Abstract
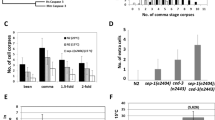
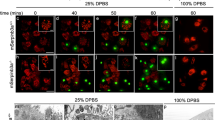
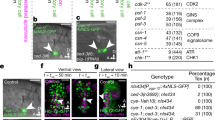
The elimination of unnecessary or defective cells from metazoans occurs during normal development and tissue homeostasis, as well as in response to infection or cellular damage1. Although many cells are removed through caspase-mediated apoptosis followed by phagocytosis by engulfing cells2, other mechanisms of cell elimination occur3, including the extrusion of cells from epithelia through a poorly understood, possibly caspase-independent, process4. Here we identify a mechanism of cell extrusion that is caspase independent and that can eliminate a subset of the Caenorhabditis elegans cells programmed to die during embryonic development. In wild-type animals, these cells die soon after their generation through caspase-mediated apoptosis. However, in mutants lacking all four C. elegans caspase genes, these cells are eliminated by being extruded from the developing embryo into the extra-embryonic space of the egg. The shed cells show apoptosis-like cytological and morphological characteristics, indicating that apoptosis can occur in the absence of caspases in C. elegans. We describe a kinase pathway required for cell extrusion involving PAR-4, STRD-1 and MOP-25.1/-25.2, the C. elegans homologues of the mammalian tumour-suppressor kinase LKB1 and its binding partners STRADα and MO25α. The AMPK-related kinase PIG-1, a possible target of the PAR-4–STRD-1–MOP-25 kinase complex, is also required for cell shedding. PIG-1 promotes shed-cell detachment by preventing the cell-surface expression of cell-adhesion molecules. Our findings reveal a mechanism for apoptotic cell elimination that is fundamentally distinct from that of canonical programmed cell death.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Fuchs, Y. & Steller, H. Programmed cell death in animal development and disease. Cell 147, 742–758 (2011)
Reddien, P. W. & Horvitz, H. R. The engulfment process of programmed cell death in Caenorhabditis elegans . Annu. Rev. Cell Dev. Biol. 20, 193–221 (2004)
Yuan, J. & Kroemer, G. Alternative cell death mechanisms in development and beyond. Genes Dev. 24, 2592–2602 (2010)
Rosenblatt, J., Raff, M. C. & Cramer, L. P. An epithelial cell destined for apoptosis signals its neighbors to extrude it by an actin- and myosin-dependent mechanism. Curr. Biol. 11, 1847–1857 (2001)
Ellis, H. M. & Horvitz, H. R. Genetic control of programmed cell death in the nematode C. elegans . Cell 44, 817–829 (1986)
Abraham, M. C., Lu, Y. & Shaham, S. A morphologically conserved nonapoptotic program promotes linker cell death in Caenorhabditis elegans . Dev. Cell 12, 73–86 (2007)
Shaham, S., Reddien, P. W., Davies, B. & Horvitz, H. R. Mutational analysis of the Caenorhabditis elegans cell-death gene ced-3 . Genetics 153, 1655–1671 (1999)
Wu, Y. C., Stanfield, G. M. & Horvitz, H. R. NUC-1, a Caenorhabditis elegans DNase II homolog, functions in an intermediate step of DNA degradation during apoptosis. Genes Dev. 14, 536–548 (2000)
Shaham, S. Identification of multiple Caenorhabditis elegans caspases and their potential roles in proteolytic cascades. J. Biol. Chem. 273, 35109–35117 (1998)
Sulston, J. E. & Horvitz, H. R. Post-embryonic cell lineages of the nematode, Caenorhabditis elegans . Dev. Biol. 56, 110–156 (1977)
Sulston, J. E., Schierenberg, E., White, J. G. & Thomson, J. N. The embryonic cell lineage of the nematode Caenorhabditis elegans . Dev. Biol. 100, 64–119 (1983)
Potts, M. B. & Cameron, S. Cell lineage and cell death: Caenorhabditis elegans and cancer research. Nature Rev. Cancer 11, 50–58 (2011)
Buechner, M. Tubes and the single C. elegans excretory cell. Trends Cell Biol. 12, 479–484 (2002)
Abdus-Saboor, I. et al. Notch and Ras promote sequential steps of excretory tube development in C. elegans . Development 138, 3545–3555 (2011)
Cordes, S., Frank, C. A. & Garriga, G. The C. elegans MELK ortholog PIG-1 regulates cell size asymmetry and daughter cell fate in asymmetric neuroblast divisions. Development 133, 2747–2756 (2006)
Ou, G., Stuurman, N., D’Ambrosio, M. & Vale, R. D. Polarized myosin produces unequal-size daughters during asymmetric cell division. Science 330, 677–680 (2010)
Shackelford, D. B. & Shaw, R. J. The LKB1–AMPK pathway: metabolism and growth control in tumour suppression. Nature Rev. Cancer 9, 563–575 (2009)
Lizcano, J. M. et al. LKB1 is a master kinase that activates 13 kinases of the AMPK subfamily, including MARK/PAR-1. EMBO J. 23, 833–843 (2004)
Boudeau, J. et al. MO25α/β interact with STRADα/β enhancing their ability to bind, activate and localize LKB1 in the cytoplasm. EMBO J. 22, 5102–5114 (2003)
Zeqiraj, E., Filippi, B. M., Deak, M., Alessi, D. R. & van Aalten, D. M. F. Structure of the LKB1-STRAD-MO25 complex reveals an allosteric mechanism of kinase activation. Science 326, 1707–1711 (2009)
Narbonne, P., Hyenne, V., Li, S., Labbé, J.-C. & Roy, R. Differential requirements for STRAD in LKB1-dependent functions in C. elegans . Development 137, 661–670 (2010)
Labouesse, M. Epithelial junctions and attachments. WormBook 13, 1–21 (2006)
Kinchen, J. M. et al. A pathway for phagosome maturation during engulfment of apoptotic cells. Nature Cell Biol. 10, 556–566 (2008)
Singhvi, A. et al. The Arf GAP CNT-2 regulates the apoptotic fate in C. elegans asymmetric neuroblast divisions. Curr. Biol. 21, 948–954 (2011)
D’Souza-Schorey, C. & Chavrier, P. ARF proteins: roles in membrane traffic and beyond. Nature Rev. Mol. Cell Biol. 7, 347–358 (2006)
D’Souza-Schorey, C. Disassembling adherens junctions: breaking up is hard to do. Trends Cell Biol. 15, 19–26 (2005)
Potten, C. S. Stem cells in gastrointestinal epithelium: numbers, characteristics and death. Phil. Trans. R. Soc. Lond. B 353, 821–830 (1998)
Watson, A. J. M., Duckworth, C. A., Guan, Y. & Montrose, M. H. Mechanisms of epithelial cell shedding in the mammalian intestine and maintenance of barrier function. Ann. NY Acad. Sci. 1165, 135–142 (2009)
Coopersmith, C. M., O’Donnell, D. & Gordon, J. I. Bcl-2 inhibits ischemia-reperfusion-induced apoptosis in the intestinal epithelium of transgenic mice. Am. J. Physiol. 276, G677–G686 (1999)
Hemminki, A. et al. A serine/threonine kinase gene defective in Peutz–Jeghers syndrome. Nature 391, 184–187 (1998)
Brenner, S. The genetics of Caenorhabditis elegans . Genetics 77, 71–94 (1974)
Hirose, T., Galvin, B. D. & Horvitz, H. R. Six and Eya promote apoptosis through direct transcriptional activation of the proapoptotic BH3-only gene egl-1 in Caenorhabditis elegans . Proc. Natl Acad. Sci. USA 107, 15479–15484 (2010)
Venegas, V. & Zhou, Z. Two alternative mechanisms that regulate the presentation of apoptotic cell engulfment signal in Caenorhabditis elegans . Mol. Biol. Cell 18, 3180–3192 (2007)
Schwartz, H. T. A protocol describing pharynx counts and a review of other assays of apoptotic cell death in the nematode worm Caenorhabditis elegans . Nature Protocols 2, 705–714 (2007)
Fraser, A. G. et al. Functional genomic analysis of C. elegans chromosome I by systematic RNA interference. Nature 408, 325–330 (2000)
Rual, J. F. et al. Toward improving Caenorhabditis elegans phenome mapping with an ORFeome-based RNAi library. Genome Res. 14, 2162–2168 (2004)
Parrish, J. Z. & Xue, D. Functional genomic analysis of apoptotic DNA degradation in C. elegans . Mol. Cell 11, 987–996 (2003)
Lu, Q. et al. elegans Rab GTPase 2 is required for the degradation of apoptotic cells. Development 135, 1069–1080 (2008)
Acknowledgements
We thank Z. Zhou, T. Hirose and S. Nakano for reporter constructs; B. Castor, E. Murphy and R. Droste for determining DNA sequences; S. Dennis for screening of the deletion library; N. An for strain management; and, J. Yuan, S. Nakano, N. Paquin, C. Engert and A. Saffer for discussions. The Caenorhabditis Genetic Center, which is funded by the National Institutes of Health National Center for Research Resources (NCRR), provided many strains. S. Mitani provided grp-1(tm1956). D.P.D. was supported by post-doctoral fellowships from the Damon Runyon Cancer Research Foundation and from the Charles A. King Trust. H.R.H. is the David H. Koch Professor of Biology at the Massachusetts Institute of Technology and an Investigator at the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
D.P.D. and H.R.H. designed the experiments, analysed the data and wrote the manuscript. D.P.D. and V.H. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1 -11, Supplementary Figures 1-8, legends for Supplementary Movies 1-3 (see separate files for Supplementary Movies), Supplementary References and the RNAi Construct Sequences. (PDF 9773 kb)
Supplementary Movie 1
This file contains a time-lapse movie of a developing ced-3(n3692) embryo (see Supplementary Information file for full legend). (MOV 5090 kb)
Supplementary Movie 2
This file contains a time-lapse movie of a developing pig-1(gm344) embryo (see Supplementary Information file for full legend). (MOV 2086 kb)
Supplementary Movie 3
This file contains a time-lapse movie of a developing pig-1(gm344) ced-3(n3692) embryo (see Supplementary Information file for full legend). (MOV 3778 kb)
Rights and permissions
About this article
Cite this article
Denning, D., Hatch, V. & Horvitz, H. Programmed elimination of cells by caspase-independent cell extrusion in C. elegans. Nature 488, 226–230 (2012). https://doi.org/10.1038/nature11240
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11240
This article is cited by
-
Replication stress promotes cell elimination by extrusion
Nature (2021)
-
Programmed cell removal by calreticulin in tissue homeostasis and cancer
Nature Communications (2018)
-
Comparative genetic, proteomic and phosphoproteomic analysis of C. elegans embryos with a focus on ham-1/STOX and pig-1/MELK in dopaminergic neuron development
Scientific Reports (2017)
-
Transcriptional control of non-apoptotic developmental cell death in C. elegans
Cell Death & Differentiation (2016)
-
Both the anti- and pro-apoptotic functions of villin regulate cell turnover and intestinal homeostasis
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.