Abstract
In female mice, two forms of X-chromosome inactivation (XCI) ensure the selective silencing of female sex chromosomes during mouse embryogenesis. Beginning at the four-cell stage, imprinted XCI (iXCI) exclusively silences the paternal X chromosome. Later, around implantation, epiblast cells of the inner cell mass that give rise to the embryo reactivate the paternal X chromosome and undergo a random form of XCI (rXCI)1,2. Xist, a long non-coding RNA crucial for both forms of XCI, is activated by the ubiquitin ligase RLIM (also known as Rnf12)3,4,5. Although RLIM is required for triggering iXCI in mice, its importance for rXCI has been controversial. Here we show that RLIM levels are downregulated in embryonic cells undergoing rXCI. Using mouse genetics we demonstrate that female cells lacking RLIM from pre-implantation stages onwards show hallmarks of XCI, including Xist clouds and H3K27me3 foci, and have full embryogenic potential. These results provide evidence that RLIM is dispensable for rXCI, indicating that in mice an RLIM-independent mechanism activates Xist in the embryo proper.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Augui, S., Nora, E. P. & Heard, E. Regulation of X-chromosome inactivation by the X-inactivation centre. Nature Rev. Genet. 12, 429–442 (2011)
Lee, J. T. Gracefully ageing at 50, X-chromosome inactivation becomes a paradigm for RNA and chromatin control. Nature Rev. Mol. Cell Biol. 12, 815–826 (2011)
Barakat, T. S. et al. RNF12 activates Xist and is essential for X chromosome inactivation. PLoS Genet. 7, e1002001 (2011)
Jonkers, I. et al. RNF12 is an X-encoded dose-dependent activator of X chromosome inactivation. Cell 139, 999–1011 (2009)
Shin, J. et al. Maternal Rnf12/RLIM is required for imprinted X-chromosome inactivation in mice. Nature 467, 977–981 (2010)
Bach, I. et al. RLIM inhibits functional activity of LIM homeodomain transcription factors via recruitment of the histone deacetylase complex. Nature Genet. 22, 394–399 (1999)
Ostendorff, H. P. et al. Ubiquitination-dependent cofactor exchange on LIM homeodomain transcription factors. Nature 416, 99–103 (2002)
Jiao, B. et al. Paternal RLIM/Rnf12 is a survival factor for milk-producing alveolar cells. Cell 149, 630–641 (2012)
Jiao, B. et al. Functional activity of RLIM/Rnf12 is regulated by phosphorylation-dependent nucleocytoplasmic shuttling. Mol. Biol. Cell 24, 3085–3096 (2013)
Gontan, C. et al. RNF12 initiates X-chromosome inactivation by targeting REX1 for degradation. Nature 485, 386–390 (2012)
Dupont, C. & Gribnau, J. Different flavors of X-chromosome inactivation in mammals. Curr. Opin. Cell Biol. 25, 314–321 (2013)
Schulz, E. G. & Heard, E. Role and control of X chromosome dosage in mammalian development. Curr. Opin. Genet. Dev. 23, 109–115 (2013)
Ostendorff, H. P. et al. Dynamic expression of LIM cofactors in the developing mouse neural tube. Dev. Dyn. 235, 786–791 (2006)
Hayashi, S., Lewis, P., Pevny, L. & McMahon, A. P. Efficient gene modulation in mouse epiblast using a Sox2Cre transgenic mouse strain. Mech. Dev. 119 (suppl. 1). S97–S101 (2002)
Hayashi, S., Tenzen, T. & McMahon, A. P. Maternal inheritance of Cre activity in a Sox2Cre deleter strain. Genesis 37, 51–53 (2003)
Avilion, A. A. et al. Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev. 17, 126–140 (2003)
Panning, B. X inactivation in mouse ES cells: histone modifications and FISH. Methods Enzymol. 376, 419–428 (2003)
Plath, K. et al. Role of histone H3 lysine 27 methylation in X inactivation. Science 300, 131–135 (2003)
Hadjantonakis, A. K., Cox, L. L., Tam, P. P. & Nagy, A. An X-linked GFP transgene reveals unexpected paternal X-chromosome activity in trophoblastic giant cells of the mouse placenta. Genesis 29, 133–140 (2001)
Tian, D., Sun, S. & Lee, J. T. The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell 143, 390–403 (2010)
Sun, S. et al. Jpx RNA activates Xist by evicting CTCF. Cell 153, 1537–1551 (2013)
Mitsui, K. et al. The homeoprotein Nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell 113, 631–642 (2003)
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007)
Zhang, L. et al. RNF12 controls embryonic stem cell fate and morphogenesis in zebrafish embryos by targeting Smad7 for degradation. Mol. Cell 46, 650–661 (2012)
Hall, L. L. et al. An ectopic human XIST gene can induce chromosome inactivation in postdifferentiation human HT-1080 cells. Proc. Natl Acad. Sci. USA 99, 8677–8682 (2002)
Griffith, G. J. et al. Yin-Yang1 is required in the mammalian oocyte for follicle expansion. Biol. Reprod. 84, 654–663 (2011)
Tursun, B. et al. The ubiquitin ligase Rnf6 regulates local LIM kinase 1 levels in axonal growth cones. Genes Dev. 19, 2307–2319 (2005)
Robertson, E. J. Teratocarcinomas and Embryonic Stem Cells—A Practical Approach Ch. 4 71–112 (IRL, 1987)
Nagy, A. et al. in Manipulating the Mouse Embryo—A Laboratory Manual 3rd edn, Ch. 8 359–397 (Cold Spring Harbor, 2003)
Acknowledgements
We are grateful to H. Ma for help in the mouse facility, M. Keeler for culturing ES cells, T. Fazzio, N. Lawson and Y. Yoon for advice, and T. Fazzio for helpful discussions and reading of the manuscript. I.B. is a member of the University of Massachusetts Diabetes-Endocrinology Research Center (DK32520). This work was supported by National Institutes of Health grants CA131158 to I.B., CA077735 to S.N.J. and GM053234 to J.B.L.
Author information
Authors and Affiliations
Contributions
J.S. and I.B. conceived and designed the experiments. M.Bo. generated the floxed Rlim mice. J.G. and S.N.J. generated the ES-cell lines and performed the tetraploid injections. M.By. and J.B.L. carried out RNA-FISH experiments and M.C.W., C.M. and J.M. performed immunohistochemistry on early embryos. All authors analysed the data. I.B. wrote the manuscript with input from J.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Strain-independent downregulation of RLIM in the mouse embryonic epiblast.
a, Test of specificity of the RLIM antibody (compare with Fig. 1b). Shown is an E4.5 Δ/Y blastocyst outgrowth stained with RLIM (red). Increasing the general signal levels (RLIM′) reveals augmented unspecific staining mainly in the cytoplasm of cells. b, Co-stainings of sections of E7–7.5 embryos within decidual tissues using antibodies directed against RLIM (red) and H3K27me3 (green). Representative C57BL/6 and SV129 embryos are shown (out of at least three that were stained). Scale bars, 75 μm.
Extended Data Figure 2 Genetic background has little or no influence on RLIM dispensability during rXCI.
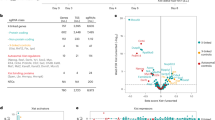
a, b, C57BL/6–SV129 (a) and C57BL/6–FVB (b) hybrid F1 parents were generated by crossings of fl/Y (C57BL/6) males with WT/WT (SV129 or FVB) females and WT/Y (SV129 or FVB) males with cKO/Δ-SC (C57BL/6) females. F1 WT/fl females and cKO/Y-SC males were then backcrossed to generate F2 cKOm offspring. Percentages of offspring (grouped into female and male) and their genotypes with respect to Rlim and SC are indicated in the abscissa and ordinate, respectively, and the total number (n) of born and genotyped F2 pups is shown. Maternally transmitted cKO alleles are indicated in black. c, No discrimination against SV129 in born cKOm/Δp pups with a mixed C57BL/6–SV129 background. Sequencing analyses of genomic DNA isolated from eight hybrid cKOm/Δp pups using 156 strain-specific SNPs distributed among all chromosomes (blue/red) or ten SNPs distributed on the X chromosome (green/orange). Note that the SV129 contribution in born cKOm/Δp pups is up to 70% (total) and 80% (on the X).
Extended Data Figure 3 Co-stainings of MEFs isolated from cKOm/Δp and WT/WT embryos using antibodies directed against H3K27me3 (green) and RLIM (red).
a, Representative images are shown. b, Summary graph of cells showing H3K27me3 foci is shown on the right. Numbers of counted cells from independent biological duplicates are 111/112 (WT/WT) and 110/104 (cKOm/Δp).
Extended Data Figure 4 Random XCI in female mice heterozygous for the Sox2-Cre-mediated deletion of Rlim.
Female mice were generated carrying either a paternal or maternal Rlim deletion and a GFP transgene on the other X chromosome (XGFP). Mating schemes to generate these females are indicated. Female littermates without the Sox2-Cre transgene were used as controls. Numbers of cells counted from independent biological duplicates are indicated.
Extended Data Figure 5 Female ES cells lacking RLIM are able to undergo rXCI in vivo.
a, Parental cross used for the generation of ES-cell lines is indicated on top. Newly generated ES-cell lines were genotyped using PCR for the presence of wild-type, floxed or knockout Rlim alleles, as well as Cre driver (SC) and Y chromosome (Zfy2). b, Newly isolated female ES cells were differentiated for 5 days in culture and stained with the H3K27me3 antibody. c, Summary of the tetraploid complementation assays showing the injected ES-cell lines, genotypes, number of deciduas and embryos obtained (see also Fig. 4c). All embryos generated via tetraploid complementation were genotyped for the presence of wild-type, floxed and knockout Rlim alleles as well as Zfy2. d, Left, MEFs isolated from an E10.5 embryo generated via tetraploid injection of Δ/Δ line IB11 ES cells were cultured for 24 h before staining with antibodies against H3K27me3. Right, summary of H3K27me3 stainings. Numbers of counted cells from independent biological duplicates are 60/62 (WT/WT) and 78/75 (cKOm/Δp). e, Summary of RNA-FISH experiments on MEFs isolated from embryos generated via tetraploid injection of Δ/Δ line IB11 ES cells (n = 109) (see Fig. 4d). MEFs isolated from WT/WT embryos (n = 106) served as control.
Rights and permissions
About this article
Cite this article
Shin, J., Wallingford, M., Gallant, J. et al. RLIM is dispensable for X-chromosome inactivation in the mouse embryonic epiblast. Nature 511, 86–89 (2014). https://doi.org/10.1038/nature13286
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature13286
This article is cited by
-
Gene regulation in time and space during X-chromosome inactivation
Nature Reviews Molecular Cell Biology (2022)
-
Activation of Xist by an evolutionarily conserved function of KDM5C demethylase
Nature Communications (2022)
-
Deletion of LBR N-terminal domains recapitulates Pelger-Huet anomaly phenotypes in mouse without disrupting X chromosome inactivation
Communications Biology (2021)
-
A novel RLIM/RNF12 variant disrupts protein stability and function to cause severe Tonne–Kalscheuer syndrome
Scientific Reports (2021)
-
A symmetric toggle switch explains the onset of random X inactivation in different mammals
Nature Structural & Molecular Biology (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.