Abstract
Infection with the malaria parasite Plasmodium falciparum leads to widely different clinical conditions in children, ranging from mild flu-like symptoms to coma and death1. Despite the immense medical implications, the genetic and molecular basis of this diversity remains largely unknown2. Studies of in vitro gene expression have found few transcriptional differences between different parasite strains3. Here we present a large study of in vivo expression profiles of parasites derived directly from blood samples from infected patients. The in vivo expression profiles define three distinct transcriptional states. The biological basis of these states can be interpreted by comparison with an extensive compendium of expression data in the yeast Saccharomyces cerevisiae. The three states in vivo closely resemble, first, active growth based on glycolytic metabolism, second, a starvation response accompanied by metabolism of alternative carbon sources, and third, an environmental stress response. The glycolytic state is highly similar to the known profile of the ring stage in vitro, but the other states have not been observed in vitro. The results reveal a previously unknown physiological diversity in the in vivo biology of the malaria parasite, in particular evidence for a functional mitochondrion in the asexual-stage parasite, and indicate in vivo and in vitro studies to determine how this variation may affect disease manifestations and treatment.
Similar content being viewed by others
Main
To study the molecular basis of disease variation in malaria after infection with P. falciparum, we analysed the expression profiles of parasites derived directly from venous blood samples4,5 of 43 patients residing in Senegal, with a diverse age range (8.3 ± 6.9 years (mean ± s.d.)), and illness severity (parasitaemia 5.5% ± 6.2%, haematocrit 32.3 ± 6.8 (means ± s.d.)). Although previous studies found little variation between expression profiles of different P. falciparum strains in vitro3, we proposed that variation in the human host environment might affect P. falciparum biology and be reflected in its transcriptional profile.
We clustered the samples’ expression profiles, using a non-negative matrix factorization (NMF) algorithm6 (Fig. 1a, Supplementary Fig. 1 and Methods) and discovered that expression profiles cluster into three distinct groups. The profiles of samples in cluster 2 were similar to early ring-stage profiles of the 3D7 strain grown in vitro7,8,9 (for example, Spearman rank correlation 0.54 on average compared with ref. 7; Supplementary Fig. 2 and Supplementary Note 1). Ring stages predominate in the peripheral blood, and these were the only stages we observed in blood smears from the 43 samples (Supplementary Fig. 3). In contrast, expression profiles of samples in clusters 1 and 3 were not similar to those of early rings (0.12 and 0.26) or late stages (0.06 and 0.01) of the asexual parasite life cycle in vitro, and were only weakly similar to profiles of other developmental states such as gametocytes9 (0.31 and 0.23) or sporozoites (0.35 and 0.33; Supplementary Fig. 2 and Supplementary Note 1). They therefore represent novel transcriptional states. Profiles in clusters 1 and 2 are internally homogeneous and diametrically opposed, possibly reflecting a global transcriptional shift. Cluster 3 represents a third, distinct, pattern, although with more heterogeneity. Computational analysis indicates that profiles in cluster 3 are not a mixture of populations in cluster 1 and cluster 2 states (Supplementary Note 2).
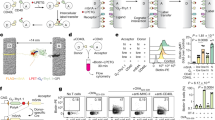
a, NMF clustering of expression profiles. The expression values for 3,937 P. falciparum genes (rows) across 43 samples (columns) are shown. Genes with very low expression were thresholded to a minimum value and filtered to exclude those that showed little variation across samples (Methods). Samples were first clustered by NMF and the genes were then sorted by their discrimination between cluster 1 versus all other samples. Each gene’s expression is normalized by mean centring and scaling (colour bar). The clustering identified three transcriptional states, two of which (clusters 1 and 2) are diametrically opposed and may represent a transcriptional shift. The number of clusters was determined objectively by the method, which does not force a structure on the data. The NMF clustering was repeated with samples derived from 2005 only (n = 31), and the cluster groups were unchanged (Supplementary Fig. 7). b, Clinical correlates of patients in each cluster. Shown are the median values and interquartile ranges of host demographic and selected laboratory values including cytokine measurements in the patients in each cluster. Statistically significant values (Mann–Whitney test with cluster 2 data as the reference group, P < 0.05) are designated by an asterisk. Cluster 3 is associated with significantly elevated inflammation markers, including duration of illness and body temperature and elevated levels of IL-6, IL-10, transforming growth factor (TGF)-α, tissue factor, vascular cell adhesion molecule (VCAM)-1 and lymphotactin. TNF, tumour necrosis factor.
The distinction between clusters 1 and 2 is not a reflection of patients’ measured parameters, of parasite genotypes or of different life cycle stages. There were no statistically significant differences between the clusters with respect to patients’ parameters, parasitological characterization (Fig. 1b), demographics or laboratory profiles. Parasite genotypes that identify distinct clones and number of clones in a single patient (MSP1/2) and chloroquine resistance (PFCRT K76T) showed no association with the clusters (data not shown). Furthermore, clusters 1 and 2 did not correlate with dates of sample collection, RNA isolation or oligonucleotide array hybridization. Examination of blood smears of each sample confirmed that only early ring stages were present (Supplementary Fig. 3) and the same clustering was observed with a set of 1,190 genes that do not vary during the parasite’s asexual life cycle7 (Supplementary Fig. 4).
To identify the physiological basis of the distinct transcriptional states, we compared the P. falciparum expression patterns with a compendium of 1,439 published expression profiles from the yeast S. cerevisiae (Methods and Supplementary Table 1). We mapped 1,247 S. cerevisiae genes to their P. falciparum orthologues (Methods) and then scored each S. cerevisiae profile for its similarity to the three expression clusters (Methods). For each cluster in P. falciparum, we identified a set of similar S. cerevisiae profiles and examined their biological annotations. We also used Gene Set Enrichment Analysis (GSEA)10 to test for the induction or repression of known pathways or functions (755 sets from P. falciparum; 328 sets from S. cerevisiae).
Each of the P. falciparum clusters was associated with a distinct set of S. cerevisiae responses (Fig. 2). Cluster 2 matched S. cerevisiae profiles associated with normal fermentative (glycolytic) growth (168/287 experiments, P = 2.3 × 10-23), cluster 1 matched profiles associated with starvation responses of S. cerevisiae (44/113, P = 1.5 × 10-7) as well as mutations in the general transcription machinery (23/53 experiments, P = 2.8 × 10-5). Cluster 3 was strongly associated with experiments on environmental stress in S. cerevisiae (278/438, P = 4.6 × 10-22).
Shown is a radial plot mapping of 1,439 array experiments from S. cerevisiae (circles) projected onto the expression space defined by the three P. falciparum NMF clusters (purple, green and brown squares corresponding to P. falciparum samples from each of clusters 1, 2 and 3, respectively). Yeast experiments associated with each cluster (Brier score ≥ 0.4) are highlighted with the corresponding colour (Methods).
This interpretation was also strongly supported by the induction of specific pathways and genes (Figs 3–5, Supplementary Table 2 and Supplementary Table 3). Cluster 2 showed induction of gene sets associated with glycolysis, amino-acid and nitrogen metabolism, and general growth processes such as nuclear transcription and cytoplasmic translation. By contrast, cluster 1 showed induction of gene sets associated with oxidative phosphorylation, respiration, mitochondrial biogenesis, the apicoplast, fatty-acid metabolism and genes involved in the uptake and metabolism of glycerol11,12,13 (Figs 3–5, Supplementary Table 2 and Supplementary Fig. 5). Thus, parasites in cluster 1 may rely on alternative pathways of energy production through the use of substrates such as glycerol, lactic acid, other carbon sources or lipids present in the patient’s blood. In addition, cluster 1 shows induction of genes related to invasion; this observation may be of clinical significance.
All the gene sets (rows) that differed significantly between cluster 1 and cluster 2 are shown, labelled by general categories. For each gene set, the mean expression of the ‘leading-edge’ genes (which supported the differential expression signature) in each experiment from the two clusters is shown (columns). The experiments are ordered as in Fig. 1. General biological categories describing the gene sets appear on the right; only gene sets with clear biological descriptions are included. Coloured bars indicate the number of genes in each gene set and in the leading edge.
Metabolic pathways derived from PlasmoCyc for glycolysis (glycolysis I), tricarboxylic acid cycle, aerobic respiration (electron donors reaction list) and glycerol degradation (glycerol degradation I) are shown13. The mean expression level for genes encoding the enzymes catalysing each reaction was calculated for cluster 1 and cluster 2. A ratio of expression for these values is indicated by colour bars. Red (blue) bars represent genes with at least twofold higher (lower) expression in cluster 1 versus cluster 2. Grey represents no change.
Relative expression of genes participating in major metabolic pathways. Hierarchical clustering of the expression values of the genes participating in glycolysis, the tricarboxylic acid cycle and fatty-acid metabolism30 in samples in cluster 1, cluster 2 and 3D7 early (E) and late (L) ring stages7. Names of glycolysis genes important for glycerol metabolism, including those encoding a glycerol transporter (PF11_0338) and aerobic glycerol catabolism enzymes (PF11_0660w, PF11_0157 and PF13_0269) are shown in red. The mean expression values for each gene in each cluster are reported in Supplementary Table 7. The relatively high expression level of genes involved in glycerol degradation and fatty-acid metabolism in cluster 1 compared with their expression in cluster 2 may suggest the use of alternative carbon sources for energy production.
Cluster 1 shows induction of cell-cycle related modules of both DNA replication and mitotic functions (Fig. 3), although the parasites in these samples were in the early ring stage (Supplementary Fig. 3). This induction explains some of the weak similarity of cluster 1 to some profiles from later stages of the asexual life cycle7,8 and from the sexual life cycle9 (Supplementary Fig. 2 and Supplementary Note 1). However, cluster 1 does not directly correspond to these developmental stages. This can be readily seen by examining key processes that are coherently induced in cluster 1. Although particular subsets of genes within these processes are induced at various points in the asexual cycle, there is no stage in the cycle that shows coherent induction of the genes within each process or of the overall collection of processes (Supplementary Note 1, Supplementary Fig. 6 and Supplementary Table 8).
What is the biological basis for the difference between clusters 1 and 2? Parasites of the reference strain are typically grown in vitro under glucose-rich and microaerophilic conditions, and they depend on anaerobic glycolysis for energy14. It has been widely assumed that exclusive reliance on anaerobic glycolysis represents the physiology of the asexual parasite in vivo. Cluster 2 is consistent with such glycolytic growth in vivo.
In contrast, cluster 1 indicates that a starvation response can lead to a metabolic shift in the asexual stage of P. falciparum and that respiration and metabolism of alternative carbon sources may be important in parasite physiology in vivo. This suggests that the metabolism of P. falciparum is consistent with that of the P. yoelii and P. berghei model systems15, which show active respiratory chains. Thus, parasites in vivo may exist in different states, as a result of varied oxygen or substrate levels. Although overall oxygen and substrate levels are tightly regulated in the human host, parasites are sequestered for half of their life cycle in the microvasculature, and oxygenation and substrate levels in this microenvironment can vary16,17. Furthermore, humans exhibit specific transcriptional changes when infected with Plasmodium18; our data indicate that the host environment may in turn affect parasite transcription.
Cluster 3 was strongly associated with S. cerevisiae profiles measured under environmental stress (for example heat shock, oxidative stress or osmotic stress) and also showed a clear correlation with the patients’ clinical phenotypes. In particular (Fig. 1b), the patients have a higher temperature, greater inflammation and elevated levels of the cytokines interleukin (IL)-6 and IL-10, which have been associated with more severe outcomes19. It has previously been demonstrated that parasite biology can change in response to environmental cues20. Additional samples from patients with severe disease will be needed to understand the clinical significance of this cluster.
Epigenetic mechanisms may have a role in the establishment of these transcriptional shifts. First, cluster 1 profiles resemble those observed in S. cerevisiae single-gene knockouts in general transcription factors (for example subunits of the Mediator, TFIID and SAGA complexes). These may be critical for the establishment of distinct transcriptional programmes. Second, the transcript encoding the CCAAT-binding protein is significantly induced in cluster 1. This protein is orthologous to the key regulator of oxidative phosphorylation genes from yeast to humans21,22. This factor may have a similar role in P. falciparum. More broadly, we found marked differences between clusters 1 and 2 in the expression of multiple genes encoding histones and chromatin modifiers (Supplementary Table 4), which may be critical for the establishment of stable and distinct transcriptional programmes in P. falciparum. Reproducing this transcriptional shift in vitro is critical for discovering its physiological and mechanistic basis.
Our observations about the apparent starvation response in samples in cluster 1 raise possible connections with gametogenesis. First, starvation responses typically cause yeast and other eukaryotic microbes to finish asexual growth and undergo meiosis. Second, respiratory and mitochondrial functions are known to be induced in gametocytes that have multiple mitochondria and higher oxygen consumption23. Third, the expression profiles in cluster 1 are more similar to late stages of in vitro gametogenesis9 than those in the other clusters, although the similarity is weak. Fourth, the expression of known gametogenesis genes9 is higher in cluster 1 samples than in cluster 2 (data not shown). Malaria parasites in the ring state choose between sexual and asexual fates long before morphological differences are apparent. Because gametocytes are isolated by the indiscriminate killing of immature sexual and asexual parasites, we know little about the metabolism or transcriptional programmes of these early sexual stages. It will be interesting to investigate whether the starvation response in cluster 1 may lead to a shift in vivo to a sexual form that allows the parasite to escape its starved host by transmitting through the mosquito vector into a new host. This hypothesis could be tested through studies of starvation in vitro and of parasite stages in vivo.
Pathogenesis studies in other systems have shown that organisms have distinct biology in vivo in comparison with in vitro models, and that some of these differences relate to virulence24. Little is known about the biology of Plasmodium residing in the human circulation. Our results show that the Plasmodium parasite exists in the human host in at least three distinct physiological states, apparently related to glycolytic growth, a starvation response and a general (non-nutritional) stress response. The relationships between these states and the course of clinical disease remain to be elucidated. Nevertheless, it is notable that cluster 1 shows strong induction of genes encoding proteins involved in invasion pathways, and cluster 3 is significantly associated with host inflammation. These novel states may result in enhanced virulence and the generation of metabolites such as reactive oxygen species, or in the consumption of substrates that could affect the host and contribute to disease severity17. Finally, if the distinct profiles represent persistent physiological differences, they may identify novel drug targets for malaria or may indicate possible alternative therapies.
Methods Summary
Patient population and sample handling
Venous blood samples from P. falciparum-infected patients in Senegal were directly added to Tri-Reagent BD (Molecular Research Center). This cohort consisted of patients who presented to the district hospital in Velingara, Senegal, with fever and symptoms suggestive of malaria. Enrolment criteria consisted of a P. falciparum infection of at least 1% of red blood cells. RNA was isolated, and steady-state parasite messenger RNA levels in 43 samples were determined with a custom-made Affymetrix chip based on the 3D7 genome as reported previously7.
Transcriptional analysis
The patient-derived transcriptional profiles were normalized with each other and with previously published in vitro data sets7,8,9 to allow direct comparisons. Samples were clustered by using NMF6, which finds a small number of gene combinations (metagenes) that best capture the behaviour of an expression data set. The number of clusters was determined using consensus clustering and maximizing the cophenetic correlation coefficient. Gene sets that are differentially expressed between clusters were identified by GSEA10, on the basis of a weighted Kolmogorov–Smirnov-like statistic. To project yeast expression data onto our parasite data set we first identified 1,247 S. cerevisiae genes that have P. falciparum orthologues. We then used metagene projection25 combined with a Support Vector Machine predictor to project 1,439 previously published26 S. cerevisiae expression profiles into the three metagene factor NMF representations described above (Supplementary Table 1) with a confidence level determined by a Brier score25. Experiments scoring highly in a given factor were associated with the P. falciparum cluster represented by that factor. We then used a hypergeometric enrichment test to identify biological conditions enriched in the profiles associated with each cluster. The complete data, gene sets, and associated analyses are available from http://carrier.gnf.org/publications/PatientProfiling and http://www.broad.mit.edu/compbio/pub/malaria
Online Methods
Patient population and study site
A field site was established in Velingara, a hyperendemic village in eastern Senegal, with peak transmission from October to December and an entomological inoculation rate of over 100 (ref. 27). Samples were collected during two transmission seasons, October to November in 2004 and 2005. Patients who required hospitalization or who appeared severely ill were enrolled in 2004. This cohort included two patients with asymptomatic hypoglycaemia, one patient with respiratory acidosis and one patient with coma. To obtain a larger sample size, all patients fulfilling enrolment criteria were enrolled in 2005, including those with minimal symptoms. Patients who presented to the hospital in Velingara were triaged by the local nurse to undergo malaria smear if they had symptoms suggestive of malaria. Enrolment criteria consisted of a P. falciparum infection without a second species noted on thin smear of 1% parasitaemia or greater. Of 1,187 patients screened for P. falciparum infection, 412 had a positive blood smear for P. falciparum and 95 fulfilled the enrolment criteria; and all consented to the study. After informed consent had been obtained, patients underwent venipuncture and one or two blood tubes (10–20 ml) coated with K3EDTA was collected. Tri-reagent BD (Molecular Research Center) was added within 5–10 min after blood collection. All samples were processed by a single person. The mixture was maintained at 4 °C until each evening, when it was placed in liquid nitrogen. Haematocrit was measured by microhaematocrit centrifugation. The remaining sample was centrifuged and divided into aliquots for serum studies, parasite cryopreservation, short-term culture, and application to filter paper for later DNA extraction. Cytokines, soluble endothelial-cell ligands and markers of inflammation were analysed from patient serum with a multiplex sandwich ELISA (Searchlight). Serum glucose levels were determined in Boston (on an Olympus AU 2700 analyser) from the transported frozen serum aliquots. Protocols were approved by the Harvard School of Public Health Human Subjects Committee and Senegal Ministry of Health Research Ethics Committee.
Detection of mRNA transcripts
The samples were shipped to Boston in liquid nitrogen, thawed at room temperature and total RNA was isolated in accordance with the manufacturer’s protocol (Tri reagent BD). Twelve samples from 2004 and 31 samples from 2005 that demonstrated sharp ribosomal bands on a denaturing agarose gel stained with ethidium bromide were selected for hybridization. Steady-state parasite mRNA levels were determined with a custom-made Affymetrix chip based on the 3D7 genome as reported previously7. Hybridizations were performed on three separate dates.
Data filtering and normalization
Each transcript was assigned a relative expression unit (EU) using MOID, used as reported previously28. A filtered gene list containing 3,937 genes was generated by thresholding gene expression levels to a minimum of 50 EU and removing any genes that varied less than threefold or 100 EU across the data set. To minimize potential effects of different dates of collection and hybridizations, the data was rank ordered by expression level and each gene was given an ordinal value. The published 3D7 reference strain data were processed in the same manner to allow comparisons7,9.
NMF clustering
The 43 P. falciparum-derived expression profiles were clustered by using NMF as described previously6 using the GenePattern software29. We chose a three-cluster solution, yielding the three distinct groups of samples, based on cluster-membership stability using consensus clustering (Supplementary Fig. 1). To determine whether the clustering is robust to the date of sample collection, we repeated NMF clustering using only the 31 samples collected in 2005 (Supplementary Fig. 7); these yielded the same results. The matrices derived from the NMF factorization give a description of the data in terms of three metagenes (three positive linear combinations of all the genes). Using the previously described metagene projection methodology25, we created an NMF projection of the data into three metagene factors, each corresponding to a compact representation of the associated cluster. To improve this projection, we equalized the number of samples to eight in each cluster. Cluster 1 had only eight samples. Because of the high degree of heterogeneity in cluster 3, we chose the eight samples in the other clusters to represent the widest range of behaviour. We then recomputed the NMF projection and used this final map to project the S. cerevisiae profiles into the same three-metagene representation.
Identification of S. cerevisiae–P. falciparum orthologues
We used the Kyoto Encyclopedia of Genes and Genomes30 SSDB database (KEGG) to find reciprocal best pairs of P. falciparum–S. cerevisiae genes with Smith–Waterman similarity scores of 100 or more. In all, 24% of the P. falciparum genome and 21% of the S. cerevisiae genome were included in the matched pairs.
Gene sets
P. falciparum gene annotations and pathways were obtained from KEGG, PlasmoDB, Hagai Ginsburg's Malaria Metabolic Pathways31 and Gene Ontology (GO)30,32,33. Yeast gene modules were constructed by following the procedure of ref. 34 using a yeast expression compendium described below and a total of 3,395 gene classes, including 1,794 from the GO35 hierarchy, 87 from KEGG, 107 from the BioCyc database36, 1,022 from the MIPS database of manually curated protein complexes37, 310 from a data set describing the genes whose promoters are bound by various transcription factors, 70 from a data set describing the genes that harbour a given cis-regulatory element in their promoter38, and 5 from a data set describing the genes whose RNA is bound by the RNA-binding proteins from the PUF family39. The yeast modules were mapped onto P. falciparum genes on the basis of the orthology relations described above.
GSEA
GSEA was performed as described previously10. In brief, the procedure assesses whether an a priori defined set of genes shows statistically significant, concordant differences between two biological states. Given a data set and two classes, genes are ranked on the basis of the correlation between their expression and the distinction between the two classes. GSEA uses a weighted Kolmogorov–Smirnov-like statistic to calculate an enrichment score that reflects the degree to which a gene set is overrepresented at the extremes of the entire ranked list. Within a gene set there is a leading-edge subset, which is defined as the genes that appear before the point in which the running sum enrichment score reaches its maximum deviation from zero. Because of the small number of samples in our study we estimated significance on the basis of a gene label (rather than class label) permutation. In our analyses we considered gene sets with a nominal P value below 0.01 and a false discovery rate (FDR) below 0.01 to be significant. The FDR for the invasion gene set is 0.07. We tested a total of 755 gene sets defined in P. falciparum and 328 sets originally defined in S. cerevisiae.
S. cerevisiae expression compendium
A compendium of 1,439 previously published S. cerevisiae expression profiles was compiled from the literature as described previously26 (Supplementary Table 1). Each experiment was manually annotated according to the experimental conditions, based on 20 categories (Supplementary Table 5); in addition each experiment was automatically annotated on the basis of the coherent induction or repression of each S. cerevisiae gene set (above) by following the procedure of ref. 34 (Supplementary Table 6).
Projection of S. cerevisiae experiments
Each S. cerevisiae expression profile was mapped into the P. falciparum gene space on the basis of the orthology assignments. Next, a Support Vector Machine predictor was used to project the S. cerevisiae expression profiles into the three-metagene-factor NMF representation described above. Experiments scoring highly in a given factor could be related to the P. falciparum cluster represented by that factor. Using a modified Brier skill score25, we measured a confidence level for each of these predictions. For each factor (cluster) we defined an associated set of the S. cerevisiae experiments that scored 0.4 or more for that factor. Next, we tested which array annotations were significantly enriched in each set of S. cerevisiae arrays by using the hypergeometric distribution to calculate a P value. The reported results were robust to the particular NMF model that we employed and to the threshold of Brier score used (ranging from 0.25 to 0.75).
Molecular analysis
The number of clones was determined by assessing MSP-1 and MSP-2 allelic variants as described previously40. Determination of the chloroquine-resistance-associated pfcrt K76T mutation was performed with PCR and restriction-fragment-length polymorphism using 3D7 chloroquine-sensitive and W2 chloroquine-resistant genomic DNA as controls41. To determine whether there were statistical differences in host features between clusters, the Mann–Whitney test was performed with Stata (version 9.0).
References
White, N. in Manson’s Tropical Diseases 21st edn (eds Cook, G. C. & Zumla, A. I.) 1205–1295 (Elsevier Science and W. B. Saunders, Edinburgh, 2002)
Greenwood, B., Marsh, K. & Snow, R. Why do some African children develop severe malaria? Parasitol. Today 7, 277–281 (1991)
Llinas, M., Bozdech, Z., Wong, E. D., Adai, A. T. & DeRisi, J. L. Comparative whole genome transcriptome analysis of three Plasmodium falciparum strains. Nucleic Acids Res. 34, 1166–1173 (2006)
Daily, J. P. et al. In vivo transcriptional profiling of P. falciparum . Malar. J. 3, 30 (2004)
Daily, J. P. et al. In vivo transcriptome of Plasmodium falciparum reveals overexpression of transcripts that encode surface proteins. J. Infect. Dis. 191, 1196–1203 (2005)
Brunet, J. P., Tamayo, P., Golub, T. R. & Mesirov, J. P. Metagenes and molecular pattern discovery using matrix factorization. Proc. Natl Acad. Sci. USA 101, 4164–4169 (2004)
Le Roch, K. G. et al. Discovery of gene function by expression profiling of the malaria parasite life cycle. Science 301, 1503–1508 (2003)
Bozdech, Z. et al. The transcriptome of the intraerythrocytic developmental cycle of Plasmodium falciparum . PLoS Biol. 1, E5 (2003)
Young, J. A. et al. The Plasmodium falciparum sexual development transcriptome: a microarray analysis using ontology-based pattern identification. Mol. Biochem. Parasitol. 143, 67–79 (2005)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Hansen, M., Kun, J. F., Schultz, J. E. & Beitz, E. A single, bi-functional aquaglyceroporin in blood-stage Plasmodium falciparum malaria parasites. J. Biol. Chem. 277, 4874–4882 (2002)
Promeneur, D. et al. Aquaglyceroporin PbAQP during intraerythrocytic development of the malaria parasite Plasmodium berghei . Proc. Natl Acad. Sci. USA 104, 2211–2216 (2007)
Yeh, I., Hanekamp, T., Tsoka, S., Karp, P. D. & Altman, R. B. Computational analysis of Plasmodium falciparum metabolism: organizing genomic information to facilitate drug discovery. Genome Res. 14, 917–924 (2004)
Lang-Unnasch, N. & Murphy, A. D. Metabolic changes of the malaria parasite during the transition from the human to the mosquito host. Annu. Rev. Microbiol. 52, 561–590 (1998)
Uyemura, S. A., Luo, S., Vieira, M., Moreno, S. N. & Docampo, R. Oxidative phosphorylation and rotenone-insensitive malate- and NADH-quinone oxidoreductases in Plasmodium yoelii yoelii mitochondria in situ . J. Biol. Chem. 279, 385–393 (2004)
Tsai, A. G., Johnson, P. C. & Intaglietta, M. Oxygen gradients in the microcirculation. Physiol. Rev. 83, 933–963 (2003)
Planche, T. & Krishna, S. Severe malaria: metabolic complications. Curr. Mol. Med. 6, 141–153 (2006)
Ockenhouse, C. F. et al. Common and divergent immune response signaling pathways discovered in peripheral blood mononuclear cell gene expression patterns in presymptomatic and clinically apparent malaria. Infect. Immun. 74, 5561–5573 (2006)
Lyke, K. E. et al. Serum levels of the proinflammatory cytokines interleukin-1β (IL-1β), IL-6, IL-8, IL-10, tumor necrosis factor α, and IL-12(p70) in Malian children with severe Plasmodium falciparum malaria and matched uncomplicated malaria or healthy controls. Infect. Immun. 72, 5630–5637 (2004)
Udomsangpetch, R. et al. Febrile temperatures induce cytoadherence of ring-stage Plasmodium falciparum-infected erythrocytes. Proc. Natl Acad. Sci. USA 99, 11825–11829 (2002)
Olesen, J., Hahn, S. & Guarente, L. Yeast HAP2 and HAP3 activators both bind to the CYC1 upstream activation site, UAS2, in an interdependent manner. Cell 51, 953–961 (1987)
Becker, D. M., Fikes, J. D. & Guarente, L. A cDNA encoding a human CCAAT-binding protein cloned by functional complementation in yeast. Proc. Natl Acad. Sci. USA 88, 1968–1972 (1991)
Krungkrai, J., Prapunwattana, P. & Krungkrai, S. R. Ultrastructure and function of mitochondria in gametocytic stage of Plasmodium falciparum . Parasite 7, 19–26 (2000)
Mahan, M. J., Slauch, J. M. & Mekalanos, J. J. Selection of bacterial virulence genes that are specifically induced in host tissues. Science 259, 686–688 (1993)
Tamayo, P. et al. Metagene projection for cross-platform, cross-species characterization of global transcriptional states. Proc. Natl Acad. Sci. USA 104, 5959–5964 (2007)
Marion, R. M. et al. Sfp1 is a stress- and nutrient-sensitive regulator of ribosomal protein gene expression. Proc. Natl Acad. Sci. USA 101, 14315–14322 (2004)
Faye, O. et al. Comparison of the transmission of malaria in 2 epidemiological patterns in Senegal: the Sahel border and the Sudan-type savanna. Dakar Med. 40, 201–207 (1995)
Zhou, Y. & Abagyan, R. Match-Only Integral Distribution (MOID) algorithm for high-density oligonucleotide array analysis. BMC Bioinformatics 3, 3 (2002)
Reich, M. et al. GenePattern 2.0. Nature Genet. 38, 500–501 (2006)
Kanehisa, M. A database for post-genome analysis. Trends Genet. 13, 375–376 (1997)
Ginsburg, H. Progress in in silico functional genomics: the Malaria Metabolic Pathways database. Trends Parasitol. 22, 238–240 (2006)
Bahl, A. et al. PlasmoDB: the Plasmodium genome resource. A database integrating experimental and computational data. Nucleic Acids Res. 31, 212–215 (2003)
Zhou, Y. et al. In silico gene function prediction using ontology-based pattern identification. Bioinformatics 21, 1237–1245 (2005)
Segal, E., Friedman, N., Koller, D. & Regev, A. A module map showing conditional activity of expression modules in cancer. Nature Genet. 36, 1090–1098 (2004)
Ashburner, M., Mungall, C. J. & Lewis, S. E. Ontologies for biologists: a community model for the annotation of genomic data. Cold Spring Harb. Symp. Quant. Biol. 68, 227–235 (2003)
Karp, P. D. et al. Expansion of the BioCyc collection of pathway/genome databases to 160 genomes. Nucleic Acids Res. 33, 6083–6089 (2005)
Mewes, H. W. et al. MIPS: analysis and annotation of proteins from whole genomes in 2005. Nucleic Acids Res. 34, D169–D172 (2006)
Harbison, C. T. et al. Transcriptional regulatory code of a eukaryotic genome. Nature 431, 99–104 (2004)
Gerber, A. P., Herschlag, D. & Brown, P. O. Extensive association of functionally and cytotopically related mRNAs with Puf family RNA-binding proteins in yeast. PLoS Biol. 2, E79 (2004)
Snounou, G. et al. Biased distribution of msp1 and msp2 allelic variants in Plasmodium falciparum populations in Thailand. Trans. R. Soc. Trop. Med. Hyg. 93, 369–374 (1999)
Happi, C. T. et al. Molecular analysis of Plasmodium falciparum recrudescent malaria infections in children treated with chloroquine in Nigeria. Am. J. Trop. Med. Hyg. 70, 20–26 (2004)
Acknowledgements
We thank the villagers and health care workers in Velingara, Senegal, for their participation in and support of this project; T. Taylor for critical advice and encouragement; G. Chechik, N. Barkai and O. Rando for providing parts of the compiled expression compendium for S. cerevisiae; and J. Bistline for assistance with the figures. J.P.D. is supported by the National Institute of Allergy and Infectious Diseases. N.P. is a Henri Benedictus Fellow of the King Baudouin Foundation and the Belgian American Educational Foundation. P.T. is supported by the National Institutes of Health (NIH). D.F.W. is supported by the Ellison Medical Foundation, the NIH, the Exxon Mobil Foundation and the Harvard School of Public Health. E.A.W. is supported by the Keck Foundation, the Novartis Research Foundation and the NIH. J.P.M. and D.S. are supported by the NIH and the National Science Foundation. A.R. is supported by a Career Award at the Scientific Interface from the Burroughs Wellcome Fund. The field work was supported by the Fogarty International from the NIH, the NIH Malaria Diversity grant, the Exxon Mobil Foundation, the Ellison Medical Foundation, the Burroughs–Welcome Fund and the Broad Institute of MIT and Harvard.
Author Contributions. D.S., N.P. and K.L.R. contributed equally to this work.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
The file contains Legends to Supplementary Figures 1-7 and Supplementary Tables 1-8; Supplementary Notes 1-2; Supplementary Figures 1-5 and 7; Supplementary Tables 1-4 and 7. (PDF 10117 kb)
Supplementary Table 5
This file contains Supplementary Table 5. (XLS 11 kb)
Supplementary Table 6
This file contains Supplementary Table 6. (XLS 108 kb)
Supplementary Table 8
This file contains Supplementary Table 8. (XLS 52 kb)
Supplementary Figure 6
This file contains Supplementary Figure 6. (PDF 30965 kb)
Rights and permissions
About this article
Cite this article
Daily, J., Scanfeld, D., Pochet, N. et al. Distinct physiological states of Plasmodium falciparum in malaria-infected patients. Nature 450, 1091–1095 (2007). https://doi.org/10.1038/nature06311
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06311
This article is cited by
-
High-throughput analysis of the transcriptional patterns of sexual genes in malaria
Parasites & Vectors (2023)
-
Parasite reliance on its host gut microbiota for nutrition and survival
The ISME Journal (2022)
-
Metabolome modulation of the host adaptive immunity in human malaria
Nature Metabolism (2021)
-
Plasmodium falciparum transcription in different clinical presentations of malaria associates with circulation time of infected erythrocytes
Nature Communications (2021)
-
Plasmodium infection inhibits tumor angiogenesis through effects on tumor-associated macrophages in a murine implanted hepatoma model
Cell Communication and Signaling (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.