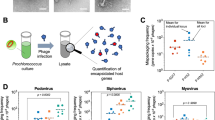
Abstract
Cyanobacteria are photosynthetic organisms responsible for ∼25% of organic carbon fixation on the Earth. These bacteria began to convert solar energy and carbon dioxide into bioenergy and oxygen more than two billion years ago. Cyanophages, which infect these bacteria, have an important role in regulating the marine ecosystem by controlling cyanobacteria community organization and mediating lateral gene transfer. Here we visualize the maturation process of cyanophage Syn5 inside its host cell, Synechococcus, using Zernike phase contrast electron cryo-tomography (cryoET)1,2. This imaging modality yields dramatic enhancement of image contrast over conventional cryoET and thus facilitates the direct identification of subcellular components, including thylakoid membranes, carboxysomes and polyribosomes, as well as phages, inside the congested cytosol of the infected cell. By correlating the structural features and relative abundance of viral progeny within cells at different stages of infection, we identify distinct Syn5 assembly intermediates. Our results indicate that the procapsid releases scaffolding proteins and expands its volume at an early stage of genome packaging. Later in the assembly process, we detected full particles with a tail either with or without an additional horn. The morphogenetic pathway we describe here is highly conserved and was probably established long before that of double-stranded DNA viruses infecting more complex organisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Danev, R. & Nagayama, K. Phase plates for transmission electron microscopy. Methods Enzymol. 481, 343–369 (2010)
Murata, K. et al. Zernike phase contrast cryo-electron microscopy and tomography for structure determination at nanometer and subnanometer resolutions. Structure 18, 903–912 (2010)
Fuller, N. J. et al. Clade-specific 16S ribosomal DNA oligonucleotides reveal the predominance of a single marine Synechococcus clade throughout a stratified water column in the red sea. Appl. Environ. Microbiol. 69, 2430–2443 (2003)
Rocap, G., Distel, D. L., Waterbury, J. B. & Chisholm, S. W. Resolution of Prochlorococcus and Synechococcus ecotypes by using 16S–23S ribosomal DNA internal transcribed spacer sequences. Appl. Environ. Microbiol. 68, 1180–1191 (2002)
Taylor, K. A. & Glaeser, R. M. Retrospective on the early development of cryoelectron microscopy of macromolecules and a prospective on opportunities for the future. J. Struct. Biol. 163, 214–223 (2008)
Danev, R., Glaeser, R. M. & Nagayama, K. Practical factors affecting the performance of a thin-film phase plate for transmission electron microscopy. Ultramicroscopy 109, 312–325 (2009)
Marko, M., Leith, A., Hsieh, C. & Danev, R. Retrofit implementation of Zernike phase plate imaging for cryo-TEM. J. Struct. Biol. 174, 400–412 (2011)
Rochat, R. H. et al. Seeing the portal in herpes simplex virus type 1 B capsids. J. Virol. 85, 1871–1874 (2011)
Liberton, M., Austin, J. R., II, Berg, R. H. & Pakrasi, H. B. Unique thylakoid membrane architecture of a unicellular N2-fixing cyanobacterium revealed by electron tomography. Plant Physiol. 155, 1656–1666 (2011)
Ting, C. S., Hsieh, C., Sundararaman, S., Mannella, C. & Marko, M. Cryo-electron tomography reveals the comparative three-dimensional architecture of Prochlorococcus, a globally important marine cyanobacterium. J. Bacteriol. 189, 4485–4493 (2007)
Iancu, C. V. et al. Organization, structure, and assembly of α-carboxysomes determined by electron cryotomography of intact cells. J. Mol. Biol. 396, 105–117 (2010)
Schmid, M. F. et al. Structure of Halothiobacillus neapolitanus carboxysomes by cryo-electron tomography. J. Mol. Biol. 364, 526–535 (2006)
Pope, W. H. et al. Genome sequence, structural proteins, and capsid organization of the cyanophage Syn5: a “horned” bacteriophage of marine synechococcus. J. Mol. Biol. 368, 966–981 (2007)
Chang, J. T. et al. Visualizing the structural changes of bacteriophage Epsilon15 and its Salmonella host during infection. J. Mol. Biol. 402, 731–740 (2010)
Hu, B., Margolin, W., Molineux, I. J. & Liu, J. The bacteriophage t7 virion undergoes extensive structural remodeling during infection. Science 339, 576–579 (2013)
Schmid, M. F. Single-particle electron cryotomography (cryoET). Adv. Protein Chem. Struct. Biol. 82, 37–65 (2011)
Schmid, M. F. & Booth, C. R. Methods for aligning and for averaging 3D volumes with missing data. J. Struct. Biol. 161, 243–248 (2008)
Raytcheva, D. A., Haase-Pettingell, C., Piret, J. M. & King, J. A. Intracellular assembly of cyanophage Syn5 proceeds through a scaffold-containing procapsid. J. Virol. 85, 2406–2415 (2011)
Chen, D. H. et al. Structural basis for scaffolding-mediated assembly and maturation of a dsDNA virus. Proc. Natl Acad. Sci. USA 108, 1355–1360 (2011)
Prevelige, P. E., Thomas, D. & King, J. Scaffolding protein regulates the polymerization of P22 coat subunits into icosahedral shells in vitro . J. Mol. Biol. 202, 743–757 (1988)
Hegde, S., Padilla-Sanchez, V., Draper, B. & Rao, V. B. Portal-large terminase interactions of the bacteriophage T4 DNA packaging machine implicate a molecular lever mechanism for coupling ATPase to DNA translocation. J. Virol. 86, 4046–4057 (2012)
Aksyuk, A. A. & Rossmann, M. G. Bacteriophage assembly. Viruses 3, 172–203 (2011)
Hendrix, R. W. Bacteriophage assembly. Nature 277, 172–173 (1979)
Fuller, D. N. et al. Measurements of single DNA molecule packaging dynamics in bacteriophage lambda reveal high forces, high motor processivity, and capsid transformations. J. Mol. Biol. 373, 1113–1122 (2007)
Machado, I. M. & Atsumi, S. Cyanobacterial biofuel production. J. Biotechnol. 162, 50–56 (2012)
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visulasualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996)
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999)
Tang, G. et al. EMAN2: an extensible image processing suite for electron microscopy. J. Struct. Biol. 157, 38–46 (2007)
Schmid, M. F. et al. A tail-like assembly at the portal vertex in intact herpes simplex type-1 virions. PLoS Pathog. 8, e1002961 (2012)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Henderson, R. et al. Outcome of the first electron microscopy validation task force meeting. Structure 20, 205–214 (2012)
Baker, M. L. et al. Validated near-atomic resolution structure of bacteriophage epsilon15 derived from cryo-EM and modeling. Proc. Natl Acad. Sci. USA 110, 12301–12306 (2013)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Acknowledgements
This research was supported by grants from the Robert Welch Foundation (Q1242) and National Institutes of Health (P41GM123832 to W.C.; AI0175208 and PN2EY016525 to W.C. and J.A.K.; GM080139 to S.J.L.; T15LM007093 through the Gulf Coast Consortia to W.D. and R.H.R.; T32GM007330 through the MSTP to R.H.R.). We thank J. G. Galaz-Montoya and R. N. Irobalieva for editing of the manuscript.
Author information
Authors and Affiliations
Contributions
W.D., D.R. and C.F. prepared the samples and conducted the infection experiments under the advice of C.H.-P. and J.P. W.D. collected the image data and reconstructed the tomograms; C.F. and H.A.K. established the Zernike phase plate imaging conditions in the microscope; K.N. provided the phase plates for imaging; R.H.R. performed the statistical analysis. W.D. and M.F.S. developed the imaging processing methods and solved the structures of the phage assembly intermediates; W.D. and X.L. refined the structures; J.F. and S.J.L. developed the symmetry-search algorithm for subvolume alignment; W.D., M.F.S., J.A.K. and W.C. interpreted the structures and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 ZPC improves contrast of cryoET images and reveals detailed structural features of Syn5-infected cells.
a, A conventional EM image of a Syn5-infected WH8109 cell. b, A ZPC image of the same cell as shown in a under the same imaging conditions.
Extended Data Figure 2 ZPC-cryoEM single-particle images of biochemically purified mature Syn5 phage.
The particles are shown with the tail pointing down and the wavy horn pointing up. The tail fibres appear to have variable conformations.
Extended Data Figure 3 General linear modelling of cellular carboxysome number with progression of infection.
The number of carboxysomes remains roughly constant as infection progresses, indicating that their variation does not correlate with progression of infection. The solid line indicates linear regression; the dashed lines indicate the 95% confidence limits.
Supplementary information
Zernike phase contrast tilt series images of a Syn5-infected WH8109 cell at an intermediate stage of infection
Tilt series of a frozen, hydrated Syn5-infected WH8109 cell was collected manually with an electron energy of 200 kV under low dose conditions on a 4kx4k CCD camera at 25,000× microscope magnification. The tilt angles ranged from -60° to 60° at 3° step increments. The accumulated electron exposure for the specimen in this tilt series was 40-50 electrons/Å2. The sampling of the data was calibrated to be 4.52Å/pixel. (MOV 11476 kb)
Volume rendering and annotation of the tomogram shown in video S1 with colour representations as in Fig. 2c
The tomogram was reconstructed using IMOD software. It is displayed both section-by-section and by volume rendering. Features are annotated by colours to designate molecular components attached and inside the cells, including cell envelope, thylakoid membrane, carboxysome, P-granules, ribosomes, infecting phages and phage assembly intermediates. (MOV 27885 kb)
Zernike phase contrast tilt series images of a Syn5-infected WH8109 cell at a late stage of infection with a ruptured cell membrane
Tilt series of frozen, hydrated Syn5-infected WH8109 cell was collected manually with an electron energy of 200 kV under low dose conditions on a 4kx4k CCD camera at 25,000× microscope magnification. The tilt angles ranged from -60° to 60° at 3° step increments. The accumulated electron exposure for the specimen in this tilt series was 40-50 electrons/Å2. The sampling of the data was calibrated to be 4.52Å/pixel. (MOV 13083 kb)
Volume rendering and annotation of the tomogram shown in video S3 with colour representations as in Fig. 1
The tomogram was reconstructed using IMOD software. It is displayed both section-by-section and by volume rendering. Features are annotated by colours to designate molecular components attached and inside the cells, including cell envelope, thylakoid membrane, carboxysome, P-granules, ribosomes, infecting phages and phage assembly intermediates. An increased number of DNA containing phage particles are visible, and overall contrast is enhanced over that in video S2, because the cell is beginning to rupture at this late stage of infection. (MOV 28142 kb)
Rights and permissions
About this article
Cite this article
Dai, W., Fu, C., Raytcheva, D. et al. Visualizing virus assembly intermediates inside marine cyanobacteria. Nature 502, 707–710 (2013). https://doi.org/10.1038/nature12604
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12604
This article is cited by
-
The application and development of electron microscopy for three-dimensional reconstruction in life science: a review
Cell and Tissue Research (2024)
-
Mortality by ribosomal sequencing (MoRS) provides a window into taxon-specific cell lysis
The ISME Journal (2023)
-
Abundant and cosmopolitan lineage of cyanopodoviruses lacking a DNA polymerase gene
The ISME Journal (2023)
-
Three-dimensional electron ptychography of organic–inorganic hybrid nanostructures
Nature Communications (2022)
-
Confocal microscopy reveals alterations of thylakoids in Limnospira fusiformis during prophage induction
Protoplasma (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.