Abstract
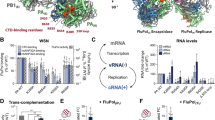
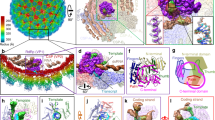
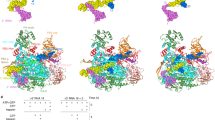
RNA polymerase (Pol) II catalyses DNA-dependent RNA synthesis during gene transcription. There is, however, evidence that Pol II also possesses RNA-dependent RNA polymerase (RdRP) activity. Pol II can use a homopolymeric RNA template1, can extend RNA by several nucleotides in the absence of DNA2, and has been implicated in the replication of the RNA genomes of hepatitis delta virus (HDV)3,4 and plant viroids5. Here we show the intrinsic RdRP activity of Pol II with only pure polymerase, an RNA template–product scaffold and nucleoside triphosphates (NTPs). Crystallography reveals the template–product duplex in the site occupied by the DNA–RNA hybrid during transcription. RdRP activity resides at the active site used during transcription, but it is slower and less processive than DNA-dependent activity. RdRP activity is also obtained with part of the HDV antigenome. The complex of transcription factor IIS (TFIIS) with Pol II can cleave one HDV strand, create a reactive stem-loop in the hybrid site, and extend the new RNA 3′ end. Short RNA stem-loops with a 5′ extension suffice for activity, but their growth to a critical length apparently impairs processivity. The RdRP activity of Pol II provides a missing link in molecular evolution, because it suggests that Pol II evolved from an ancient replicase that duplicated RNA genomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Dezelee, S., Sentenac, A. & Fromageot, P. Role of deoxyribonucleic acid–ribonucleic acid hybrids in eukaryotes. J. Biol. Chem. 249, 5978–5983 (1974)
Johnson, T. L. & Chamberlin, M. J. Complexes of yeast RNA polymerase II and RNA are substrates for TFIIS-induced RNA cleavage. Cell 77, 217–224 (1994)
Lai, M. M. C. RNA replication without RNA-dependent RNA polymerase: surprises from hepatitis delta virus. J. Virol. 79, 7951–7958 (2005)
Taylor, J. M. Replication of human hepatitis delta virus: recent developments. Trends Microbiol. 11, 185–190 (2003)
Rackwitz, H. R., Rohde, W. & Sanger, H. L. DNA-dependent RNA polymerase II of plant origin transcribes viroid RNA into full-length copies. Nature 291, 297–301 (1981)
Gnatt, A. L., Cramer, P., Fu, J., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: an RNA polymerase II elongation complex at 3.3 Å resolution. Science 292, 1876–1882 (2001)
Kettenberger, H., Armache, K.-J. & Cramer, P. Complete RNA polymerase II elongation complex structure and its interactions with NTP and TFIIS. Mol. Cell 16, 955–965 (2004)
Westover, K. D., Bushnell, D. A. & Kornberg, R. D. Structural basis of transcription: nucleotide selection by rotation in the RNA polymerase II active center. Cell 119, 481–489 (2004)
Kettenberger, H. et al. Structure of an RNA polymerase II–RNA inhibitor complex elucidates transcription regulation by noncoding RNAs. Nature Struct. Mol. Biol. 13, 44–48 (2006)
Kettenberger, H., Armache, K.-J. & Cramer, P. Architecture of the RNA polymerase II–TFIIS complex and implications for mRNA cleavage. Cell 114, 347–357 (2003)
Filipovska, J. & Konarska, M. M. Specific HDV RNA-templated transcription by pol II in vitro . RNA 6, 41–54 (2000)
Yamaguchi, Y. et al. Stimulation of RNA polymerase II elongation by hepatitis delta antigen. Science 293, 124–127 (2001)
Kireeva, M. L., Komissarova, N. & Kashlev, M. Overextended RNA:DNA hybrid as a negative regulator of RNA polymerase II processivity. J. Mol. Biol. 299, 325–335 (2000)
Toulokhonov, I. & Landick, R. The role of the lid element in transcription by E. coli RNA polymerase. J. Mol. Biol. 361, 644–658 (2006)
Naryshkina, T., Kuznedelov, K. & Severinov, K. The role of the largest RNA polymerase subunit lid element in preventing the formation of extended RNA–DNA hybrid. J. Mol. Biol. 361, 634–643 (2006)
Yamaguchi, Y., Mura, T., Chanarat, S., Okamoto, S. & Handa, H. Hepatitis delta antigen binds to the clamp of RNA polymerase II and affects transcriptional fidelity. Genes Cells 12, 863–875 (2007)
Kashlev, M. & Komissarova, N. Transcription termination: primary intermediates and secondary adducts. J. Biol. Chem. 277, 14501–14508 (2002)
Poole, A. M. & Logan, D. T. Modern mRNA proofreading and repair: clues that the last universal common ancestor possessed and RNA genome? Mol. Biol. Evol. 22, 1444–1455 (2005)
Wassarman, K. M. & Saecker, R. M. Synthesis-mediated release of a small RNA inhibitor of RNA polymerase. Science 314, 1601–1603 (2006)
Gildehaus, N., Neusser, T., Wurm, R. & Wagner, R. Studies on the function of the riboregulator 6S RNA from E. coli: RNA polymerase binding, inhibition of in vitro transcription and synthesis of RNA-directed de novo transcripts. Nucleic Acids Res. 35, 1885–1896 (2007)
Zenkin, N., Yuzenkova, Y. & Severinov, K. Transcript-assisted transcriptional proofreading. Science 313, 518–520 (2006)
Salgado, P. S. et al. The structure of an RNAi polymerase links RNA silencing and transcription. PLoS Biol. 4, e434 (2006)
Armache, K.-J., Kettenberger, H. & Cramer, P. Architecture of the initiation-competent 12-subunit RNA polymerase II. Proc. Natl Acad. Sci. USA 100, 6964–6968 (2003)
Brueckner, F., Hennecke, U., Carell, T. & Cramer, P. CPD damage recognition by transcribing RNA polymerase II. Science 315, 859–862 (2007)
Acknowledgements
We thank U. Hennecke and members of the Cramer laboratory for help, and J. Doudna, K. Förstemann, G. Meister and R. Schroeder for discussions. This work was supported by the Deutsche Forschungsgemeinschaft, the SFB646, the Nanoinitiative Munich, the Elitenetzwerk Bayern, the EU grant 3D repertoire, and the Fonds der Chemischen Industrie. The coordinates and structure factors for the RdRP EC and the HDVEC have been deposited in the Protein Data Bank under accession codes 2R92 and 2R93, respectively.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Tables 1-2, Supplementary Figures 1-6 with Legends and additional references. (PDF 491 kb)
Rights and permissions
About this article
Cite this article
Lehmann, E., Brueckner, F. & Cramer, P. Molecular basis of RNA-dependent RNA polymerase II activity. Nature 450, 445–449 (2007). https://doi.org/10.1038/nature06290
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature06290
This article is cited by
-
Structure of transcribing RNA polymerase II-nucleosome complex
Nature Communications (2018)
-
The Hepatitis Delta Virus accumulation requires paraspeckle components and affects NEAT1 level and PSP1 localization
Scientific Reports (2018)
-
The emerging role of RNAs in DNA damage repair
Cell Death & Differentiation (2017)
-
An RNA-dependent RNA polymerase gene in bat genomes derived from an ancient negative-strand RNA virus
Scientific Reports (2016)
-
Shared active site architecture between archaeal PolD and multi-subunit RNA polymerases revealed by X-ray crystallography
Nature Communications (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.