Abstract
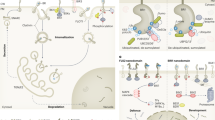
Both animals and plants use steroids as signalling molecules during growth and development. Animal steroids are principally recognized by members of the nuclear receptor superfamily of transcription factors1. In plants, BRI1, a leucine-rich repeat (LRR) receptor kinase localized to the plasma membrane, is a critical component of a receptor complex for brassinosteroids2,3. Here, we present the first evidence for direct binding of active brassinosteroids to BRI1 using a biotin-tagged photoaffinity castasterone (BPCS), a biosynthetic precursor of brassinolide (the most active of the brassinosteroids). Binding studies using BPCS, 3H-labelled brassinolide and recombinant BRI1 fragments show that the minimal binding domain for brassinosteroids consists of a 70-amino acid island domain (ID) located between LRR21 and LRR22 in the extracellular domain of BRI1, together with the carboxy-terminal flanking LRR (ID-LRR22). Our results demonstrate that brassinosteroids bind directly to the 94 amino acids comprising ID-LRR22 in the extracellular domain of BRI1, and define a new binding domain for steroid hormones.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Aranda, A. & Pascual, A. Nuclear hormone receptors and gene expression. Physiol. Rev. 81, 1269–1304 (2001)
Li, J. & Chory, J. A putative leucine-rich repeat receptor kinase involved in brassinosteroid signal transduction. Cell 90, 929–938 (1997)
Wang, Z. Y. et al. BRI1 is a critical component of a plasma-membrane receptor for plant steroids. Nature 410, 380–383 (2001)
Clouse, S. D. Brassinosteroid signal transduction: clarifying the pathway from ligand perception to gene expression. Mol. Cell 10, 973–982 (2002)
Thummel, C. S. & Chory, J. Steroid signaling in plants and insects—common themes, different pathways. Genes Dev. 16, 3113–3129 (2002)
Clouse, S. D., Langford, M. & McMorris, T. C. A brassinosteroid-insensitive mutant in Arabidopsis thaliana exhibits multiple defects in growth and development. Plant Physiol. 111, 671–678 (1996)
Schumacher, K. & Chory, J. Brassinosteroid signal transduction: still casting the actors. Curr. Opin. Plant Biol. 3, 79–84 (2000)
Altmann, T. Molecular physiology of brassinosteroids revealed by the analysis of mutants. Planta 208, 1–11 (1999)
Clouse, S. & Feldmann, K. in Brassinosteroids: Steroidal Plant Hormones (eds Sakurai, A., Yokota, T. & Clouse, S.) 163–190 (Springer-Verlag, Tokyo, 1999)
Kauschmann, A. et al. Genetic evidence for an essential role of brassinosteroids in plant development. Plant J. 9, 701–713 (1996)
Noguchi, T. et al. Brassinosteroid-insensitive dwarf mutants of Arabidopsis accumulate brassinosteroids. Plant Physiol. 121, 743–752 (1999)
He, Z. et al. Perception of brassinosteroids by the extracellular domain of the receptor kinase BRI1. Science 288, 2360–2363 (2000)
Caño-Delgado, A. et al. BRL1 and BRL3 are novel brassinosteroid receptors that function in vascular differentiation in Arabidopsis. Development 131, 5341–5351 (2004)
Li, J. et al. A role for brassinosteroids in light-dependent development of Arabidopsis. Science 272, 398–401 (1996)
Friedrichsen, D. M. et al. Brassinosteroid-insensitive-1 is a ubiquitously expressed leucine-rich repeat receptor serine/threonine kinase. Plant Physiol. 123, 1247–1255 (2000)
Yin, Y. et al. BES1 accumulates in the nucleus in response to brassinosteroids to regulate gene expression and promote stem elongation. Cell 109, 181–191 (2002)
Seto, H. et al. Preparation, conformational analysis, and biological evaluation of 6a-carbabrassinolide and related compounds. Tetrahedron 58, 9741–9749 (2002)
Yin, Y., Wu, D. & Chory, J. Plant receptor kinases: systemin receptor identified. Proc. Natl Acad. Sci. USA 99, 9090–9092 (2002)
Li, J. et al. BAK1, an Arabidopsis LRR receptor-like protein kinase, interacts with BRI1 and modulates brassinosteroid signaling. Cell 110, 213–222 (2002)
Nam, K. H. & Li, J. BRI1/BAK1, a receptor kinase pair mediating brassinosteroid signaling. Cell 110, 203–212 (2002)
Wurtz, J.-M. et al. A canonical structure for the ligand-binding domain of nuclear receptors. Nature Struct. Biol. 3, 87–94 (1996)
Kobe, B. & Deisenhofer, J. Crystal structure of porcine ribonuclease inhibitor, a protein with leucine-rich repeats. Nature 366, 751–756 (1993)
Di Matteo, A. et al. The crystal structure of polygalacturonase-inhibiting protein (PGIP), a leucine-rich repeat protein involved in plant defense. Proc. Natl Acad. Sci. USA 100, 10124–10128 (2003)
Ward, C. W. & Garrett, T. P. J. The relationship between the L1 and L2 domains of the insulin and epidermal growth factor receptors and leucine-rich repeat modules. BMC Bioinformatics 2, 4 (2001)
Seto, H. et al. A general approach to synthesis of labeled brassinosteroids: preparation of [25,26,27-2H7]brassinolide with 60% isotopic purity from the parent brassinolide. Tetrahedr. Lett. 39, 7525–7528 (1998)
Konoki, K. et al. Development of biotin–avidin technology to investigate okadaic acid-promoted cell signaling pathway. Tetrahedron 56, 9003–9014 (2000)
Gallagher, S., Winston, S. E. & Hurrell, J. G. R. in Current Protocols in Molecular Biology (eds Ausubel, F. M. et al.) 10.8.1–10.8.16 (Greene Publishing & Wiley-Interscience, New York, 1992)
Acknowledgements
We thank S. Richard and J. Noel for discussions, and Y. Zhao and S. Mora-Garcı´a for reading the manuscript and providing critical comments. This work was funded by grants from the USDA and HFSP to J.C., by a Grant-in-Aid for Young Scientist (A) to T.K. and Grant-in-Aid for scientific research (C) to H.S. from the Ministry of Education, Culture, Sports, Science and Technology of Japan, and by an HFSP fellowship to A.C.D. J.C. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Supplementary Figure S1
Preparation of biotin-tagged photoaffinity castasterone (BPCS). (PDF 341 kb)
Supplementary Figure S1 Legend
Text to accompany the above Supplementary Figure. (PDF 50 kb)
Supplementary Figure S2
Competition for [3H]-labelled BL binding to membrane fractions of BRI1–GFP plants by BL and BPCS. Includes legend for Supplementary Figure S2. (PDF 301 kb)
Supplementary Figure S3
3H-labelled BL binding in membrane fractions from bri1-119. Includes legend for Supplementary Figure S3. (PDF 241 kb)
Supplementary Figure S4
3H–BL finding is unaltered in a bak1 null mutant. Includes legend for Supplementary Figure S4. (PDF 61 kb)
Rights and permissions
About this article
Cite this article
Kinoshita, T., Caño-Delgado, A., Seto, H. et al. Binding of brassinosteroids to the extracellular domain of plant receptor kinase BRI1. Nature 433, 167–171 (2005). https://doi.org/10.1038/nature03227
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature03227
This article is cited by
-
The LRR receptor-like kinase ALR1 is a plant aluminum ion sensor
Cell Research (2024)
-
How the Wild Sugarcane Resource Miscanthus floridulus Responds to Low-Temperature Stress: A Physiological and Transcriptomic Analysis
Sugar Tech (2023)
-
Cloning, characterization and expression analysis of a brassinosteroids biosynthetic gene VvDET2 in Cabernet Sauvignon (Vitis vinifera L.)
Plant Cell, Tissue and Organ Culture (PCTOC) (2023)
-
The kinesin-13 protein BR HYPERSENSITIVE 1 is a negative brassinosteroid signaling component regulating rice growth and development
Theoretical and Applied Genetics (2022)
-
Integrated genetic mapping and transcriptome analysis reveal the BnaA03.IAA7 protein regulates plant architecture and gibberellin signaling in Brassica napus L.
Theoretical and Applied Genetics (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.