.Z. performed RNA sequence analysis. B.A.G. performed initial proteomic analysis. V.K.R., G.K.S. and J.H.H. provided the uveal melanoma cell line. L.P., T.K., M.S.B., V.M., K.K., L.H.W., J.H., E.H.K., K.M., S.K.B., K.P., O.F., M.J., S.K., M.A.H., P.M.G., K.J.L., J.M.W., A.N. and W.R.J. provided and prepared human samples. H.Z., A.J.M. and P.S. read the manuscript and provided feedback. C.M.G., I.M. and H.P. discussed the hypothesis and contributed to data interpretation and wrote the manuscript. Y.K., M.d.S. and M.J.B. contributed to discussing the hypothesis, interpretation of data. J.B. coordinated the project, interpreted data and wrote the manuscript. D.L. conceived the hypothesis, led the project, interpreted the data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Characterization of organotropic exosome properties and biodistribution.
a, Human cancer exosome biodistribution in lung and liver. Exosomes (10 μg) derived from each cell line were labelled with lipophilic PKH26 dye (red) and injected retro-orbitally into nude mice 24 h before culling. Top, representative NIR whole-lung image by Odyssey imaging (n = 3). Middle and bottom, represent exosome biodistribution in the lung and liver as determined by immunofluorescence microscopy. Arrows indicate exosome foci (n = 3, three independent experiments). b, Biodistribution of exosomes isolated from mouse cell lines E0771 and Pan02. Mouse exosome biodistribution in the lung and liver was determined by immunofluorescence microscopy. Exosomes (10 μg) derived from each cell line were labelled with lipophilic PKH26 dye (red) and injected retro-orbitally into nude mice 24 h before culling. Top, lung at 40× magnification. Bottom, liver at 40× magnification. Arrows indicate exosome foci. Graph represents the quantification of exosome distribution by counting exosome-positive cells. An average of five random fields per sample were counted at 20× magnification (three independent experiments, each with n = 3). **P < 0.01 by two-tailed Student’s t-test. c, Analysis of organotropic cell-derived exosomes. MDA-MB-231 organotropic cell-line-derived exosomes were analysed for size distribution by NanoSight and phenotype (purity and shape) by electron microscopy; black arrows indicate representative exosomes. Technical triplicates were analysed, at least 10 images per sample. d, Flow cytometric analysis of exosome+ cells in lung. Exosomes (10 μg) derived from MDA-MB-231 organotropic cell lines were labelled with lipophilic PKH67 dye (green) and injected retro-orbitally into nude mice 24 h before culling. FITC-channel-positive cells were acquired on a FACS Calibur, and the percentage of exosome-positive cells was quantified (representing data pooled from two independent experiments, a total of n = 12). ***P < 0.001 by one-way ANOVA. e, Flow cytometric analysis of exosome-positive cells in the bone marrow. Exosomes (10 μm) derived from MDA-MB-231 organotropic cell lines were labelled with lipophilic PKH67 dye (green) and injected retro-orbitally into nude mice 24 h before culling. FITC-channel-positive cells were acquired on a FACS Calibur, and the percentage of exosome-positive cells was quantified (representative data pooled from two independent experiments, a total of n = 6). **P < 0.01 and *P < 0.05 by one-way ANOVA for the 831-BrT to 1833-BoT and 4175-LuT comparisons, respectively. Data are mean ± s.e.m. Scale bars, 5 mm (a, top), 50 μm (a, middle and bottom, b) and 100 nm (c).
Extended Data Figure 2 4175-LuT cell-derived exosomes localize to lung and dictate future metastatic sites.
a, Electron microscopy imaging of FM1-43-labelled 4175-LuT exosomes. Red arrows, FM1-43-positive exogenous exosomes; black arrows, endogenous exosomes. Two mice were tested, images were taken for several sections from each organ (n = 30 images in total). b, Representative NIR imaging of lung whole mount after daily exosome injections. Exosomes (10 μg) derived from 4175-LuT cells were injected daily for three consecutive days via the retro-orbital sinus and the whole lung was imaged by Odyssey imaging (n = 4). c, Representative haematoxylin/eosin staining of the lung from Fig. 1f at 20× magnification; n = 5 for all, except for LuT exo/LuT cells, in which n = 4; data representative of two independent experiments. Arrows indicate lung metastasis. d, Analysis of 1833-BoT cell metastasis to the lung, after 3 weeks of continuous treatment with PBS or 4175-LuT exosomes, followed by intracardiac injection of 1 × 105 tumour cells. Mice were injected retro-orbitally with exosomes every other day for 3 weeks, before tumour cell injection. Quantitative bioluminescence imaging of luciferase activity by IVIS imaging. Metastasis was quantified 3 weeks after tumour cell injection (n = 4). Scale bars, 100 nm (a), 5 mm (b, d) and 500 μm (c). Data are mean ± s.e.m. ***P < 0.001 by two-tailed Student’s t-test.
Extended Data Figure 3 Characterization of organotropic exosome protein cargo.
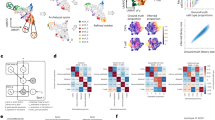
a, Top 40 adhesion molecules packaged in exosomes isolated from organ tropic cell lines. Heat map of adhesion molecule signals based on Z-scored LFQ values obtained from quantitative mass spectrometry analysis. PEP (posterior error probability), MS/MS count is a number of fragmentation spectra (spectral counting), Razor + unique peptides refers to the number of peptides, and sequence coverage refers to percentage of peptide counts identified. b, Ponceau staining of exosome lysates isolated from organ tropic cell lines. Representative Ponceau staining of total protein from the organ tropic cell-line-derived exosomes. Exosomal protein (10 μg) was loaded in each well (n = 2, three independent experiments). c, Western blot analysis comparison of ITGα6 and ITGβ4 levels in cell lysates versus exosomes derived from organotropic breast cancer and pancreatic cancer cell lines. Graph represents the relative ratios of integrin to GAPDH signals as determined by densitometry. For western blot source data, see Supplementary Fig. 1i–k.
Extended Data Figure 4 Functional characterization of organotropic exosomes.
a, Quantification of organotropic exosome uptake by target cells in vivo. Top graph, flow cytometric quantification of the frequency of 4175-LuT exosome-positive fibroblasts and epithelial cells (n = 4). Left bottom graph, flow cytometric quantification of the frequency of BxPC-3 exosome-positive macrophages (n = 3). Right bottom graph, quantification of the frequency of 831-BrT exosome-positive endothelial cells by immunofluorescence microscopy (n = 5). b, Organotropic cell-line-derived exosomes induce vascular leakiness in the lung. Leakiness in the lung 24 h after retro-orbital injection of 10 μg of normal mammary fat pad or MDA-MB-231 organotropic cell-line-derived exosomes was quantified by imaging the presence of fluorescent dextran (red) outside of blood vessels, in the lung parenchyma. Left top panel, 40× magnification of representative lung image after PBS injection. Left bottom panel, representative lung image after 4175-LuT exosome injection. Scale bar, 50 μm. Right graph depicts the quantification of five random areas at 20× magnification in arbitrary units (data representative of two independent experiments; n = 3). Data are mean ± s.e.m. *P < 0.05; **P < 0.01 by one-way ANOVA.
Extended Data Figure 5 Exosome co-localization with specific cell types within target tissues.
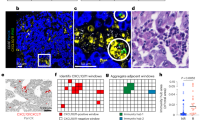
a, Immunofluorescence analysis of resident cells in lung, liver and brain after labelled exosome injection. Analysis of exosome (red) co-staining with markers (green) for tissue-specific stromal cell types. Top, representative images of immunofluorescence microscopy of 4175-LuT exosome co-staining with F4/80, CD31 and EpCAM. Middle, liver sections from mice injected with BxPC-3-LiT-derived exosomes were co-stained with CD31, S100A4 and EpCAM. Bottom, brain sections from mice injected with 831-BrT exosome were co-stained with F4/80, S100A4 and EpCAM (n = 3 per experiment for two independent experiments). b, Exosome biodistribution and co-localization with extracellular matrix proteins. Left top, representative immunofluorescence microscopy images of lung tissue, depicting 4175-LuT exosome (red) co-staining with laminin (green). Right top, laminin (green) co-staining with S100A4 (red). Left bottom, representative immunofluorescence microscopy of liver tissue co-stained for fibronectin (green) and BxPC-3-LiT exosomes (red). Right bottom, fibronectin (green) co-staining with F4/80 (red) (n = 3, two independent experiments). Scale bars, 30 μm.
Extended Data Figure 6 ITGs functionally regulate organotropic exosome uptake and exosome-mediated metastasis.
a, Representative western blot analysis of integrin expression in 4175-LuT and 4175β4KD cells and exosomes (representative of three independent experiments). For western blot source data, see Supplementary Fig. 1l. b, In vitro uptake of 4175-LuT exosomes by WI-38 lung fibroblasts. The WI-38 cell membrane was labelled with PKH67 green dye and 4175-LuT exosomes were labelled with PKH26 red dye. Exosomes (10 μg ml−1) were first incubated with PBS or HYD-1 peptide for 30 min at 37 °C, followed by 1-h incubation with WI-38 cells at 37 °C. Excess exosomes were washed and cells were imaged (n = 4 for two independent experiments). c, Representative western blot of ITGβ4 expression in exosomes isolated from wild-type or ITGβ4-overexpressing 1833-BoT cells (representative of two independent experiments). For western blot source data, see Supplementary Fig. 1m. d, Representative haematoxylin/eosin staining of lungs from Fig. 3e. Arrows indicate lung metastasis; n = 6, data representative of two independent experiments. e, Representative western blot analysis of integrin expression in BxPC-3-LiT and BxPC-3β5KD cells and exosomes. For western blot source data, see Supplementary Fig. 1n. f, Immunofluorescence analysis of BxPC-3-LiT control and BxPC-3β5KD-derived exosome biodistribution in the liver. Exosomes (10 μg) isolated from each cell line were labelled with lipophilic PKH26 dye (red) and injected retro-orbitally into nude mice 24 h before culling. Left, 40× magnification. Arrows indicate exosome foci. Scale bar, 50 μm. Right, quantification of exosome distribution by exosome-positive cells. An average of five random fields were counted at 20× magnification (data representative of two independent experiments; n = 3). g, Flow cytometry analysis of exosome-positive cells in the liver 24 h after exosome injection. Labelled BxPC-3-LiT exosomes (5 μg) per mouse were incubated with PBS, RGD, HYD-1 or ITGαvβ5 antibody for 30 min at 37 °C before retro-orbital injection into nude mice. Livers were collected and analysed for exosome-positive cells by flow cytometry 24 h after injection (n = 4, except for the ITGαvβ5 antibody group, in which n = 5). Scale bars, 10 μm (b), 500 μm (d) and 500 μm (f). **P < 0.01, ***P < 0.001 by two-tailed Student’s t-test (f) and one-way ANOVA (g). Data are mean ± s.e.m.
Extended Data Figure 7 Functional contribution of exosomes to metastasis.
a, Microscopic analysis of exosome-positive cells in the livers of mice injected with liver metastatic Pan02-LiT-derived exosomes. Before injection, Pan02-LiT exosomes were pre-incubated with RGD peptide for 30 min at 37 °C. Pan02-LiT exosomes (10 μg) were labelled with lipophilic PKH67 green dye and injected retro-orbitally into C57BL/6 mice 24 h before culling. Livers were digested and exosome-positive cells were quantified by flow cytometry (n = 3). b, Analysis of Pan02-LiT liver metastasis after 3 weeks of continuous treatment with PBS, Pan02-LiT-derived exosomes, or Pan02-LiT-derived exosomes pre-incubated with RGD peptide for 30 min at 37 °C. Pan02-LiT cells were injected intrasplenically. Mice were injected retro-orbitally with 5 μg exosome every other day for 3 weeks. Top, representative liver images showing metastasis taken at culling. Bottom, liver weight quantification (n = 4 except for the control and peptide group for which n = 3 of one experiment). c, Functional analysis of lung fibroblasts educated with 4175-LuT-derived exosomes. Proliferation of lung fibroblasts educated with exosomes every other day for 2 weeks. Three days after cells were plated at equal density, cell numbers were counted using a haemocytometer (n = 3; three independent experiments). d, Migration of lung fibroblasts educated with exosomes every other day for 2 weeks was measured as follows. Fibroblasts were plated in 24-well transwell chamber inserts, and after 6 h the number of cells that migrated was counted using haematoxylin staining. Nine random fields were counted at 20× magnification and the average number of cells per field was calculated (total of n = 4 from two independent experiments). e, Representative image of the lung stained for S100A4. Mice were treated every other day with PBS, 4175-LuT or 4175β4KD exosomes for 3 weeks. Scale bar, 50 μm; n = 4 mice. f, In situ (in-cell western) protein expression analysis of WI-38 fibroblasts treated with PBS, 4175-LuT or 4175ITGβ4KD exosomes. Relative expression levels of Src and phosphorylated (p-) Src (n = 3, three independent experiments). Data are mean ± s.e.m. *P < 0.05; **P < 0.01; ***P < 0.001 by two-tailed Student’s t-test (a, c, d) and one-way ANOVA (b, f).
Extended Data Figure 8 Exosomal integrin expression as a potential metastatic site biomarker.
a, Exosomal ITGβ4 levels in the plasma of mice bearing orthotopic 4175-LuT tumours, as a function of tumour progression. Blood plasma was collected for exosome isolation 6 weeks after intra-mammary fat pad tumour injection, then again 1 week after tumour resection, from mice that were deemed to be either free of tumour or presenting with recurring tumours based on IVIS bioluminescence imaging (n = 5 were pooled for each group, based on one experiment). For western blot source data, see Supplementary Fig. 1o. b, Exosomal ITGβ4 in healthy control subjects (Ctrl) (n = 13); patients with breast cancer (BrCa) and no metastasis (n = 3), liver metastasis (n = 1), or lung metastasis (n = 3); patients with rhabdomyosarcoma (RMS) and no metastasis (n = 1) or lung metastasis (n = 3); patients with pancreatic cancer (PDAC) with liver metastasis (n = 14) and lung metastasis (n = 3); and patients with melanoma (Mel) with lung metastasis (n = 2). c, Exosomal ITGαV in healthy control subjects (n = 13); patients with rhabdomyosarcoma and no metastasis (n = 1) or lung metastasis (n = 3); patients with breast cancer and lung metastasis (n = 3) or liver metastasis (n = 1); and patients with pancreatic cancer and liver metastasis (n = 15). Data are mean ± s.e.m. *P < 0.05; ***P < 0.001 by one-way ANOVA.
Supplementary information
Supplementary Figure
This file contains western blot source data. a-h, Analysis of ITGβ4, α6, β1, α2, β3, αV, β5 and α3 expression, respectively, in organotropic cell line-derived exosomes in Fig. 2b. i-k, Analysis of ITGβ4, ITGα6 and GAPDH expression, respectively, in cell lysates versus exosomes derived from organotropic breast cancer and pancreatic cancer cell lines in Extended Fig. 3c. l, Analysis of ITGβ4 and GAPDH expression in 4175-LuT and 4175β4KD cells and exosomes in Extended Fig. 6a. m, Analysis of ITGβ4 and GAPDH expression in exosomes isolated from WT or ITGβ4-overexpressing1833-BoT cells in Extended Fig. 6c. n, Analysis of ITGβ5 and GAPDH expression in BxPC-3 and BxPC-3β5KD cells and exosomes in Extended Fig. 6e. o, Exosomal ITGβ4 and Alix levels in the plasma of mice bearing orthotopic 4175-Lu-T tumors in Extended Fig. 8a. LC, loading control. (PDF 502 kb)
Source data
Rights and permissions
About this article
Cite this article
Hoshino, A., Costa-Silva, B., Shen, TL. et al. Tumour exosome integrins determine organotropic metastasis. Nature 527, 329–335 (2015). https://doi.org/10.1038/nature15756
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15756
This article is cited by
-
Integrinβ-1 in disorders and cancers: molecular mechanisms and therapeutic targets
Cell Communication and Signaling (2024)
-
Protein cargo in extracellular vesicles as the key mediator in the progression of cancer
Cell Communication and Signaling (2024)
-
Tumor-derived exosomal ADAM17 promotes pre-metastatic niche formation by enhancing vascular permeability in colorectal cancer
Journal of Experimental & Clinical Cancer Research (2024)
-
Versatile extracellular vesicle-mediated information transfer: intercellular synchronization of differentiation and of cellular phenotypes, and future perspectives
Inflammation and Regeneration (2024)
-
Tumor-derived small extracellular vesicles in cancer invasion and metastasis: molecular mechanisms, and clinical significance
Molecular Cancer (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.