Abstract
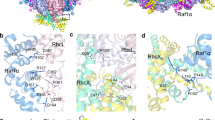
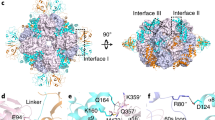
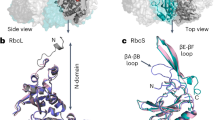
Ribulose 1,5-bisphosphate carboxylase/oxygenase (Rubisco) catalyses the fixation of atmospheric CO2 in photosynthesis, but tends to form inactive complexes with its substrate ribulose 1,5-bisphosphate (RuBP). In plants, Rubisco is reactivated by the AAA+ (ATPases associated with various cellular activities) protein Rubisco activase (Rca), but no such protein is known for the Rubisco of red algae. Here we identify the protein CbbX as an activase of red-type Rubisco. The 3.0-Å crystal structure of unassembled CbbX from Rhodobacter sphaeroides revealed an AAA+ protein architecture. Electron microscopy and biochemical analysis showed that ATP and RuBP must bind to convert CbbX into functionally active, hexameric rings. The CbbX ATPase is strongly stimulated by RuBP and Rubisco. Mutational analysis suggests that CbbX functions by transiently pulling the carboxy-terminal peptide of the Rubisco large subunit into the hexamer pore, resulting in the release of the inhibitory RuBP. Understanding Rubisco activation may facilitate efforts to improve CO2 uptake and biomass production by photosynthetic organisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
Coordinates and structure factor amplitudes for CbbX crystal structures are deposited in the Protein Data Bank (PDB) under accession codes 3SYL and 3SYK; the hexamer model and the electron microscopy density are deposited in the PDB under accession code 3ZUH and in the Electron Microscopy Database (http://www.ebi.ac.uk/pdbe/emdb/) under accession code EMD-1932, respectively.
References
Spreitzer, R. J. & Salvucci, M. E. Rubisco: structure, regulatory interactions, and possibilities for a better enzyme. Annu. Rev. Plant Biol. 53, 449–475 (2002)
Andersson, I. & Backlund, A. Structure and function of Rubisco. Plant Physiol. Biochem. 46, 275–291 (2008)
Tabita, F. R. Microbial ribulose 1,5-bisphosphate carboxylase/oxygenase: a different perspective. Photosynth. Res. 60, 1–28 (1999)
Tabita, F. R., Satagopan, S., Hanson, T. E., Kreel, N. E. & Scott, S. S. Distinct form I, II, III, and IV Rubisco proteins from the three kingdoms of life provide clues about Rubisco evolution and structure/function relationships. J. Exp. Bot. 59, 1515–1524 (2008)
Badger, M. R. & Bek, E. J. Multiple Rubisco forms in proteobacteria: their functional significance in relation to CO2 acquisition by the CBB cycle. J. Exp. Bot. 59, 1525–1541 (2008)
Whitney, S. M., Baldet, P., Hudson, G. S. & Andrews, T. J. Form I Rubiscos from non-green algae are expressed abundantly but not assembled in tobacco chloroplasts. Plant J. 26, 535–547 (2001)
Falkowski, P. G. et al. The evolution of modern eukaryotic phytoplankton. Science 305, 354–360 (2004)
Lorimer, G. H., Badger, M. R. & Andrews, T. J. The activation of ribulose-1,5-bisphosphate carboxylase by carbon dioxide and magnesium ions. Equilibria, kinetics, a suggested mechanism, and physiological implications. Biochemistry 15, 529–536 (1976)
Jordan, D. B. & Chollet, R. Inhibition of ribulose bisphosphate carboxylase by substrate ribulose 1,5-bisphosphate. J. Biol. Chem. 258, 13752–13758 (1983)
Portis, A. R., Jr Rubisco activase—Rubisco’s catalytic chaperone. Photosynth. Res. 75, 11–27 (2003)
Hanson, P. I. & Whiteheart, S. W. AAA+ proteins: have engine, will work. Nature Rev. Mol. Cell Biol. 6, 519–529 (2005)
Pearce, F. G. Catalytic by-product formation and ligand binding by ribulose bisphosphate carboxylases from different phylogenies. Biochem. J. 399, 525–534 (2006)
Gibson, J. L. & Tabita, F. R. Analysis of the cbbXYZ operon in Rhodobacter sphaeroides. J. Bacteriol. 179, 663–669 (1997)
Maier, U. G., Fraunholz, M., Zauner, S., Penny, S. & Douglas, S. A nucleomorph-encoded CbbX and the phylogeny of RuBisCo regulators. Mol. Biol. Evol. 17, 576–583 (2000)
Fujita, K., Tanaka, K., Sadaie, Y. & Ohta, N. Functional analysis of the plastid and nuclear encoded CbbX proteins of Cyanidioschyzon merolae. Genes Genet. Syst. 83, 135–142 (2008)
Bowien, B. & Kusian, B. Genetics and control of CO2 assimilation in the chemoautotroph Ralstonia eutropha. Arch. Microbiol. 178, 85–93 (2002)
Saschenbrecker, S. et al. Structure and function of RbcX, an assembly chaperone for hexadecameric Rubisco. Cell 129, 1189–1200 (2007)
Liu, C. et al. Coupled chaperone action in folding and assembly of hexadecameric Rubisco. Nature 463, 197–202 (2010)
Gibson, J. L. & Tabita, F. R. Activation of ribulose 1,5-bisphosphate carboxylase from Rhodopseudomonas sphaeroides: probable role of the small subunit. J. Bacteriol. 140, 1023–1027 (1979)
Robinson, S. P. & Portis, A. R., Jr Adenosine triphosphate hydrolysis by purified rubisco activase. Arch. Biochem. Biophys. 268, 93–99 (1989)
Sugawara, H. et al. Crystal structure of carboxylase reaction-oriented ribulose 1,5-bisphosphate carboxylase/oxygenase from a thermophilic red alga, Galdieria partita. J. Biol. Chem. 274, 15655–15661 (1999)
Okano, Y. et al. X-ray structure of Galdieria Rubisco complexed with one sulfate ion per active site. FEBS Lett. 527, 33–36 (2002)
Von Caemmerer, S. & Edmondson, D. L. Relationship between steady-state gas exchange in vivo ribulose bisphosphate carboxylase activity and some carbon reduction cycle intermediates in Raphanus sativus. Aust. J. Plant Physiol. 13, 669–688 (1986)
Sousa, M. C. et al. Crystal and solution structures of an HslUV protease–chaperone complex. Cell 103, 633–643 (2000)
Massey, T. H., Mercogliano, C. P., Yates, J., Sherratt, D. J. & Lowe, J. Double-stranded DNA translocation: structure and mechanism of hexameric FtsK. Mol. Cell 23, 457–469 (2006)
Matias, P. M., Gorynia, S., Donner, P. & Carrondo, M. A. Crystal structure of the human AAA+ protein RuvBL1. J. Biol. Chem. 281, 38918–38929 (2006)
Davies, J. M., Brunger, A. T. & Weis, W. I. Improved structures of full-length p97, an AAA ATPase: implications for mechanisms of nucleotide-dependent conformational change. Structure 16, 715–726 (2008)
Glynn, S. E., Martin, A., Nager, A. R., Baker, T. A. & Sauer, R. T. Structures of asymmetric ClpX hexamers reveal nucleotide-dependent motions in a AAA+ protein-unfolding machine. Cell 139, 744–756 (2009)
Weibezahn, J. et al. Thermotolerance requires refolding of aggregated proteins by substrate translocation through the central pore of ClpB. Cell 119, 653–665 (2004)
Hinnerwisch, J., Fenton, W. A., Furtak, K. J., Farr, G. W. & Horwich, A. L. Loops in the central channel of ClpA chaperone mediate protein binding, unfolding, and translocation. Cell 121, 1029–1041 (2005)
Martin, A., Baker, T. A. & Sauer, R. T. Pore loops of the AAA+ ClpX machine grip substrates to drive translocation and unfolding. Nature Struct. Mol. Biol. 15, 1147–1151 (2008)
Roll-Mecak, A. & Vale, R. D. Structural basis of microtubule severing by the hereditary spastic paraplegia protein spastin. Nature 451, 363–367 (2008)
Andersson, I. Catalysis and regulation in Rubisco. J. Exp. Bot. 59, 1555–1568 (2008)
Catanzariti, A.-M., Soboleva, T. A., Jans, D. A., Board, P. G. & Baker, R. T. An efficient system for high-level expression and easy purification of authentic recombinant proteins. Protein Sci. 13, 1331–1339 (2004)
Baker, R. T. et al. Using deubiquitylating enzymes as research tools. Methods Enzymol. 398, 540–554 (2005)
Esau, B. D., Snyder, G. W. & Portis, A. R., Jr Differential effects of N- and C-terminal deletions on the two activities of rubisco activase. Arch. Biochem. Biophys. 326, 100–105 (1996)
Kreuzer, K. N. & Jongeneel, C. V. Escherichia coli phage T4 topoisomerase. Methods Enzymol. 100, 144–160 (1983)
Parry, M. A. J., Keys, A. J., Madgwick, P. J., Carmo-Silva, A. E. & Andralojc, P. J. Rubisco regulation: a role for inhibitors. J. Exp. Bot. 59, 1569–1580 (2008)
Pitcher, D. G., Saunders, N. A. & Owen, R. J. Rapid extraction of bacterial genomic DNA with guanidium thiocyanate. Lett. Appl. Microbiol. 8, 151–156 (1989)
Gibson, J. L., Falcone, D. L. & Tabita, F. R. Nucleotide sequence, transcriptional analysis, and expression of genes encoded within the form I CO2 fixation operon of Rhodobacter sphaeroides. J. Biol. Chem. 266, 14646–14653 (1991)
Guzman, L. M., Belin, D., Carson, M. J. & Beckwith, J. Tight regulation, modulation, and high-level expression by vectors containing the arabinose p-BAD promoter. J. Bacteriol. 177, 4121–4130 (1995)
Edmondson, D. L., Badger, M. R. & Andrews, T. J. A kinetic characterization of slow inactivation of ribulosebisphosphate carboxylase during catalysis. Plant Physiol. 93, 1376–1382 (1990)
Smith, J. M. Ximdisp—a visualization tool to aid structure determination from electron microscope images. J. Struct. Biol. 125, 223–228 (1999)
Mindell, J. A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003)
Frank, J. et al. SPIDER and WEB: processing and visualization of images in 3D electron microscopy and related fields. J. Struct. Biol. 116, 190–199 (1996)
Shaikh, T. R. et al. SPIDER image processing for single-particle reconstruction of biological macromolecules from electron micrographs. Nature Protocols 3, 1941–1974 (2008)
van Heel, M., Harauz, G., Orlova, E. V., Schmidt, R. & Schatz, M. A new generation of the IMAGIC image processing system. J. Struct. Biol. 116, 17–24 (1996)
Pettersen, E. F. et al. UCSF chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010)
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D Biol. Crystallogr. 62, 72–82 (2006)
Evans, P. R. Scala. CCP4 ESF-EACBM Newsl. Prot. Crystallogr. 33, 22–24 (1997)
Collaborative Computational Project No. 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994)
French, G. & Wilson, K. On the treatment of negative intensity observations. Acta Crystallogr. A 34, 517–525. (1978)
Van Duyne, G. D., Standaert, R. F., Karplus, P. A., Schreiber, S. L. & Clardy, J. Atomic structures of the human immunophilin FKBP-12 complexes with FK506 and rapamycin. J. Mol. Biol. 229, 105–124 (1993)
Schneider, T. R. & Sheldrick, G. M. Substructure solution with SHELXD. Acta Crystallogr. D Biol. Crystallogr. 58, 1772–1779 (2002)
de la Fortelle, E. & Bricogne, G. Maximum-likelihood heavy atom parameter refinement for multiple isomorphous replacement and multiwavelength anomalous diffraction methods. Methods Enzymol. 276, 472–494 (1997)
Terwilliger, T. C. Maximum-likelihood density modification. Acta Crystallogr. D Biol. Crystallogr. 56, 965–972 (2000)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997)
Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Cryst. 26, 283–291 (1993)
Kleywegt, G. T. & Jones, T. A. A super position. CCP4/ESF-EACBM Newsl. Prot. Crystallogr. 31, 9–14 (1994)
Gouet, P., Courcelle, E., Stuart, D. I. & Metoz, F. ESPript: multiple sequence alignments in PostScript. Bioinformatics 15, 305–308 (1999)
Acknowledgements
We thank S. Kaplan for providing the R. sphaeroides strain 2.4.1, S. Whitney for providing the pHue protein expression system, and R. Lange and N. Wischnewski for technical assistance. Support by the Max Planck Institute of Biochemistry (MPIB) Core Facility, the MPIB Crystallization Facility and the Joint Structural Biology Group staff at the European Synchrotron Radiation Facility beamlines is gratefully acknowledged. We thank the Deutsche Forschungsgemeinschaft (DFG) (SFB 594; DFG grant WE4628/1 to P.W.) and the Körber Foundation for financial support.
Author information
Authors and Affiliations
Contributions
O.M.-C. designed and performed all the biochemical experiments. O.M.-C., M.S. and A.B. obtained the CbbX crystals and solved the structure. P.W. performed the electron microscopy and three-dimensional image analysis. All authors contributed to data interpretation and manuscript preparation. O.M.-C., A.B., F.U.H. and M.H.-H. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Tables 1-2 and Supplementary Figures 1-5 with legends. (PDF 1551 kb)
Rights and permissions
About this article
Cite this article
Mueller-Cajar, O., Stotz, M., Wendler, P. et al. Structure and function of the AAA+ protein CbbX, a red-type Rubisco activase. Nature 479, 194–199 (2011). https://doi.org/10.1038/nature10568
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10568
This article is cited by
-
Grafting Rhodobacter sphaeroides with red algae Rubisco to accelerate catalysis and plant growth
Nature Plants (2023)
-
Elucidating the picocyanobacteria salinity divide through ecogenomics of new freshwater isolates
BMC Biology (2022)
-
The reliance of glycerol utilization by Cupriavidus necator on CO2 fixation and improved glycerol catabolism
Applied Microbiology and Biotechnology (2022)
-
Cloning and Characterization of the RubisCO Activase Gene from Pinus massoniana
Plant Molecular Biology Reporter (2022)
-
Gains and losses of metabolic function inferred from a phylotranscriptomic analysis of algae
Scientific Reports (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.