Abstract
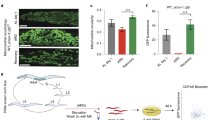
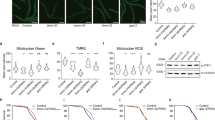
Longevity is regulated by a network of closely linked metabolic systems. We used a combination of mouse population genetics and RNA interference in Caenorhabditis elegans to identify mitochondrial ribosomal protein S5 (Mrps5) and other mitochondrial ribosomal proteins as metabolic and longevity regulators. MRP knockdown triggers mitonuclear protein imbalance, reducing mitochondrial respiration and activating the mitochondrial unfolded protein response. Specific antibiotics targeting mitochondrial translation and ethidium bromide (which impairs mitochondrial DNA transcription) pharmacologically mimic mrp knockdown and extend worm lifespan by inducing mitonuclear protein imbalance, a stoichiometric imbalance between nuclear and mitochondrially encoded proteins. This mechanism was also conserved in mammalian cells. In addition, resveratrol and rapamycin, longevity compounds acting on different molecular targets, similarly induced mitonuclear protein imbalance, the mitochondrial unfolded protein response and lifespan extension in C. elegans. Collectively these data demonstrate that MRPs represent an evolutionarily conserved protein family that ties the mitochondrial ribosome and mitonuclear protein imbalance to the mitochondrial unfolded protein response, an overarching longevity pathway across many species.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Fontana, L., Partridge, L. & Longo, V. D. Extending healthy life span–from yeast to humans. Science 328, 321–326 (2010)
Houtkooper, R. H., Williams, R. W. & Auwerx, J. Metabolic networks of longevity. Cell 142, 9–14 (2010)
Kenyon, C. J. The genetics of ageing. Nature 464, 504–512 (2010)
Mair, W. & Dillin, A. Aging and survival: the genetics of life span extension by dietary restriction. Annu. Rev. Biochem. 77, 727–754 (2008)
Pagliarini, D. J. et al. A mitochondrial protein compendium elucidates complex I disease biology. Cell 134, 112–123 (2008)
Sharma, M. R. et al. Structure of the mammalian mitochondrial ribosome reveals an expanded functional role for its component proteins. Cell 115, 97–108 (2003)
Anderson, S. et al. Sequence and organization of the human mitochondrial genome. Nature 290, 457–465 (1981)
Liao, C. Y., Rikke, B. A., Johnson, T. E., Diaz, V. & Nelson, J. F. Genetic variation in the murine lifespan response to dietary restriction: from life extension to life shortening. Aging Cell 9, 92–95 (2010)
Argmann, C. A., Chambon, P. & Auwerx, J. Mouse phenogenomics: the fast track to “systems metabolism”. Cell Metab. 2, 349–360 (2005)
Andreux, P. A. et al. Systems genetics of metabolism: the use of the BXD murine reference panel for multiscalar integration of traits. Cell 150, 1287–1299 (2012)
Peirce, J. L., Lu, L., Gu, J., Silver, L. M. & Williams, R. W. A new set of BXD recombinant inbred lines from advanced intercross populations in mice. BMC Genet. 5, 7 (2004)
Keane, T. M. et al. Mouse genomic variation and its effect on phenotypes and gene regulation. Nature 477, 289–294 (2011)
De Haan, G. & Van Zant, G. Genetic analysis of hemopoietic cell cycling in mice suggests its involvement in organismal life span. FASEB J. 13, 707–713 (1999)
Shifman, S. et al. A high-resolution single nucleotide polymorphism genetic map of the mouse genome. PLoS Biol. 4, e395 (2006)
Edwards, M. G. et al. Gene expression profiling of aging reveals activation of a p53-mediated transcriptional program. BMC Genomics 8, 80 (2007)
Geisert, E. E. et al. Gene expression in the mouse eye: an online resource for genetics using 103 strains of mice. Mol. Vis. 15, 1730–1763 (2009)
Chen, Y. et al. Variations in DNA elucidate molecular networks that cause disease. Nature 452, 429–435 (2008)
Lee, S. S. et al. A systematic RNAi screen identifies a critical role for mitochondria in C. elegans longevity. Nature Genet. 33, 40–48 (2003)
Hamilton, B. et al. A systematic RNAi screen for longevity genes in C. elegans. Genes Dev. 19, 1544–1555 (2005)
Hansen, M., Hsu, A. L., Dillin, A. & Kenyon, C. New genes tied to endocrine, metabolic, and dietary regulation of lifespan from a Caenorhabditis elegans genomic RNAi screen. PLoS Genet. 1, e17 (2005)
Wong, A., Boutis, P. & Hekimi, S. Mutations in the clk-1 gene of Caenorhabditis elegans affect developmental and behavioral timing. Genetics 139, 1247–1259 (1995)
Dillin, A. et al. Rates of behavior and aging specified by mitochondrial function during development. Science 298, 2398–2401 (2002)
Kenyon, C., Chang, J., Gensch, E., Rudner, A. & Tabtiang, R. A. C. elegans mutant that lives twice as long as wild type. Nature 366, 461–464 (1993)
Lakowski, B. & Hekimi, S. The genetics of caloric restriction in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 95, 13091–13096 (1998)
Durieux, J., Wolff, S. & Dillin, A. The cell-non-autonomous nature of electron transport chain-mediated longevity. Cell 144, 79–91 (2011)
Haynes, C. M. & Ron, D. The mitochondrial UPR - protecting organelle protein homeostasis. J. Cell Sci. 123, 3849–3855 (2010)
Zhao, Q. et al. A mitochondrial specific stress response in mammalian cells. EMBO J. 21, 4411–4419 (2002)
Yoneda, T. et al. Compartment-specific perturbation of protein handling activates genes encoding mitochondrial chaperones. J. Cell Sci. 117, 4055–4066 (2004)
Haynes, C. M., Yang, Y., Blais, S. P., Neubert, T. A. & Ron, D. The matrix peptide exporter HAF-1 signals a mitochondrial UPR by activating the transcription factor ZC376.7 in C. elegans.. Mol. Cell 37, 529–540 (2010)
Benedetti, C., Haynes, C. M., Yang, Y., Harding, H. P. & Ron, D. Ubiquitin-like protein 5 positively regulates chaperone gene expression in the mitochondrial unfolded protein response. Genetics 174, 229–239 (2006)
Overall, R. W. et al. Genetics of the hippocampal transcriptome in mouse: a systematic survey and online neurogenomics resource. Front. Neurosci. 3, 55 (2009)
Zylbee, E., Vesco, C. & Penman, S. Selective inhibition of the synthesis of mitochondria-associated RNA by ethidium bromide. J. Mol. Biol. 44, 195–204 (1969)
Harrison, D. E. et al. Rapamycin fed late in life extends lifespan in genetically heterogeneous mice. Nature 460, 392–395 (2009)
Robida-Stubbs, S. et al. TOR signaling and rapamycin influence longevity by regulating SKN-1/Nrf and DAF-16/FoxO. Cell Metab. 15, 713–724 (2012)
Zid, B. M. et al. 4E-BP extends lifespan upon dietary restriction by enhancing mitochondrial activity in Drosophila. Cell 139, 149–160 (2009)
Schulz, T. J. et al. Glucose restriction extends Caenorhabditis elegans life span by inducing mitochondrial respiration and increasing oxidative stress. Cell Metab. 6, 280–293 (2007)
Zarse, K. et al. Impaired insulin/IGF1 signaling extends life span by promoting mitochondrial L-proline catabolism to induce a transient ROS signal. Cell Metab. 15, 451–465 (2012)
Schägger, H. & Pfeiffer, K. Supercomplexes in the respiratory chains of yeast and mammalian mitochondria. EMBO J. 19, 1777–1783 (2000)
De Haan, G. & Van Zant, G. Genetic analysis of hemopoietic cell cycling in mice suggests its involvement in organismal life span. FASEB J. 13, 707–713 (1999)
Geisert, E. E. et al. Gene expression in the mouse eye: an online resource for genetics using 103 strains of mice. Mol. Vis. 15, 1730–1763 (2009)
Chen, Y. et al. Variations in DNA elucidate molecular networks that cause disease. Nature 452, 429–435 (2008)
Reich, M. et al. GenePattern 2.0. Nature Genet. 38, 500–501 (2006)
de Hoon, M. J., Imoto, S., Nolan, J. & Miyano, S. Open source clustering software. Bioinformatics 20, 1453–1454 (2004)
Kamath, R. S., Martinez-Campos, M., Zipperlen, P., Fraser, A. G. & Ahringer, J. Effectiveness of specific RNA-mediated interference through ingested double-stranded RNA in Caenorhabditis elegans. Genome Biol. 2, research0002––research0002.10 (2000)
Mouchiroud, L. et al. Pyruvate imbalance mediates metabolic reprogramming and mimics lifespan extension by dietary restriction in Caenorhabditis elegans. Aging Cell 10, 39–54 (2011)
Wood, J. G. et al. Sirtuin activators mimic caloric restriction and delay ageing in metazoans. Nature 430, 686–689 (2004)
Durieux, J., Wolff, S. & Dillin, A. The cell-non-autonomous nature of electron transport chain-mediated longevity. Cell 144, 79–91 (2011)
Yang, W. & Hekimi, S. A mitochondrial superoxide signal triggers increased longevity in Caenorhabditis elegans. PLoS Biol. 8, e1000556 (2010)
Watanabe, M. et al. Bile acids induce energy expenditure by promoting intracellular thyroid hormone activation. Nature 439, 484–489 (2006)
Lagouge, M. et al. Resveratrol improves mitochondrial function and protects against metabolic disease by activating SIRT1 and PGC-1α. Cell 127, 1109–1122 (2006)
Ryu, D. et al. Endoplasmic reticulum stress promotes LIPIN2-dependent hepatic insulin resistance. Diabetes 60, 1072–1081 (2011)
Noriega, L. G. et al. CREB and ChREBP oppositely regulate SIRT1 expression in response to energy availability. EMBO Rep. 12, 1069–1076 (2011)
Acknowledgements
We thank P. Gönczy and the Caenorhabditis Genetics Center for sharing or providing reagents. R.H.H. is supported by fellowships from NWO-Rubicon and AMC, and L.M. by an FRM fellowship. J.A. is the Nestlé Chair in Energy Metabolism and supported by EPFL, ERC (2008-AdG-23138), Velux Stiftung and SNSF. R.W.W. and GeneNetwork are supported by the National Institutes of Health (NIH) (P20-DA 21131, UO1AA13499 and U01AA14425), and the UT Center for Integrative and Translational Genomics. R.W.W. and J.A. are supported by NIH grant R01AG043930.
Author information
Authors and Affiliations
Contributions
D.R., N.M. and E.K. contributed equally to this work. R.H.H., L.M. and J.A. conceived and designed the project. R.H.H. and R.W.W. performed QTL mapping and sequence analyses. R.H.H., L.M., E.K., D.R., N.M., G.K. performed experiments. R.H.H. and J.A. wrote the manuscript with contributions from all other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-4 and Supplementary Figures 1-10. (PDF 4703 kb)
Movement of wild type worms at day 13 of adulthood
Time lapse video of 13-day old N2 worms treated with HT115 control bacteria. (MOV 22 kb)
Movement of mrps-5 RNAi worms at day 13 of adulthood
Time lapse video of 13-day old N2 worms treated with mrps-5 RNAi bacteria. These worms move more than HT115 controls (see Supplementary Video 1). (MOV 19 kb)
Movement of wild type worms at day 20 of adulthood
Time lapse video of 20-day old N2 worms treated with HT115 control bacteria. (MOV 13 kb)
Movement of mrps-5 RNAi worms at day 20 of adulthood
Time lapse video of 20-day old N2 worms treated with mrps-5 RNAi bacteria. These worms move more than HT115 controls (see Supplementary Video 3). (MOV 15 kb)
Rights and permissions
About this article
Cite this article
Houtkooper, R., Mouchiroud, L., Ryu, D. et al. Mitonuclear protein imbalance as a conserved longevity mechanism. Nature 497, 451–457 (2013). https://doi.org/10.1038/nature12188
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12188
This article is cited by
-
Mitochondrial perturbation in immune cells enhances cell-mediated innate immunity in Drosophila
BMC Biology (2024)
-
N1-methylation of adenosine (m1A) in ND5 mRNA leads to complex I dysfunction in Alzheimer’s disease
Molecular Psychiatry (2024)
-
NAD+ dependent UPRmt activation underlies intestinal aging caused by mitochondrial DNA mutations
Nature Communications (2024)
-
Reducing mitochondrial ribosomal gene expression does not alter metabolic health or lifespan in mice
Scientific Reports (2023)
-
Coupling of autophagy and the mitochondrial intrinsic apoptosis pathway modulates proteostasis and ageing in Caenorhabditis elegans
Cell Death & Disease (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.