Abstract
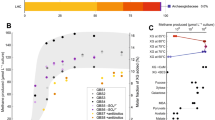
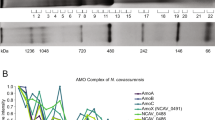
The anaerobic oxidation of methane (AOM) with sulphate, an area currently generating great interest in microbiology, is accomplished by consortia of methanotrophic archaea (ANME) and sulphate-reducing bacteria1,2. The enzyme activating methane in methanotrophic archaea has tentatively been identified as a homologue of methyl-coenzyme M reductase (MCR) that catalyses the methane-forming step in methanogenic archaea3,4. Here we report an X-ray structure of the 280 kDa heterohexameric ANME-1 MCR complex. It was crystallized uniquely from a protein ensemble purified from consortia of microorganisms collected with a submersible from a Black Sea mat catalysing AOM with sulphate4. Crystals grown from the heterogeneous sample diffract to 2.1 Å resolution and consist of a single ANME-1 MCR population, demonstrating the strong selective power of crystallization. The structure revealed ANME-1 MCR in complex with coenzyme M and coenzyme B, indicating the same substrates for MCR from methanotrophic and methanogenic archaea. Differences between the highly similar structures of ANME-1 MCR and methanogenic MCR include a F430 modification, a cysteine-rich patch and an altered post-translational amino acid modification pattern, which may tune the enzymes for their functions in different biological contexts.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Reeburgh, W. S. Oceanic methane biogeochemistry. Chem. Rev. 107, 486–513 (2007)
Hallam, S. J. et al. Reverse methanogenesis: testing the hypothesis with environmental genomics. Science 305, 1457–1462 (2004)
Scheller, S., Goenrich, M., Boecher, R., Thauer, R. K. & Jaun, B. The key nickel enzyme of methanogenesis catalyses the anaerobic oxidation of methane. Nature 465, 606–608 (2010)
Krüger, M. et al. A conspicuous nickel protein in microbial mats that oxidize methane anaerobically. Nature 426, 878–881 (2003)
Meyerdierks, A. et al. Metagenome and mRNA expression analyses of anaerobic methanotrophic archaea of the ANME-1 group. Environ. Microbiol. 12, 422–439 (2010)
Knittel, K. & Boetius, A. Anaerobic oxidation of methane: progress with an unknown process. Annu. Rev. Microbiol. 63, 311–334 (2009)
Losekann, T. et al. Diversity and abundance of aerobic and anaerobic methane oxidizers at the Haakon Mosby Mud Volcano, Barents Sea. Appl. Environ. Microbiol. 73, 3348–3362 (2007)
Thauer, R. K. Anaerobic oxidation of methane with sulfate: on the reversibility of the reactions that are catalyzed by enzymes also involved in methanogenesis from CO2 . Curr. Opin. Microbiol. 14, 292–299 (2011)
Thauer, R. K. & Shima, S. Methane as fuel for anaerobic microorganisms. Ann. NY Acad. Sci. 1125, 158–170 (2008)
Nauhaus, K., Treude, T., Boetius, A. & Krüger, M. Environmental regulation of the anaerobic oxidation of methane: a comparison of ANME-I and ANME-II communities. Environ. Microbiol. 7, 98–106 (2005)
Heller, C., Hoppert, M. & Reitner, J. Immunological localization of coenzyme M reductase in anaerobic methane-oxidizing archaea of ANME 1 and ANME 2 type. Geomicrobiol. J. 25, 149–156 (2008)
Mayr, S. et al. Structure of an F430 variant from archaea associated with anaerobic oxidation of methane. J. Am. Chem. Soc. 130, 10758–10767 (2008)
Moran, J. J. et al. Methyl sulfides as intermediates in the anaerobic oxidation of methane. Environ. Microbiol. 10, 162–173 (2008)
Kahnt, J. et al. Post-translational modifications in the active site region of methyl-coenzyme M reductase from methanogenic and methanotrophic archaea. FEBS J. 274, 4913–4921 (2007)
Knittel, K., Losekann, T., Boetius, A., Kort, R. & Amann, R. Diversity and distribution of methanotrophic archaea at cold seeps. Appl. Environ. Microbiol. 71, 467–479 (2005)
Grabarse, W. et al. On the mechanism of biological methane formation: structural evidence for conformational changes in methyl-coenzyme M reductase upon substrate binding. J. Mol. Biol. 309, 315–330 (2001)
Scheller, S., Goenrich, M., Mayr, S., Thauer, R. K. & Jaun, B. Intermediates in the catalytic cycle of methyl coenzyme M reductase: isotope exchange is consistent with formation of a σ-alkane–nickel complex. Angew. Chem. Int. Edn Engl. 49, 8112–8115 (2010)
Dey, M., Li, X., Kunz, R. C. & Ragsdale, S. W. Detection of organometallic and radical intermediates in the catalytic mechanism of methyl-coenzyme M reductase using the natural substrate methyl-coenzyme M and a coenzyme B substrate analogue. Biochemistry 49, 10902–10911 (2010)
Thauer, R. K. Biochemistry of methanogenesis: a tribute to Marjory Stephenson. 1998 Marjory Stephenson Prize Lecture. Microbiology 144, 2377–2406 (1998)
Satzger, H. et al. Photoswitchable elements within a peptide backbone-ultrafast spectroscopy of thioxylated amides. J. Phys. Chem. B 109, 4770–4775 (2005)
Grove, T. L. et al. A radically different mechanism for S-adenosylmethionine-dependent methyltransferases. Science 332, 604–607 (2011)
Krüger, M. et al. A novel, multi-layered methanotrophic microbial mat system growing on the sediment of the Black Sea. Environ. Microbiol. 10, 1934–1947 (2008)
Michaelis, W. et al. Microbial reefs in the Black Sea fueled by anaerobic oxidation of methane. Science 297, 1013–1015 (2002)
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Cryst. 26, 795–800 (1993)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Afonine, P. V. et al. phenix. model_vs_data: a high-level tool for the calculation of crystallographic model and data statistics. J. Appl. Cryst. 43, 669–676 (2010)
Grabarse, W., Mahlert, F., Shima, S., Thauer, R. K. & Ermler, U. Comparison of three methyl-coenzyme M reductases from phylogenetically distant organisms: unusual amino acid modification, conservation and adaptation. J. Mol. Biol. 303, 329–344 (2000)
Jensen, L. M., Sanishvili, R., Davidson, V. L. & Wilmot, C. M. In crystallo posttranslational modification within a MauG/pre-methylamine dehydrogenase complex. Science 327, 1392–1394 (2010)
Acknowledgements
This work was supported by the Max Planck Society, the Fonds der Chemischen Industrie, the Deutsche Forschungsgemeinschaft (SPP 1319, ER 222/5-1, KR 3311/6-2) and the projects MUMM, and BEBOP (Biogeochemistry and Microbiology of Bioherms Prospering in the Black Sea funded by the University of Hamburg). S.S. was financed by the PRESTO program, Japan Science and Technology Agency (JST). We are indebted to W. Michaelis and R. Seifert for access to samples. We thank the crews of RV Poseidon with JAGO for help during field work, M. Brefort and L. Känel for the electron spin resonance measurement and Edman sequencing, respectively. We are grateful to H. Michel for continuous support, the staff of the PXII beamline at the Swiss-Light-Source, Villigen for assistance during data collection, and K. Davies for critical reading of the manuscript.
Author information
Authors and Affiliations
Contributions
S.S., R.K.T. and U.E. designed the experiments, interpreted the data and wrote the paper. The mat sample from the Black Sea was collected and maintained by M.K. The enzyme was purified and characterized by M.K. and S.S. Crystallization was performed by S.S. and U.D.; U.E. collected the X-ray data. T.W. refined the structure and found that the primary structure conformed with that of the D1JBK2-4 sequences in the metagenome. J.K. performed the mass-spectrometric analyses.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Atomic coordinates and structure factors of ANME-1 MCR have been deposited in the Protein Data Bank with the code 3SQG.
Supplementary information
Supplementary Information
The file contains Supplementary Tables 1-2, Supplementary Figures 1-5 with legends and additional references. (PDF 646 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Shima, S., Krueger, M., Weinert, T. et al. Structure of a methyl-coenzyme M reductase from Black Sea mats that oxidize methane anaerobically. Nature 481, 98–101 (2012). https://doi.org/10.1038/nature10663
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10663
This article is cited by
-
A microbe that uses crude oil to make methane
Nature (2022)
-
Expression of divergent methyl/alkyl coenzyme M reductases from uncultured archaea
Communications Biology (2022)
-
Insight into anaerobic methanotrophy from 13C/12C- amino acids and 14C/12C-ANME cells in seafloor microbial ecology
Scientific Reports (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.